"This web page was produced as an assignment for an undergraduate course at Davidson College."
Krista's Review of:
Nature. Vol 434: 759-763. April 2005.
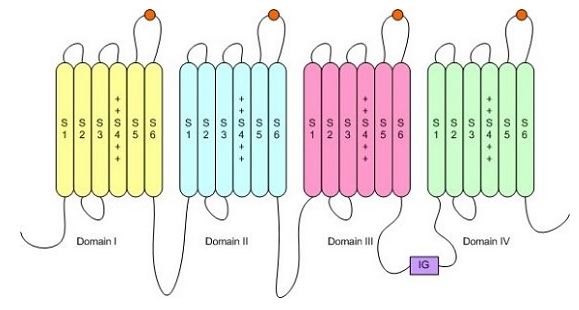
This paper examined the genetic and molecular basis for an adaptation which makes garter snakes (Thamnophis sirtalis) resistant to tetrodotoxin (TTX) that is found in newts (Taricha granulosa) that they have co-evolved with. TTX is a potent toxin and causes paralysis by binding to the outer pore of sodium channels and blocking nerve and muscle fibre activity (Geffeney et al., 2005). Sodium channels are essential to nerve and muscle function by allowing sodium ions to flow into the cell and thus propagating the action potential.The sodium channel is composed of four domains, each of which contain multiple transmembrane segments. TTX binds to the sodium channel on the connecting segments in each domain as depicted by the orange circles (Smith, 2005).

Figure 1: Structure of the sodium ion channel. Orange circles depict places where TTX binds. http://www.chemsoc.org/exemplarchem/entries/2002/Tim_Smith/about/ (permission pending)
The garter snake preys on the newt, and both have co-evolved in a sort of "arms race" with the newts producing more toxic TTX as the garter snake becomes more resistant. There is high selective pressure for this resistance and it is believed that TTX resistance has occurred at least twice in garter snakes (Geffeney et al., 2005). It is known that TTX-sensitive animals have an aromatic amino acid in the outer pore of domain I of the sodium ion channel, which has a high affinity for TTX. TTX-resistant animals have a non-aromatic amino acid in domain I, which inhibits binding of TTX and causes resistance (Geffeney et al., 2005). This paper is testing populations of garter snakes that have an aromatic amino acid in domain I, but are still resistant to TTX. Therefore, it is testing the possibility that additional mutations in the amino acid sequence of the sodium ion channel, change the structure, inhibit the binding of TTX, and provide resistance to the toxin.
To test this, they first isolated the TTX-resistant sodium channel gene by making a cDNA library of TTX-resistant snakes and screening the library with probes of skeletal and cardiac sodium ion channel genes. Then they sequenced the corresponding cDNA fragments and found a gene with 5,625 nucleotides that translates into 1,875 amino acids (Geffeney et al., 2005). This gene is very similar to other sodium ion channel genes called NAv1.4, so this gene was named tsNAv1.4. They found that this gene contained amino-acid substitutions in the pore helix and B-strand of domain IV which are also important regions for TTX binding (Geffeney et al., 2005). To test the functional significance of these mutations they expressed TTX-resistant sodium ion channels in Xenopus oocytes and measured the changes in TTX sensitivity. To do this, they made two types of channels: one that is completely snake (tsNAv1.4) and one that is a human (NAv1.4) and snake (tsNAv1.4) chimera. They used the tsNAv1.4 sequence from the Benton population to begin with and then point mutated individual amino acids to make the protein constructs that matched the mutations in the other snake populations.
Figure 1a: Figure 1a is a chart comparing the tsNAv1.4 sequences of 4 snake populations that all have different levels of resistance to TTX, but all have an aromatic amino acid in domain I, which under normal conditions would indicate a high affinity for TTX. TTX resistance is reported in mass adjusted mouse units (MAMU), which is measured by calculating the amount of mass adjusted TTX that caused a 50% decrease in the sprint speed in the snakes. The Bear Lake population is not resistant to TTX and therefore serves as a negative control in this figure. Willow Creek had the strongest resistance with 730 MAMU, followed by Benton with 34.1 MAMU, and Warrenton with 15.2 MAMU. This figure also shows the phylogenetic relationship of the snake populations based on mitochondria DNA analysis. It shows that the Willow Creek population diverged first with TTX resistance, and the Warrenton and Benton populations evolved TTX resistance separately after they diverged from the Bear Lake population.
