
*This website was produced as an assignment for an undergratuate course at Davidson College.*
Sec61/Y - The Evolutionary Story of a Protein Essential to All Forms of Life
As explained here, Sec61p is an essential protein in the S. cerevisiae translocon complex. In yeast, the translocon complex allows for newly translated polypepties to be transported across or embedded into the Endoplasmic Reticulum (ER) membrane. Translocation is an essential element of the secretory pathway, which ensures the proper localization and membrane orientation of proteins. Proper localization of proteins is necessary for any functional cell, and thus it is not surprising that an ortholog of Sec61 is found in every living organism that has had its genome sequenced (Cao and Saier, 2002). This set of orthologs is referred to as the Sec61/Y family (upon discovery, this protein was named Sec61 in eukaryotes and SecY in bacteria), and the sequences of this family of proteins are highly conserved across all organisms (Cao and Saier, 2002). Once again, this is not surprising, due both to the necessity of Sec61/Y’s role in the cell, as well as the structural components of Sec61/Y that allow it to accomplish its function. These include surprisingly hydrophillic transmembrane regions that, together, form an aqueous pore used for polypeptide transportation (Wilkinson, et al., 1996), as well as cytosolic loops designed to bind to ribosomal components that are also necessary for the translocation process (Cheng et al., 2005). More information on these structures can be found here.
Where is it found, and what does it do?
In eukaryotes, Sec61/Y family proteins are found in the ER membrane, and serve to transport across and embed proteins within this membrane (Stephenson, 2005). In bacteria, Sec61/Y family proteins are located in the cytoplasmic membrane and serve to transport proteins across and embed proteins within this membrane (Stephenson, 2005). Sec61/Y family proteins are also found in archaea. Though relatively little is known about the specific details of the translocaiton process in archaea, the Sec61/Y protein in archaea still serves the purpose of transporting soluble proteins across a lipid membrane (Bolhuis, 2004). Additionally, Sec61/Y proteins are always found as part of a trimeric complex. In eukaryotes and archaea, this complex is referred to as the Sec61αβγ complex, and in prokaryotes it is referred to as the SecYEG complex (Cao and Saier, 2002).
Which segments are most conserved?
Generally speaking, the amino acid sequence of Sec61/Y family proteins is highly conserved across all organisms. For example, the Sec61/Y protein found in S. cerevisiae (Sec61p) is 56% identical to the human ortholog (Secalpha), 56% identical to the Mouse ortholog (Sec61alpha), 57% identical to the Drosophila ortholog (GH1329) and 54% identical to the Arabidopsis ortholog (Sec61) (Data obtained from Protein Blast results using sequences obtained from Uniprot.org). All of these proteins have the same topologies, containing 10 transmembrane segments (Cao and Saier, 2002).
There are, however, some segments that are more highly conserved than others. Cytoplasmic loops located between transmembrane segments 2-3, 4-5, 6-7 and 8-9 are well conserved, whereas the extracytoplasmic loops located between transmembrane segments 1-2, 3-4, 5-6, 7-8 and 9-10 are less well conserved (Cao and Saier, 2002). This makes sense, as evolutionary pressures would ensure that functionally significant loops were conserved. Because the ribosome interacts with the cytoplasmic loops of Sec61/Y proteins in order to begin translocation, the cytoplasmic loops are more likely to be conserved. It is also possible that the lack of conservation among the extracytoplasmic loops is due to the need for those loops to have specific “protein-protein recognition” abilities (Cao and Saier, 2002). Because extracytoplasmic proteins will vary among species, it may not be advantageous for the structures of these extracytoplasmic loops to be universally conserved (Cao and Saier, 2002).
The most conserved region of the protein is found in the cytoplasmic loop between transmembrane segments 4-5, and within the 5th transmembrane segment (Cao and Saier, 2002). "The consensus sequence for this region is:
G (L I V) G N G (L I V) S (L I V)3 F (L I V A) G (L I V)2 (S A T)2 (L I V) P.
(Alternative residues at a single position are indicated in parentheses)” (Cao and Saier, 2002).
Once again, this high degree of conservation makes sense given the function of this segment of Sec61/Y proteins. As described here, transmembrane segment 5 is one of the surprisingly hydrophillic regions that is necessary for Sec61/Y to form the aqeous, translocation pore (Johnson, van Waes, 1999).
Protein Sequences and Clustal W Alignment
When analyzing orthologs, it is useful to compare the amino acid sequences of a conserved protein between many different organisms. I obtained amino acid sequences for Sec61/Y family proteins using Uniprot.org, and aligned these sequences using Clustal W (http://www.ebi.ac.uk/Tools/clustalw2/index.html).
The organisms that correspond to each protein code are listed below:
Saccaromyces cerevisiae |
P32915-1 |
Homo sapiens |
P61619-1 |
Mus musculus |
P61620-1 |
Caenorhabditis elegans |
O18239-1 |
Drosophila grimshawi |
B4JQ75-1 |
Arabidopsis thaliana |
Q9ZV90-1 |
Escherichia coli (strain K12) |
P0AGA2-1 |
Fig 1. This shows an alignment of amino acid sequences for Sec61/Y family proteins found in select eukaryotes. Note the highly conserved region located roughly between A.A.’s 165-200. This segment aligns with the predicted 5th transmembrane segment (Uniprot.org amino acid sequence of Sec61), which is noted to be the most highly conserved segment of the Sec61/Y family proteins (Cao and Saier, 2002).
