This web page was produced as an assignment for an undergraduate course at Davidson College
Song et al. Figure 2

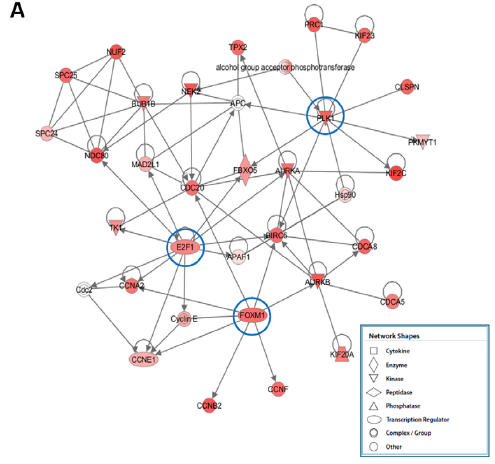
Figure 2. A) Diagram shows top-scoring network of genes enhanced in the PAH condition, including multiple important transcription regulators. Network score determined via network analysis in Ingenuity Pathway Analysis. Red nodes show up-regulation.
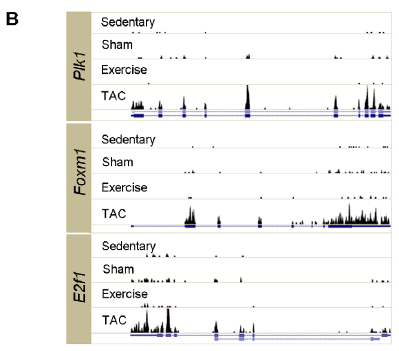
Figure 2. B) Histograms show increased gene expression for three transcription regulatory genes (Plk1, Foxm1, and E2f1) in TAC as compared to the other three test conditions. UCSC gene structures below histograms show exons as boxes.
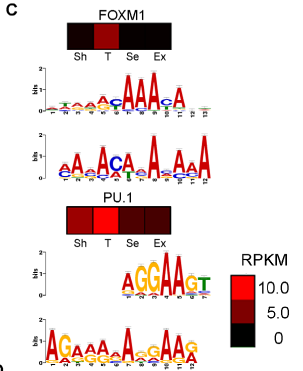
Figure 2. C) Heat maps show relative expression in RPKM of two putative transcription factors (FOXM1 and PU.1) in the four test conditions. Directly below each heat map is a MEME predicted binding motif determined by upstream sequence analysis for TAC enhanced genes. Below the predicted binging motif is the consensus motif for the specific transcription factor (FOXM1 or PU.1).
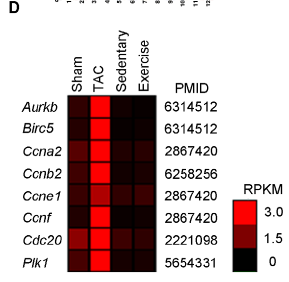
Figure 2. D) Heat map of gene expression (in RPKM) for eight known targets of the FOXM1 transcription factor shows elevated expression in TAC versus other test conditions.
Images reprinted under a Creative Commons license from Song et al. 2012