
*This website was produced as an assignment for an undergratuate course at Davidson College.*
The Structure and Function of Sec61p
An Essential Protein in Saccharomyces cerevisiae Translocons

Fig 1. Structure of Sec61p
Image retrieved from ModBase
Why is Sec61p needed by the cell?
Translation:
In eukaryotes, all ribosomes are found in the cytoplasm. Thus, translation of all proteins begins in the cytoplasm, even though as many as half of the proteins made in a cell will ultimately be localized somewhere other than the cytoplasm (e.g. in the plasma membrane or the nucleus), and will have to cross as many as three membranes to get to their final destination (Lodish, et al., 2008). As a result, there is a mechanism for directing nascent proteins (proteins that are still being formed by a ribosome, and have not yet folded into their native configuration) to their proper locations, as well as a mechanism for transporting proteins across membranes.
Signal Sequences:
Proteins that must be transported out of the cytoplasm contain signal sequences in their amino acid sequences. During translation, signal recognition particles (SRPs) bind to these newly translated signal sequences, bring translation to a temporary halt, and direct the proteins to the target organelles indicated by their signal sequences. One common destination for nascent proteins is the endoplasmic reticulum (ER). Aside from serving as the final location for some proteins, the ER is also the first stop on what is known as the secretory pathway. On this pathway, proteins are transported to the ER and through the golgi apparatus, undergoing further processing before they are transported to their final destinations. Proteins targeted to the ER contain a signal sequence at their N-terminus – the first part of the protein that is translated (Lodish, et al., 2008). Surprisingly, these signal sequences have little in common aside from containing “one or more positively charged amino acids adjacent to a continuous stretch of 6-12 hydrophobic residues” (Lodish, et al., 2008).
Translocons:
Many of the proteins targeted to the ER are luminal or transmembrane proteins, which means they contain hydrophilic regions in their structures. Like all membranes, the ER membrane is composed of a phospholipid bilayer that is non-permeable to hydrophillic structures. So – how are hydrophillic proteins transported from the cytoplasm to the ER? Even more puzzling – how are transmembrane proteins, which contain alternating hydrophillic and hydrophobic regions, properly placed within membranes such that they contain all their necessary domains? Translocon complexes, one of which is called the Sec61 complex, facilitate this transportation and positioning.
The Sec61 Complex and Sec61p
Sec61 Complex:
In Saccharomyces cerevisiae, the Sec61 complex comprises Sec61p, Sss1p and Sbh1p, and is located on the ER membrane (Wilkinson, et al., 1996). This complex “interacts with a second membrane protein complex (comprising Sec62p, Sec63p, Sec71p, and Sec72p” to facilitate “efficient post-translational translocation” (Wilkinson, et al., 1996). Of all the proteins in these complexes, the protein most specifically targeted by nascent proteins and thought to contain the pore through which these proteins travel is Sec61p (Johnson, van Waes, 1999).
Sec61p
Sec61p is an integral membrane protein located on the surface of the ER. It is made up of one polypeptide chain containing 480 amino acids, and has 10 transmembrane regions (Uniprot.com, 2010). Within each Sec61 complex, an estimated 3-4 Sec61p heterotrimers assemble in a ring formation to create a central, aqueous pore (Uniprot.com, 2010). When the Sec complex is not bound to a ribosome, a protein called BiP blocks the aqueous pore formed by the complex, maintaining the membrane barrier of the ER. When the nascent polypeptide chain comes into contact with BiP, it changes BiP’s conformation, causing it to remove itself from the pore of the translocon (Johnson, 1999). This allows the polypeptide chain to pass through the pore into the lumen of the ER.
