This web page was produced as an assignment for an undergraduate course at Davidson College
 CD4: T-cell surface glycoprotein
CD4: T-cell surface glycoprotein
The Function and Structure of CD4
The Human Immune System: Our Defense Mechanism
The human immune system protects the body from foreign molecules such as bacteria, viruses, toxins, and parasites that invade the body and may cause harm. The immune system consists of highly specialized cells as well as a circulatory system that is separate from the blood vessels in order to rid the body of infection. The immune system functions by being able to recognize molecules that are not "self"; that is to say molecules that are not a part of any proteins normally produced in the human body. The cells of the immune system that perform this function are lymphocytes, which exist in two forms: T cells and B cells. B cells make antibodies, which can bind to a foreign molecule and trigger many resposes to destroy the invader. T cells can be subdivided into two groups: T helper cells and cytotoxic T lyphocytes. Cytotoxic T lymphocytes kill foreign cells directly through membrane-to-membrane contact while T helper cells aid the other immune system cells to perform their functions efficiently. For example, T helper cells help B cells to generate antibodies in the humoral immune response, they help cytotoxic T lymphocytes in killing foreign cells, and they release lymphokines which produce activation signals within the immune system. The T helper cells are particularly important because they play an integral role in ALL immune responses.
CD4: A receptor protein on the surface of T helper cells
Figure 1: Chime image of CD4 generated by the University of Rochester Medical Center
CD4 is a 55kd cell surface molecule found on a subset of T lymphocytes utilizing MHC class II antigens as a restriction element for antigen recognition (Camerini et al., 1990). The mature CD4 protein consists of consists of a 372 amino acid extracellular segment composed of four tandem immunoglobulin-like VJ regions, a 23 amino acid transmembrane domain, and a 33 amino acid cytoplasmic segment. The cDNA sequence is composed 1742 base pairs and 450 amino acids are present in the mature protein. The CD4 locus maps to 12p12-pter and contains 10 exons. CD stands for "cluster of differentiation" and and CD4 is the official destination for the T cell antigen T4/leu3. CD4 is not linked to any known member of the immunoglobin gene family, including its counterpart gene CD8 which is located on human chromosome 2. The overall structure of T4 consists of an N-terminal variable (V)-like domain, a joining (J)-like region, a third extracellular domain, a membrane-spanning region homologous to class II major histocompatibility complex (MHC) beta-chains, and a highly charged cytoplasmic domain (Maddon et al., 1985).
As seen in Figure 1 CD4 is a single chain molecule composed of four immunoglobin-like domains. CD4 binds to MHC class II molecules through a region that is mainly on the lateral face of the first domain. CD4 also interacts with a cytoplasmic tyrosine kinase known as Lck. When CD4 and the T-cell receptor can simultaneously bind to the same MCH class II:peptide complex, the sensitivity of a T cell to an antigen is strongly increased. CD4 binding to an MHC class II molecule on its own is very weak and it is possible that this binding would not be strong enought to transmit a signal to the interior of a T cell. The CD4 molecule is also capable of forming homodimers and it is not yet clear whether or not the dimerization of CD4 plays a role in its function as a coreceptor on the T cell surface. T helper cells are also known as CD4 positive T cells (CD4+ T cells) and these cells alert B cells to start making antibodies; they also can activate other T cells and immune system scavenger cells called macrophages and influence which type of antibody is produced. T helper cells are activated when a macrophage engulfs and processes an antigen and then displays fragments of that antigen in combination with a class II MHC Protein on its cell surface. This antigen-protein combination attracts a T helper cell and promotes its activation (National Institute of Allegy and Infectious Diseases). This concept is diagramed in Figure 1.
Figure 2: Activation of a T helper cell by an antigen presenting macrophage.
