This web page was produced as an assignment for an undergraduate course at Davidson College.
IDP3
Accession Number : NC 001146
Six-Cutter Restriction Sites :
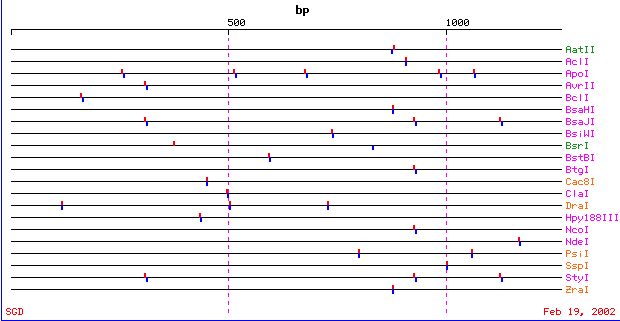
Figure 5. Restriction map of IDP3 showing the six-cutter restriction sites. Figure adapted from <http://genome-www2.stanford.edu/cgi-bin/SGD/PATMATCH/RESTmap?id=20144&beg=1&type=6base>
PCR Primers :
forward: 5' ATGAGTAAAATTAAAGTTGT 3' Tm = 44 degrees Celsius
reverse: 5' TTATAGTTTGCACATACCTT 3' Tm = 44 degrees Celsius
In-Frame Fusion Protein of His6 + IDP3 :
MRGSHHHHHH-GSHVISSIASMSKIKVVHPI
(AA sequence of MRGS-His epitope = MRGSHHHHHH)
Coenzyme: NADP+
Experiment Design (to insure proper orientation) :
The IDP3 gene can insert in either the forward or reverse orientation into the expression vector. The only functional orientation is the forward orientation, so we need to design an experiment that selects for only this orientation. The experiment is as follows: Transfect E. coli cells with the expression vectorand allow the E. coli to proliferate so that many copies of the desired vector are produced. Then, digest selected colonies of the E. coli cells with a restriction enzyme that will cut once inside the gene and once in the rest of the vector. In the case of IDP3, we had to digest with two restriction enzymes ClaI(cut once inside the gene but not in the rest of the vector) and EcoRV (cut once in the vector but not in the gene) Run the restriction fragments of these selected colonies on a gel electrophoresis. If the fragments are ~510 bp and ~4250 bp long, you have the forward orientation. Separate these colonies and allow them to continue to proliferate. The reverse orientation would give fragments of ~780 bp and ~3980 bp. These colonies can be discarded.
If you have any questions or comments about this web page, please e-mail me, Chris Norbet