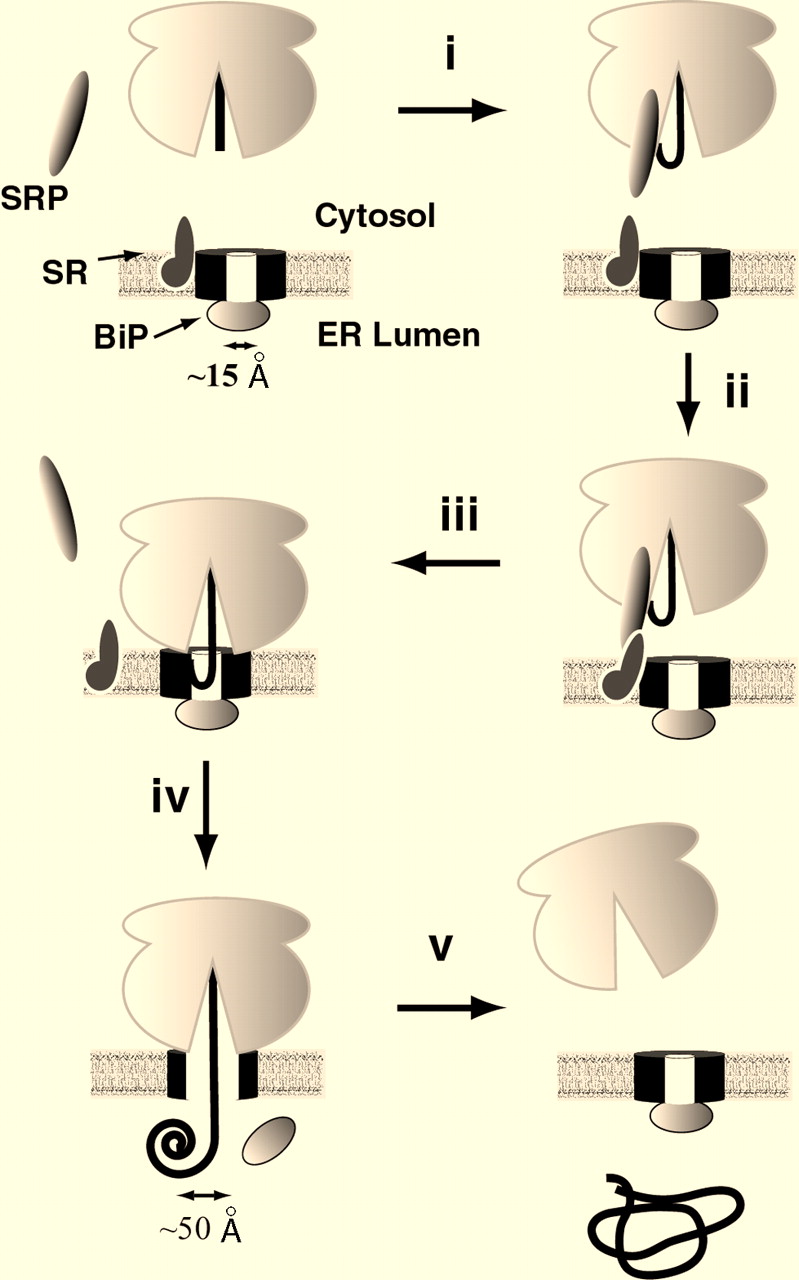
Fig 2. Depiction of ribosome binding to the Sec61 translocon complex. The polypeptide strand interacts with BiP and gains access to the ER lumen.
Image retrieved from Johnson and van Waes, 1999, with permission.
Structure Leads to Function
The unique structure of Sec61p enables the SRP-bound ribosome complex to target an available translocation site, and also allows for the formation of an aqueous pore in the ER membrane
Ribosome Binding:
The formation of the complete translocation complex, including the nascent polypeptide strand, the ribosome, SRP, the Sec61 complex and other ER membrane proteins, is a multistep process, involving the interaction of many molecules. One element of this process is the need for the ribosome to be able to identify translocation pores that are not currently “in use.” A 2005 study by Cheng et al. indicates that the cytoplasmic loops six (L6) and eight (L8) serve as a recognition site for the ribosome during translocation, arguing, “Cytosolic loops of Sec61p would be the optimal marker for an unoccupied translocon, as these segments will be occluded upon attachment of a ribosome to the translocation channel.” These loops are “highly exposed to the cytoplasmic surface of the Sec61 complex,” and point mutations in the coding regions for these loops “cause reductions in growth rates and defects in the translocation of nascent polypeptides that use the cotranslational translocation pathway” (Cheng et al., 2005). Additionally, “the lengths and, to a lesser extent, the sequences of L6 and L8 are well conserved between the archae and eukaryotic translocation channels,” indicating that the presence of these loops plays an important role in Sec61’s function (Cheng et al., 2005).