Research involving the CD4 gene began in the early 1980s whcn the human aquired immune deficiency virus (AIDS) began to create a national epidemic. In 1986, it became clear that the T4 gene (which later became known as the CD4 gene) may not only serve as a receptor for molecules on the surface of target cells, but also as a receptor for the AIDS virus. Monoclonal antibodies directed against T4 were able to block AIDS virus infection, thus it became clear that CD4 could potentially play an important role in acquiring more information about the AIDS virus (Maddon et. al., 1986). In these early studies, the T4 gene was sequenced and cloned so that the interaction between the molecules on the surface of the HIV virus and the T4 molecule could be investigated. It was initially hypothesized that the HIV virus could enter the human body by two mechanisms: some viruses fuse directly with the plasma membrane whereas others are internalized by receptor mediated endocytosis. A study done in 1986 was able to provide good evidence to suggest that AIDS virus infection occurred as a direct result of interaction of the viral molecules with T4 molecules on the cell surface (Maddon et al., 1986). This information was determined after the T4 gene had been sequenced in previous investigations (Maddon et. al., 1985).
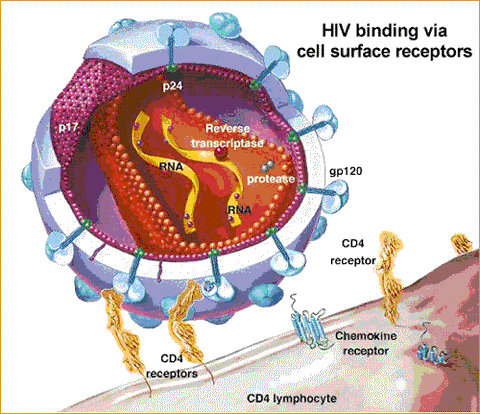
Viruses that enter the body often target certain cells based upon binding capabilities between the viral envelop and and the membrane bound protiens on a cell. CD4, a protein found on the surface of T helper cells, is the protein targeted by the HIV virus. While CD4 alone is not sufficient to induce binding with the HIV virus, it has been proven to be necessary for binding. It has been found that CD4 as well as CXCR4 or CCR5 are necessary for binding. The diagram below provides a view of these interactions.
Figure 3: The binding of HIV to a T helper cell which possesses the CD4 receptor and a coreceptor (either CXCR4 or CCR5).
CXCR4 and CCR5 are coreceptors, that is they participate in the binding of the HIV virus to the surface of the T cell. The protein on the surface of the HIV viral envelop that binds to CD4 is known as gp120.
What Happens When the CD4 Gene is Mutated?
While mutations within the CD4 gene have not been investigated in great detail, mutations are often induced in the gene for purposes of study. For example, in the determination of the interaction between CD4 and gp120, many mutations where induced in the gene to determine which region of the gene was crucial to binding. One study concluded that that residues 41-43 and 51-55 were important for gp120 binding. Also, the study concluded that substitution in the first region of valine for phenylalanine at position 43 results in a striking reduction in binding. This binding is further reduced when the serine at poition 42 is changed to glycine. Binding was also disrupted when amino acids were substituted in the 51-55 region and also showed reduced binding upon deletion of these regions (Arthos et al., 1989).
Mutations which contribute to HIV resistance do not involve the CD4 gene, however, the mutation effects the ability of CD4 to bind to gp120. The gene under study encodes a cell surface receptor called CCR5 that HIV-1 usually requires (in addition to the CD4 receptor) to invade CD4+ T cells. Scientists have long been aware that a mutation called CCR5-32 results in a faulty receptor, and that people who inherit two copies of the mutated gene (and, thus, have no working CCR5 receptors) are highly resistant to HIV infection (Stephenson, 2001). The study suggests that strategies for blocking CCR5 receptors may have utility for reducing [HIV] transmission, and if so it might not be necessary to block all of the receptors to obtain some protection. Reducing the number of receptor sites per cell may be adequate to provide an imperfect but important degree of protection.