Figure 1b: Figure 1b displays the amino acid sequence of TTX binding site on domain IV for each of the snake populations. Dots indicate amino acids that are conserved from the Bear Lake population which is TTX-sensitive. According to the text, there are no sequence differences between the four populations in coding regions of the pore helix, selectivity filter and B-strand of domains I, II, and III. Domain IV of the Bear Lake population matches the human and rat sequences, which are TTX sensitive, and therefore validates it as a negative control. All TTX-resistant snakes have a valine substitution of isoleucine at position 1561 in the pore helix, which appears to be an important region for TTX binding. The Benton population has an alanine substitution for glycine at position 1566 in the B-strand. The Willow Creek population has an arginine substitution for aspartic acid and valine substitution for glycine at position 1568 and 1569 in the B-strand, and a leucine substitution for isoleucine at position 1556 in the pore helix. This figure is very effective at showing how the amino acid sequences differ between the populations and correlating those changes in amino acid sequence to differences in TTX resistance and phylogentic relationships. However, a positive control would have been a helpful comparison. Perhaps, the amino acid sequence of a snake with a non-aromatic amino acid in domain I, that is known to be TTX resistant, could have been added for comparison so that we could determine if these new mutations are separate from that known mutation or if they are part of the whole mutated phenotype. It also could be useful to see the rest of the sodium ion channel amino acid sequence compared between the four populations or to know how closely the amino acids were conserved in order to ensure that it is this mutation that is causing the TTX-resistant phenotype and not another mutation.
Figure 2a: Figure 2a shows the comparison of TTX sensitivity, measured by the ratio of unblocked channels to total current over various TTX concentrations, of entirely snake channels vs. human/snake chimeric channels. The orange line of the graph shows the Benton population and the purple line shows the Bear Lake population. Within those two populations, the open squares are the entirely snake channels and the filled squares are the human/snake chimeric channels. The graph shows that there is no significant difference in TTX sensitivity between the snake only and human/snake channels, because the 2 lines are well within each others 95% confidence limits. There is a significant difference between TTX sensitivity of the Bear Lake and the Benton populations, which corresponds to the previous figure where the Bear Lake population had a lower MAMU value than the Benton population. This demonstrates that it is the variations in domain IV of the snake sequence that cause TTX resistance because there was no difference in TTX sensitivity when the other portions of the channel were composed of TTX-sensitive human channel. It might have been helpful to see the TTX sensitivity for a normal Xenopus oocyte without the TTX-resistant sodium ion channel expressed, to serve as a base line so that we could determine what the normal level of TTX sensitivity is. Similarly, it might also have been helpful to see a Xenopus oocyte with an entirely human channel expressed, so that we could directly compare the effects of adding the snake portions of the chimeric protein.
Figure 2b: Figure 2b is comparing the TTX sensitivity, again measured by the ratio of unblocked channels to total current over various TTX concentrations, of the chimeric channels with mutations that correspond to the mutations found in the four different snake populations. The most sensitive is the Bear Lake population, followed by the Warrenton, Benton, and then Willow Creek populations, which corresponds to the data presented in Figure 1. From this figure it appears that more mutations in the gene correspond with greater TTX resistance, as seen in the Willow Creek population, which has the greatest number of mutations and the highest resistance. Again, normal Xenopus oocytes would have been a good negative control and given a good base line value for TTX sensitivity.
Figure 2c: Figure 2c is showing the effect of a single amino acid substitution at position 1561 from isoleucine to valine in the Willow Creek population only. The graph shows that when the isoleucine is present the TTX resistance decreases, but not substantially. This shows that even though this mutation is present in all of the TTX-resistant populations it is not the main cause of the resistance, and other mutations are also involved. This figure would have been more informative if the substitution of valine for isoleucine in all snake populations was compared, because this would help determine if the mutations are cumulative and all contribute to resistance in different ways or if one mutation augments the other mutations.
Figure 2d: Figure 2d shows the current recordings through the chimeric channels in the Xenopus oocytes. The gray line is the base line negative control without any current. The black line is the positive control, which is the chimeric channels operating under normal conditions prior to the addition of TTX. The blue line is the current through Bear Lake channel after the addition of 100 nM TTX. This line is very similar to the negative control and shows that the TTX blocked the channels and prevented the current from passing through. The red line is the current through the Willow Creek channel after the addition of 100 nM TTX. This line is very similar to the positive control and shows that the TTX did not block the channels and the current passed through normally. This graph is very effective, because it displayed positive and negative controls, which makes it very convincing that the Bear Lake and Willow Creek channels are different in their ability to bind TTX and thus transmit a charge.
