*This web page was created as an assignment for an undergraduate course at Davidson College*
HFE Orthologs
Introduction
Orthologs are genes from multiple species that can be traced to a common ancestor. These genes usually serve similar functions in their respective organisms. Scientists can analyze their nucleotide or amino acid sequences to identify portions of the gene or protein that are highly conserved. Overall, the more highly conserved a gene/protein, the more recently the two species diverged. Specific portions that are highly conserved within a gene or protein sequence often correspond to important sites (i.e. - binding sites) in the final protein or are important to the final configuration of the protein. Analyzing ortholog sequences can help further understanding of the relationship between structure and function in proteins.
HFE
Since the role of HFE revolves around iron absorption from the blood, this protein is present in a narrower range of species than some. Also, HFE is a relatively recently characterized protein, and thus, its function is not completely understood. Currently, scientists have only found and characterized mammalian HFE sequences (Lambert Bioinformatics Group at Chatham).
I found human HFE orthologs from five other mammalian species: Canis familiaris (dog), Mus musculus (mouse), Rattus norvegicus (rat), Pan troglodytes (monkey), and Bos taurus (cow). Overall, the sequences were fairly well conserved (Figure 1).
Canis familiaris
67% similar to Homo sapiens
![]()
Mus musculus
61% similar to Homo sapiens
![]()
Rattus norvegicus
63% similar to Homo sapiens
![]()
Pan troglodytes
100% similar to Homo sapiens (for portion of sequence available)
![]()
Bos taurus
62% similar to Homo sapiens
![]()
Figure 1. Cartoon comparison of the amino acid sequences of humans and each of the orthologs. Red strips represent missing amino acids that would have to be "added" to make the sequences align. Black lines represent portions of the sequences on each end that do not correspond. The Pan troglodytes amino acid sequence is only partially known. Overall, the sequences are very similar. Blast 2 generated the images and percentages.
Conserved Sequences
As a reminder, HFE consists of three alpha domains, a transmembrane domain, and a cytoplasmic domain. The alpha-3 domain also interacts with beta2M (Figure 2). Structure is important to remember since amino acids that affect structure significantly are more likely to be highly conserved among species.
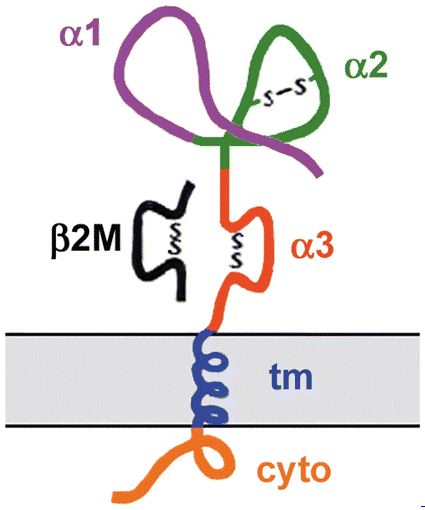
Figure 2. A cartoon depiction of the HFE protein. Image from Fleming and Sly, 2002. Permission pending.
Studies on the function and structure of HFE have found several mutations that cause or potentially cause disfunction of the protein (Figure 3). Mutations that cause the protein to cease functioning properly presumably cause important changes in the structure of the protein. Thus, these amino acids are likely candidates to be highly conserved.
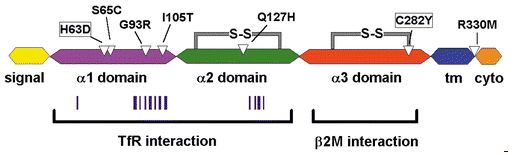
Figure 3. A cartoon depiction of the six exons of the HFE protein. White triangles indicate amino acids at which known missense mutations occur. Vertical bars indicate positions of amino acids likely to participate in HFE-TfR interaction. In general, the alpha-1 and alpha-2 domains interact with TfR and the alpha-3 domain interacts with beta2M. "tm" represents "transmembrane region", and "cyto" represents "cytoplasmic region." Image from Fleming and Sly, 2002. Permission pending.
C282Y mutation
In this mutation, tyrosine replaces the cysteine at amino acid position 282. This mutation is the most common cause of hereditary hemochromatosis (Zhang et al., 2003). The change interferes with intracellular transport of HFE, restricts the ability of the alpha-3 domain to interact with beta2M, and prevents the formation of a disulfide bond with C225 (Lebron et al., 1998). One explanation that exists for the relatively common occurrence of this mutation is that it could have been helpful for populations with limited dietary iron intake to absorb extra iron (Fleming and Sly, 2002). C282 is conserved in all six species, and C225 is conserved in all but the monkey (Figure 4; highlighted in aqua). Thus, these amino acids are likely important for structure, and mutations in other species probably cause similar problems.
H63D mutation
In this mutation, aspartic acid replaces histidine at amino acid position 63. Although this mutation causes an increased chance of developing hereditary hemochromatosis, it is present in approximately 15-40% of the Caucasian population. Consequently, the change is too common to cause major malfunction of the protein. In the wild type, H63 forms a salt bridge in the alpha-2 domain that binds TfR (Fleming and Sly, 2002). H63 is conserved in all six orthologs, which provides further evidence for the importance of the salt bridge to HFE structure and function (Figure 4; highlighted in yellow).
Other mutations
Researchers have found other amino acid mutations in patients with hereditary hemochromatosis although they have not determined the causal relationship. These mutations are S65C, G93R, I105T, Q127H, and R330M. S65C is the most common mutation after C282Y and H63D. G93R and I105T occur in the area of the alpha-1 domain responsible for TfR binding; these two mutations likely alter the structure and hinder HFE's ability to bind TfR (Fleming and Sly, 2002). S65, G93, and I105 are conserved in all six species while Q127 is only conserved in humans, dogs, and monkeys, and R330 is conserved in all but cows (Figure 4; highlighted in green). Further research will elucidate the specific role each of these proteins plays in HFE structure and function.
Transmembrane region
Amino acids 307 through 329 make up the transmembrane region of HFE in humans (Ensembl). HFE's position as a transmembrane protein likely is a crucial factor in its ability to function. Located elsewhere in the cell, HFE is probably useless. Thus, this sequence is particularly important, and it is partially conserved across all six orthologs (Figure 4; highlighted in pink). This conservation may be sufficient to retain function.
Histidine/tyrosine cluster
Four histidines (H109, H111, H116, H145) cluster with Y140 in the alpha-1 domain. This cluster resembles iron binding sites in other proteins although researchers have not proven it performs this function in HFE (Lebron et al., 1998). Y140, H16, and H145 are conserved across all six species; H109 is conserved in human, dog, and cow, and H111 is conserved in all but cow and monkey (Figure 4; highlighted in red). Presumably then these five amino acids are also important to HFE structure and function.
P188
In all other class 1 MHC proteins (the broader group of which HFE is a member) with a proline at this position, a kink is present in the structure. This kink occurs in the groove for peptide binding, and the absence of the kink in HFE possibly explains why HFE's groove cannot bind peptides (Lebron et al., 1998). This amino acid is conserved in all six species (Figure 4; highlighted in dark turquoise), which indicates that it does play an important role in HFE structure and function.
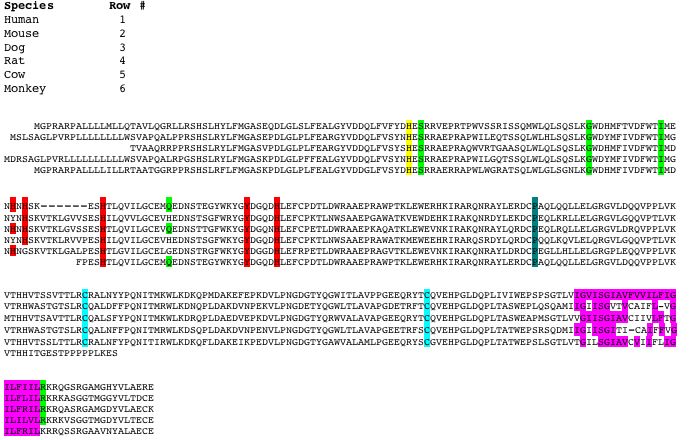
Figure 4. Orthologs of HFE from six species: Homo sapiens (human), Mus musculus (mouse), Canis familiaris (dog), Rattus norvegicus (rat), Bos taurus (cow), and Pan troglodytes (monkey). Amino acid sequences are aligned to best show conservation among species. Aqua highlights C282 and C225. Yellow highlights H63. Green highlights S65C, G93R, I105T, Q127H, R330M. Pink highlights the transmembrane region. Red highlights the histidine/tyrosine cluster. Dark turquoise highlights P188. Sequences provided by Ensembl.
References
Blast 2. http://www.ncbi.nlm.nih.gov/blast/bl2seq/wblast2.cgi. Accessed 6 March 2005.
Ensembl. http://www.ensembl.org. Accessed 6 March 2005.
Fleming RE, Sly WS. 2002. Mechanisms of iron accumulation in hereditary hemochromatosis. Annual Review of Physiology 64: 663-680.
Lambert Bioinformatics Group. 2005 February 21. Chatham College. http://www.chatham.edu/undergraduate/bio/lambert/research.htm#Hfe. Accessed 7 March 2005.
Lebron JA, et al. 1998. Crystal Structure of the Hemochromatosis Protein HFE and Characterization of Its Interaction with Transferrin Receptor. Cell 93: 111-123.
Zhang A, et al. 2003. Mechanisms of HFE-induced regulation of iron homeostasis: Insights from the W81A HFE mutation. PNAS 100 (16): 9500-95005.