Formation of Aqueous Pores:
The presence of aqueous pores in the ER membrane was first detected by conductivity measurements, and later explored using water-sensitive fluorescent probes (Johnson, 1999). Hydopathic analysis of Sex61p indicates the presence of 10 transmembrane domains, but, more importantly, several of these domains have “surprisingly limited hydrophobicity” (Wilkinson, et al., 1996). For example, the fifth domain predicted to span the membrane contains “12 residues, in which only 11 are nopolar” (Wilkinson, et al., 1996). Using a C-terminal reporter approach, Wilkinson et al. provide evidence that the weakly hydrophobic domains of Sec61p, such as the fifth membrane spanning region, require “C-terminal sequences for their correct topogenesis.” Wilkinson et al. also propose that the fifth membrane spanning region is stabilized when it forms an alpha helix or beta sheet within the bilayer.
A 1999 report on the translocon by Johnson and van Waes elaborates on Wilkinson et al., concluded that the available data suggest that “the walls facing the aqueous pore are formed largely by the alpha-helices of Sec61alpha [the mammalian homologue to S. cerevisiae Sec61p].” Wilkinson et al. propose that membrane spanning regions 5, 6, 9 and 10 interact in order for Sec61p to function. Thus, it is possible that these regions with limited hydrophobicity form alpha helices that together form the aqueous pore earlier detected in the ER membrane. Additionally, as indicated by the stabilizing structures identified by Wilkinson et al., suggest that overall structure of Sec61p may serve as the means of stabilization for these transmembrane regions of limited hydrophobicity, and ultimately allows for the creation of aqueous pores.
Size of the Aqueous Pore:
The size of the aqueous pore formed by the Sec61 complex depends on whether the complex is bound to a ribosome or not (Johnson, van Waes, 1999). While unbound, the pore has a diameter between 9-15 angstroms, “large enough to pass a hydrated iodide ion, but too small to pass a hydrated ADP molecule” (Johnson, van Waes, 1999). When bound to a ribosome, the Sec61 complex shifts in formation such that the pore formed by the Sec61p heterotrimers is 40-60 angstroms in diameter (Johnson, van Waes, 1999). In this formation, the ribosome maintains the permeability barrier of the ER membrane, but the polypeptide it is creating is still able to pass through the translocon into the ER lumen (Johnson, van Waes, 1999). Though the pore undergoes a threefold increase in diameter, the diameter of the Sec61p complex only increases 10%, suggesting that protein movement within the complex, as well as the removal of BiP from the pore, produce the enlarged pore diameter (Jonson, van Waes, 1999).
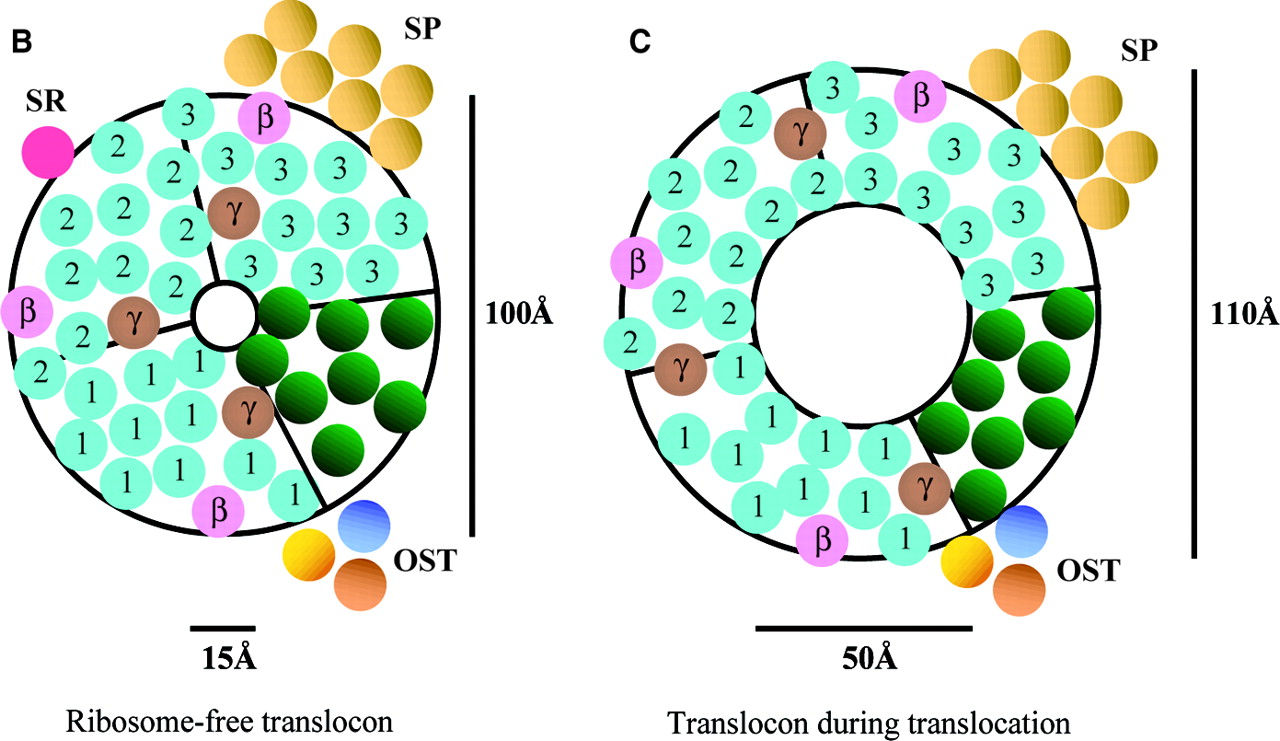
Fig 3. Depicts pore structure of the Sec61 translocon complex in mammals. Three Sec61alpha (homologue to Sec61p) heterotrimers are represented by blue numbers.