Table 1: Table 1 shows the Kd values for the various combinations of chimeric channels. The Kd value is the TTX concentration that blocked 50% of the channel. It shows that there is no significant difference between the entirely snake channel and the snake/human chimera in either the Bear Lake or Willow Creek populations, which corresponds to Figure 2a. It also shows that Willow Creek has higher resistance then Warrenton, followed by Benton and Bear Lake, which corresponds with Figure 2b. Finally, it shows that The Willow Creek chimera with the isoleucine (V1561) has about half the resistance as the Willow Creek chimera with the valine, which corresponds to Figure 2c. This table is a very effective way of demonstrating this information and is a nice supplement to the graphs in Figure 2.
Figure 3: Figure 3 compares the Kd of the chimeric channels to the Kd of the skeletal muscle. The Kd for the skeletal muscle was calculated by looking at the rates of rise in the action potential of the skeletal muscle fibers over varying TTX concentrations. The graph shows that there is a significant relationship, with a p value < 0.0001, between the Kd value of the chimeric channel and the Kd value of the skeletal muscle. This indicates that TTX resistance in the channel is a good indicator of TTX resistance in the whole animal. This is an important figure because it shows that the data can be generalized and the TTX resistance of the whole animal is probably a result of the TTX resistance of the sodium ion channel.
Critique: Overall this paper was well written, and the data given support the author's claims. The fact that they used functional tests to determine TTX resistance was very convincing, and comparing those functional differences to genetic differences with good molecular explanations of why the populations are functionally different was very helpful. They also did a nice job of showing that the TTX resistance seen in the expressed sodium ion channels could be generalized to whole animal TTX resistance. In most of their experiments they had fairly good positive and negative controls that allowed for comparison. However, in a few circumstances, like with the Xenopus oocyte expression experiments, a baseline value would have been helpful in order to compare TTX sensitivity between the four populations. Also, it would have been nice to compare the TTX resistance and the sequence of the mutated gene to other known TTX resistant animals, that could serve as a positive control.
There are a few limitations to their experiments that should be mentioned. All of their work was done on the cDNA sequences of the sodium ion gene, and therefore can not be generalized to the genomic DNA. It could be that the sodium ion channel is TTX resistant as a result of some change in the non-coding region of the DNA, which was not tested in this experiment. It is possible that instead of just a change in the structure of the sodium ion channel, TTX resistance occurs because of a change in the expression of the sodium ion channel gene which may produce less sodium ion channels for the TTX to bind to. Similarly, there could be a difference in the amount of mRNA that is produced, which could affect TTX binding. These possibilities were not accounted for in this paper. However, in general from their experiments it is clear that changes in the structure of the sodium ion channel do affect the TTX sensitivity of the populations, but whether this is the only explanation for the TTX resistance is left to be determined.
Future Research: There are lots of areas of future research that could expand upon the work done here. To begin, it would be nice to know if the TTX resistant phenotype is just a result of changes in the coding regions of the sodium ion channel gene or if it could also be the result of changes in gene regulation or expression. One way to test this would be to look at the expression level of the mRNA transcript of the different TTX resistant populations compared to a TTX sensitive control. In order to compare relative amounts of mRNA, a loading control that shows the RNA expression of some other regularly expressed gene like myosin would need to be included. Correspondingly, the non-coding regions of gene could be sequenced and compared to determine if TTX resistance is a result of mutations in promoters or enhancers. This research research might provide insights as to why TTX resistance varies within a population of snakes. If each snake population has the same mutation that changes the structure of the sodium ion channel then there must be some other change, such as the regulation or expression of the gene, which would cause slight variations within the population.
Additional research could also be done on the molecular mechanism of the TTX resistance. The paper speculates on how the amino acid changes might effect the TTX binding, but this has not been thoroughly tested. One way to test this would be to mutate specific amino acids and perform functional tests on the protein, as was done in this experiment, but with more attention to how the specific mutations change the TTX binding capabilities.
Finally, future research might also be conducted on whether these mutations, and therefore TTX resistance, could be translated to other species of animals, such as mice or humans. One way to test this would be to use the mouse or human sodium ion channel genes, point mutate the corresponding amio acids, and functionally test the TTX resistance of the channel. This could lead to valuable information about how the sodium ion channel works in humans, and potentially lead to conclusions about how TTX poisoning in humans could be remedied.
Geffeney, Shana L, Esther Fujimoto, Edmund D. Brodie III, Edmund D. Brodie Jr, and Peter C. Ruben. 2005. Evolutionary diversification of TTX-resistant sodium channels in a predator-prey interaction. Nature 434: 759-763.
Smith, Tim. Ion Channels in Biological Membranes. http://www.chemsoc.org/exemplarchem/entries/2002/Tim_Smith/about/. April 2005.
© Copyright 2005 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: krheiner@davidson.edu