Fig 2. This shows an alignment of amino acid sequences for Sec61/Y proteins found in eukaryote S. Cerevisiae versus prokaryote E. coli. Contrary to much of the reading I have done, these sequences do not appear particularly similar. It is possible that, upon further examination, though the exact amino acids do not match, their properties are similar. One statement that leads me to this conclusion is the analysis of cytosolic regions 6 and 8 by Cheng et al. in 2005, which notes that the lengths of these loops, more so than the exact sequences, are the conserved elements.
Phylogenetic Tree
In 2002, Dr.’s Thien Cao and Milton Saier created a phylogenetic tree for the Sec61/Y family proteins.
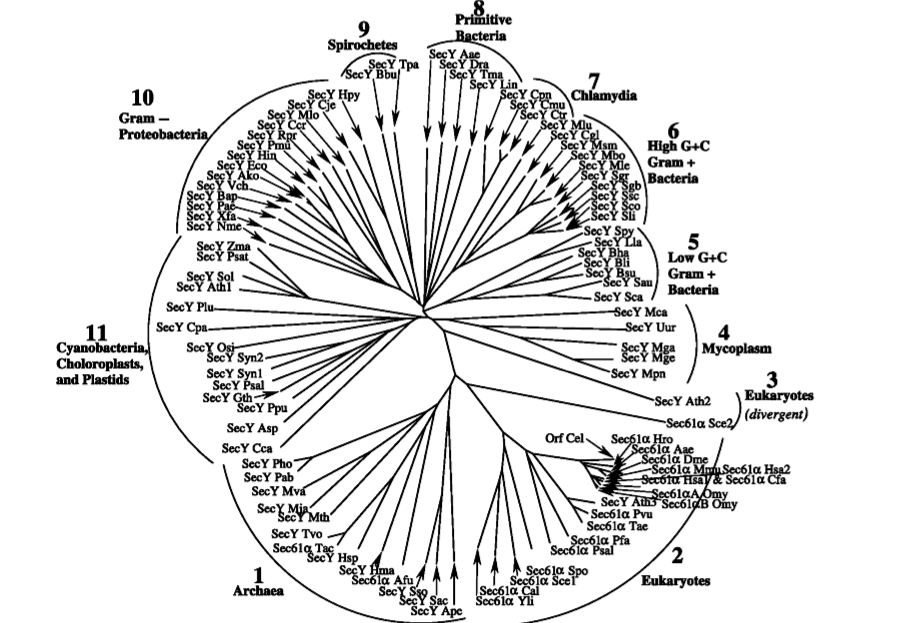
Fig. 3. This phylogenetic tree shows all the sequenced members of the Sec61/Y protein family as of 2002. Note the low levels of protein diversity found in eukaryotic and archaeal species, in contrast with the high levels of protein diversity found in prokaryotes. Image retrieved from Cao and Saier, 2003, Permission Pending. Link to Article
Where is there the most variation? Why? What does this tell us about evolution?
As demonstrated by the phylogenetic tree, the eukaryotic and archaeal Sec61/Y orthologs make up only a fraction of the total orthologs found in nature. Most of the variety in the Sec61/Y family lies in bacterial organisms. This pattern in diversity indicated that “the bacterial domain is more diverse at the molecular level than either the archaeal or the eukaryotic domain” (Cao and Saier, 2002). This is “consistent with the suggestion that bacteria were the evolutionary precursors of the archaea and eukaryotes” (Cao and Saier, 2002).
In addition to the patterns of overall diversity, it is important to note that a Sec61/Y family proteins only occurs once within the genomes of most organisms (Cao and Saier, 2002). These minimal levels of Sec paralogs indicate that there is a strong evolutionary pressure to conserve a specific Sec gene, which minimizes "gene duplication events leading to the establishment [of these] paralogs" (Cao and Saier, 2002).
Conclusion
The presence of a protein from the Sec61/Y family is clearly essential for any functional cell. This necessity has led to interesting evolutionary patterns; conserved regions demonstrate which segments of the protein are essential to function, and the lack of paralogues within species demonstrates the evolutionary pressure to maintain one, conserved Sec61/Y protein within the genome.
Works Cited
Cao, T., Saier M. The General Protein Secretory Pathway: Phylogenetic Analysis Leading to Evolutionary Conclusions. Biochimica et Biophysica Acta 2003; 1609: 115-125 Link to Article
Cheng Z., Jang Y., Mandon E., Gilmore R. Identification of Cytoplasmic Residues of Sec61p Involved in Ribosome Binding and Cotranslational Translocation. Journal of Cell Biology 2005; 168.1: 67-77. Link to Article
Johnson A., van Waes M. The Translocon: A Dynamic Gateway at the ER Membrane. Annu. Rev. Cell Dev. Bio. 1999; 15: 799-842. Link to Article
Stephenson, K. Sec-dependent Protein Translocation Across Biological Membranes: Evolutionary Conservation of an Essential Protein Transport Pathway (Review). Molecular Membrane Biology 2005; 22(1-2): 17-28. Link to Article
Wilkinson B., Critchley A., Stirling C. Determination of the Transmembrane Topology of Yeast Sec61p, an Essential Component of the Endoplasmic Reticulum Translocation Complex. The Journal of Biology Chemistry 1996; 271.41: 25590-25597. Link to Article
Please direct questions or comments to alvalauriorton@davidson.edu