Left image depicts translocon that is unbound. Right image depicts the translocon bound to a ribosome.
Image retrieved from Johnson and van Waes, 1999, with permission.
Positioning of Transmembrane Proteins:
As mentioned above, transmembrane proteins have both hydrophillic and hydrophobic domains, which must be positioned appropriately within a membrane. When a ribosome is bound to a translocon and is translating an amino acid sequence, SRP, in conjunction with the Sec61 complex, recognizes signal anchor sequences in the growing polypeptide strand (Johnson, van Waes, 1999). These sequences indicate which regions of a growing protein must be embedded in a membrane. Photocross-linking studies followed the path of a transmembrane segment as it moved through a translocon. Initially the transmembrane segment was adjacent to Sec61alpha (the mammalian homologue of Sec61p), but as translation continued, the segment moved away from Sec61alpha, towards hydrophobic regions of the translocon complex (Johnson, van Waes, 1999). While the exact mechanism of this movement is unknown, it is likely that the aqueous pore formed by alpha helices of Sec61p is structured such that the alpha helices may shift in relation to the growing polypeptide strand in order to let transmembrane regions escape the pore and become embedded in the ER membrane.
In Conclusion...
Sec61p, and its role in the Sec61 translocon complex, is necessary for proper protein localization within the cell. It’s unique structure, with limited hydrophobicity and easily targeted cytosolic domains, lends to Sec61p’s ability to do this. Proteins are only fully functional if they are in the right orientation and location, and thus Sec61p is necessary for any functional cell.
Want More Information?
You can find the complete amino acid sequence and identification of transmembrane regions of Sec61p at Uniprot.org
Works Cited
Cheng Z., Jang Y., Mandon E., Gilmore R. Identification of Cytoplasmic Residues of Sec61p Involved in Ribosome Binding and Cotranslational Translocation. Journal of Cell Biology 2005; 168.1: 67-77.
Johnson A., van Waes M. The Translocon: A Dynamic Gateway at the ER Membrane. Annu. Rev. Cell Dev. Bio. 1999; 15:799-842.
Lodish H., Berk A., Kaiser C., Krier M., Scott M., Bretscher A., Ploegh H., Matsudaira P. Molecular Cell Biology. Sixth Edition. New York: W. H. Freeman and Company; 2008.
Uniprot. Protein Transport Protein Sec61 - Saccharomyces cerevisiae. 2010. Uniprot.org. February 15 2010. http://www.uniprot.org/uniprot/P32915
Wilkinson B., Critchley A., Stirling C. Determination of the Transmembrane Topology of Yeast Sec61p, an Essential Component of the Endoplasmic Reticulum Translocation Complex. The Journal of Biology Chemistry 1996; 271.41: 25590-25597.
Please direct questions or comments to alvalauriorton@davidson.edu