CD4 Orthology in Other Species
Phylogenetic studies have shown that lymphocytes from distant primate species are susceptible to HIV infection in vitro. In vivo, however, infection and persistent viremia have only been demonstrated in chimpanzees. Despite the fact that chimpanzees are susceptible to infection, they do not appear to develop the fatal immunodeficiency that occurs in humans. One difference between human and nonhuman primate CD4 is the inability of the latter to support syncytium formation. The study concluded that this was due to one amino acid difference (at position 87) between human and chimpanzee CD4. The study also reported that Rhesus CD4 shares 92% identity with human CD4. Below is a comparison of the amino acid sequences (Camerini et. al., 1990).
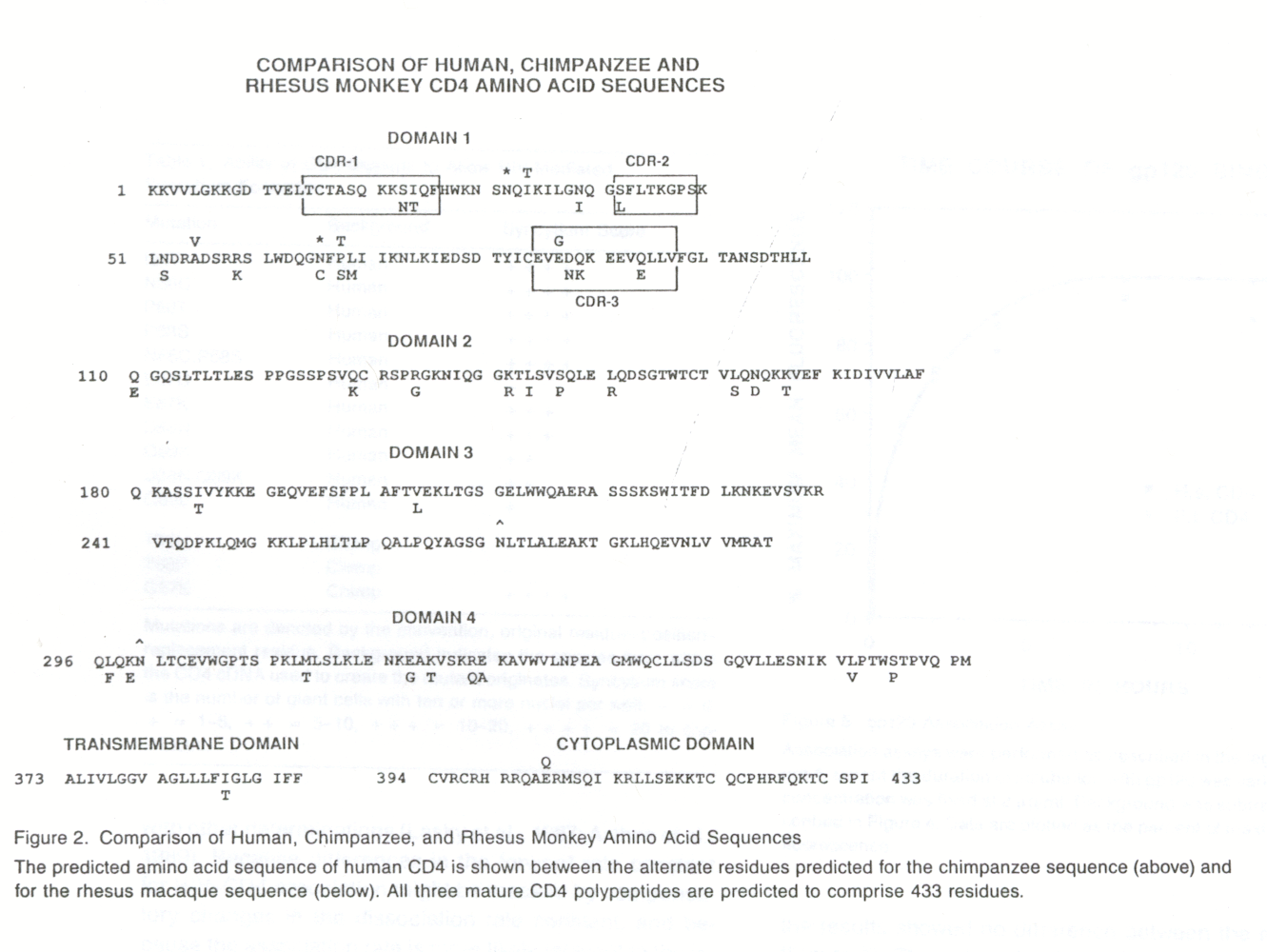
Figure 4: See figure caption above marked Figure 2.
