This page was written by Elizabeth Canfield '99
Sanger Method for DNA Sequencing
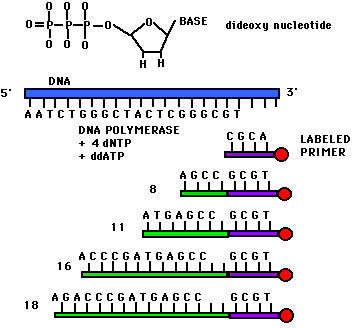
Dideoxynucleotide sequencing represents only one method of sequencing DNA. It is commonly called Sanger sequencing since Sanger devised the method. This technique utilizes 2',3'-dideoxynucleotide triphospates (ddNTPs), molecules that differ from deoxynucleotides by the having a hydrogen atom attached to the 3' carbon rather than an OH group. (Figure 1). These molecules terminate DNA chain elongation because they cannot form a phosphodiester bond with the next deoxynucleotide.
In order to perform the sequencing, one must first convert double stranded DNA into single stranded DNA. This can be done by denaturing the double stranded DNA with NaOH. A Sanger reaction consists of the following: a strand to be sequenced (one of the single strands which was denatured using NaOH), DNA primers (short pieces of DNA that are both complementary to the strand which is to be sequenced and radioactively labelled at the 5' end), a mixture of a particular ddNTP (such as ddATP) with its normal dNTP (dATP in this case), and the other three dNTPs (dCTP, dGTP, and dTTP). The concentration of ddATP should be 1% of the concentration of dATP. The logic behind this ratio is that after DNA polymerase is added, the polymerization will take place and will terminate whenever a ddATP is incorporated into the growing strand. If the ddATP is only 1% of the total concentration of dATP, a whole series of labeled strands will result (Figure 1). Note that the lengths of these strands are dependent on the location of the base relative to the 5' end.
This reaction is performed four times using a different ddNTP for each reaction. When these reactions are completed, a polyacrylamide gel electrophoresis (PAGE) is performed. One reaction is loaded into one lane for a total of four lanes (Figure 2). The gel is transferred to a nitrocellulose filter and autoradiography is performed so that only the bands with the radioactive label on the 5' end will appear. In PAGE, the shortest fragments will migrate the farthest. Therefore, the bottom-most band indicates that its particular dideoxynucleotide was added first to the labeled primer. In Figure 2, for example, the band that migrated the farthest was in the ddATP reaction mixture. Therefore, ddATP must have been added first to the primer, and its complementary base, thymine, must have been the base present on the 3' end of the sequenced strand. One can continue reading in this fashion. Note in Figure 2 that if one reads the bases from the bottom up, one is reading the 5' to 3' sequence of the strand complementary to the sequenced strand. The sequenced strand can be read 5' to 3' by reading top to bottom the bases complementary to the those on the gel.

Figure 1. This figure shows the structure of a dideoxynucleotide (notice the H atom attached to the 3' carbon). Also depicted in this figure are the ingredients for a Sanger reaction. Notice the different lengths of labeled strands produced in this reaction.
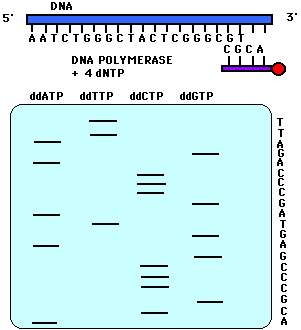
Figure 2. This figure is a representation of an acrylamide sequencing gel. Notice that the sequence of the strand of DNA complementary to the sequenced strand is 5' to 3' ACGCCCGAGTAGCCCAGATT while the sequence of the sequenced strand, 5' to 3', is AATCTGGGCTACTCGGGCGT.
This figure was taken with permission from <http://www.plattsburgh.edu/acadvp/artsci/biology/bio401/DNASeq.html>
If you would like to learn about a variation of Sanger sequencing, click on cycle sequencing.
© Copyright 2001 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu