*This website was produced as an assignment for an undergraduate course at Davidson College.*
My Favorite Protein: NGF
The Structure-Function Relationship
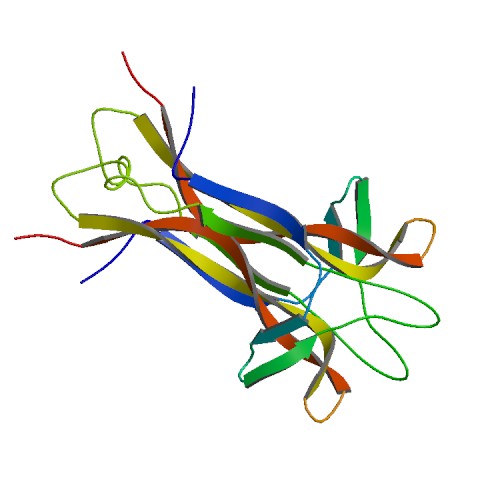
Figure 1. Basic structure of NGF. Image from Protein Data Bank.
Introduction:
Neurotrophins, including Nerve Growth Factor (NGF), brain-derived neurotrophic factor (BDNF), neurotrophin-3 (NT-3), neurotrophin-4/5 (NT-4/5), and neurotrophin-6, comprise a family that plays a critical role in the development and maintenance of the neuronal networks in the peripheral and central nervous systems. Due to the fundamental neutrophic functions, many researchers are considering their potential therapeutic qualities, particularly in the treatment of certain neurodegenerative diseases including Alzheimer’s disease and Parkinson’s disease (McDonald et al., 1991). However, an incomplete understanding of the protein-receptor complexes has impeded the utility of neurotrophins as clinical agents. Nerve Growth Factor is the most studied and best understood member of the neurotrophin family, and as an individual interested in the neuroscience field, I chose NGF to be my favorite protein.
Structure:
An understanding of the structure-function relationship of a protein is dependent upon basic knowledge of the structure. Specifically, NGF functions by binding to cell surface receptors. Thus, complete identification of not only the structure of the protein, but also of the NGF binding sites and receptors are required. Research has recently incorporated chemical modification, site-directed mutagenesis, and X-ray crystal structuring to gain insight into the structure of NGF (Bradshaw et al., 1994).
Nerve Growth Factor is a homodimeric molecule, consisting of two structurally similar subunits. This dimer formation is thought to be essential for NGF’s biological functioning (Bradshaw et al., 1994). The two protomers of NGF, assembled around a twofold axis, are comprised of 118 amino acids each and run parallel to each other. The protein contains antiparallel β-pleated strands, and the residues from these β-strands play a fundamental role in the stabilization of the dimer. One end of the molecule contains three hairpin loop structures, loops that may be responsible for the functional differences between the proteins of the neurotrophin family (McDonald et al., 1991). The other end of the molecule displays a cystein-knot motif, stabilizing the fold and locking the molecule into its specific conformation. Three disulfide bridges, two of which form a ring with the protein backbone while the other penetrates the ring, form the cysteine-knot motif (Wiesmann et al., 1999).
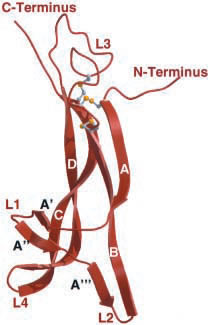
Figure 2. NGF monomer depicting termini, looping regions (L1-L4), and cysteine-knot motif (grey and yellow ball-and-stick).
Image from Wiesmann et al., 2001. Permission pending.
View Jmol structure of Nerve Growth Factor.
Function:
As mentioned above, neurotrophins function by binding to two different classes of cell surface receptors, p75 and Trk receptors. Through the mediation of these two classes of cell surface receptors, NGF and the other neutrophins can control cell survival, cell differentiation, sensory neuron apoptosis, and growth cessation (Wiesmann et al., 1999). The first class of cell surface receptors, p75, are low affinity receptors belonging to the family of the tumor necrosis factor receptors. This receptor is not specific to NGF but is activated by all neurotrophins. When NGF binds and activates this receptor, apoptosis is initiated in the absence of TrkA. However, activation promotes cell survival if TrkA is expressed on the surface of the cell. The second receptor for NGF, TrkA, displays a high level of affinity and specificity. After NGF binds, the receptor dimerizes, autophosphorylates, and triggers a cascade of signaling.
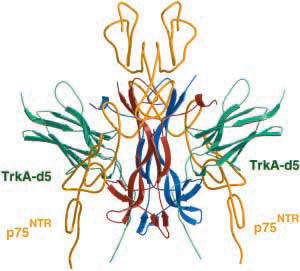
Figure 3. Complex between NGF, TrkA receptor (green), and p75NTR receptor (yellow).
Image from Weismann et al., 2001. Permission pending.
Structure & Function:
NGF has two binding epitopes for the p75 receptor. One epitope involves positively charged amino acids in L1 and L4. This epitope can be divided into two patches. The first patch is a "knob-in-hole configuration," and the second patch consists of a salt bridge between NGF and p75 (He et al., 2004). The second epitope, consisting of hydrophilic amino acids, is found in L3 and the C terminus (Wiesmann et al., 1999). The complex between NGF and p75 is unique because it binds in a way that leaves NGF with an open binding site. Perhaps, this mechanism makes it possible for NGF to participate in another binding complex, such as with the TrkA receptor (He et al., 2004).
The TrkA receptor is comprised of five domains. It has been found that the region proximal to the membrane, the TrkA-d5 domain, is "necessary and sufficient" to bind to NGF (Wiesmann et al., 1999). The complex between NGF and the TrkA receptor has two interfaces. The first involves the central β-sheet region of NGF. The second interface is found at the terminal regions of NGF which form a helical conformation when binding. It is hypothesized that the first interface is conserved between all neurotrophins, but the second interface is specific for the NGF-TrkA interaction (Wiesmann et al., 1999).
In the proposed structure of the binding complex between NGF and its receptors, NGF has minor contact with TrkA-d5. This orientation then places the cysteine knot, N terminus, and C terminus of NGF near the top of the molecule. In turn, the areas responsible for binding to p75 receptors are partially exposed in this orentation. This finding is consistent with the notion that NGF can bind to both TrkA and p75 receptors simultaneously (Wiesmann et al., 1999). This trimolecular binding complex makes it possible for NGF to control various activities, by binding to p75, TrkA, or both receptors.
References:
Bradshaw R, Murray-Rust J, Ibanez C, McDonald, Lapatto R, Blundell T. 1994. Nerve growth factor: structure/function relationships. Protein Science 3: 1901-1913.
He X, Garcia K. 2004. Structure of nerve growth factor complexed with the shared neurotrophin receptor p75. Science 304: 870-875.
McDonald N, Lapatto R, Murray-Rust J, Gunning J, Wlodawer A, Blundell T. 1991. New protein fold revealed by a 2.3-Å resolution crystal structure of nerve growth factor. Nature 354: 411-414.
Wiesmann C, Ultsch M, Bass S, de Vos A. 1999. Crystal structure of nerve growth factor in complex with the ligand-binding domain of the TrkA receptor. Nature 401: 184-188.
Wiesmann C, de Vos A. 2001. Nerve growth factor: structure and function. Cellular and Moleuclar Life Sciences 58: 748-759.
Davidson College Molecular Biology Homepage
Please send any questions, comments, or suggestions to moheinzelmann@davidson.edu.