There is also a mouse ortholog to CD4 which is located on the mouse chromosome 6 (Ansari-Lari et al., 1998). The amino acid sequence of the mouse ortholog can also be compared to the human sequence.
Important Links Related to HIV Infection
Official Journal of the International AIDS Society
References
Ansari-Lari MA, Muzny DM, Lu J, Lu F, Lilley CE, Spanos S, Malley
T, Gibbs RA. 1996, A gene-rich cluster between the CD4 and triosephosphate
isomerase genes at human chromosome 12p13 [abstract]. In Genome Research 6:314-326.
PubMed Database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=8723724&dopt=Abstract>.
Accessed 2003 March 12.
Arthos J, Deen FA, Chaikin MA, Fornwald JA, Sathe G, Sattentau QJ, Clapham PR, Weiss SJ, Pietropaolo RA, Truneh A, Maddon PJ, Sweet RW. 1989 May, Indentification of the Residues In Human CD4 Critical for the Binding of HIV [abstract]. In Cell 57:469-481. PubMed Database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=2541915&dopt=Abstract> Accessed 2003 March 12.
Biores Structure of CD4. <http://bioresearch.ac.uk/whatsnew/detail/1099769.html>. Accessed 2003 March 13.
Camerini D, Seed B. 1990 March, A CD4 Domain Important for HIV-Mediated Synytium Formation Lies outside the Virus Binding Site [abstract]. In Cell 60:747-754. PubMed Database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=2107024&dopt=Abstract>. Accessed 2003 March 12.
Campbell, A.M., Bernd, K. and Serie, J. Introductory Biology 111: Cell and Molecular Biology Study Guide. Davidson College: Revised June 2001.
JAMA Medical News and Perspectives. 2001. JAMA Medical News and Perspectives Archives. <http://jama.ama-assn.org/issues/v286n12/ffull/jmn0926-3.html>. Accessed 2003 March 13.
Janeway, Charles A.; Travers, Paul; Walport, Mark. Shlomchik, Mark. Immunobiology. New York: Garland Publishing, 2001.
Maddon PJ, Littman DR, Godfrey M, Maddon DE, Chess L, Axel R. 1985 August, The isolation and nucleotide sequence of a cDNA encoding the T cell surface protein T4: a new member of the immunoglobulin gene family. [abstract]. In Cell 42:93-104. PubMed Database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=2990730&dopt=Abstract>. Accessed 2003 March 12.
Maddon PJ, Dalgleish A, McDougal JS, Clapham PR, Weiss RA, Axel R. 1986 November. The T4 gene encodes the AIDS virus receptor and is expressed in the immune system and the brain [abstract]. In Cell 47:333-348. PubMed Database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=3094962&dopt=Abstract>. Accessed 2003 March 12.
NIAID Net News, The Immune System. <http://www.niaid.nih.gov/final/immun/immun.htm> Accessed 2003 March 13.
National Center for Biotechnology Information Database. <http://www.ncbi.nlm.nih.gov/>. Accessed 2003 March 12.
University of Rochester Medical Center Structure of CD4. <http://www.urmc.rochester.edu/smd/mbi/chime/cd4frame.html>. Accessed 2003 March 14.
Davidson College Biology Department
Davidson College Molecular Biology Home
Questions or comments contact: Erin Cobain