-This web page was produced as an assignment for an undergraduate course at Davidson College-
Toll-like Receptor 2
Introduction: Toll-like Receptors
Toll-like receptors (TLRs) are highly conserved type-I transmembrane receptors which were first discovered in fruit flies (Drosophila melanogaster), where they play a critical role in dorsal-ventral orientation and development. Since then, their homologs have been identified in insects, mammal, and plants, where they function in innate immune responses and microbial recognition. Toll-like receptors belong to the interleukin-1/Toll-like receptor superfamily, based on the shared cytosolic intracellular signaling domain that is known as the Toll/IL-1R (TIR) domain. The superfamily is divided into two subgroups based on the extracellular characteristics of the receptor, namely whether they possess an immunoglobulin-like domain or leucine-rich repeats (LRRs). Toll-like receptors are characterized by extracellular LRRs. The LRRs of the TLR extracellular domain contain 24-29 amino acid repeats with the pattern xxLxLxx (Akira and Sato, 2003).
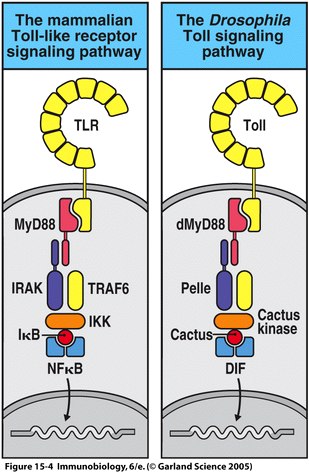
Homology between IL-1R and the Drosophila Toll cytosolic domain was first reported in 1991 by Gay and Keith. Since then, significant homology has been determined between other Drosophila and human proteins involved in the TLR signaling pathway. These proteins include dMyD88, Pelle, dTRAF, Cactus kinase, Cactus and DIF in Drosophila, which correspond with mammalian MyD88, IRAK, TRAF6, IKK, and IκB respectively (Fitzgerald and O'Neill, 2000); See Figure 1. In Drosophila embryos, activation of the Toll pathway leads to dorval-ventral axis development, while in adult Drosophila the same pathway is responsible for the formation of antimicrobial peptides (Janeway et al., 2005).
 |
Figure 1 depicts the homologous proteins described in the Drosophila Toll signaling pathway and the mammalian TLR signaling pathway. Figure 15.4 from Janeway, et al., 2005. |
Given the highly conserved nature of the Toll pathway, scientists had reason to suspect that TLRs in other organisms likewise played a role in innate immunity. Unlike the known mammalian models, Toll in Drosophila does not appear to function as a direct pathogen recognition receptor (PRR), and the actual PRR remains to be identified (Fitzgerald and O'Neill, 2000). While Drosophila possesses no adaptive immunity, the activation of the TLR pathway in mammals has been shown to initiate the first steps leading to an adaptive immune response (Takeda and Akira, 2003). Ten members of the TLR family are expressed in humans and mice and are designated TLR1 through TLR10; each receptor is responsible for recognition of a specific set of pathogen-associated molecular patterns (PAMPs) that are common to numerous microbial organisms (Janeway et al., 2005). The specificity of each TLR to different ligands and the differential response to pathogens suggests that individual TLRs have their own signaling pathway that relates to their activity, but contains many common steps. TLRs can be found on numerous cell types, but are believed to be most efficient at pathogen recognition when expressed on antigen presenting cells (APCs), which include dendritic cells and macrophages (Beg, 2002). The Toll pathway is not confined to mammalian systems; in plants, more than 100 genes encode for the TIR domain, where the Toll pathway confers resistance to a range of viral pathogens. The receptors in plants do not possess a transmembrane domain, but rather consists of a TIR domain with LRRs and a nucleotide binding site (NBS) (Takeda and Akira, 2003).
TLR2 is among the better characterized and studied Toll-like receptors, and has an essential role in detection of invading pathogens. TLR2's primary function is LP (lipoprotein) mediated signaling. Specifically, TLR2 recognizes peptidoglycans (PGN) and lipoproteins associated with Gram-positive bacteria, as well as a host of other microbial and endogenous ligands including peptidoglycan (Staphylococcus aureus), GPI anchors (Trypanosoma cruzi), lipoarabinomannan (Mycobacterium tuberculosis), porins (Neisseria meningitides), and zymosan (yeast cell wall component) (Akira and Sato, 2003). A full list of the ligands of TLR2 and their origins can be found in Table 1. Research suggests that TLR2 forms heterodimers with TLR1 and TLR6. Using TLR1, TLR2, and TLR6 knockout mice, scientists were able to determine that the different heterodimers recognize unique peptides; TLR2/6 recognizes diacylated forms of mycoplasmal lipoproteins and TLR2/1 recognizes triacylated bacterial lipopeptides. These heterodimers are advantageous as they expand the range of ligands that may be recognized via TLR2. Interestingly, TLR2 homodimers do not appear to activate signaling (Wetzler, 2000). The structure of the TIR cytosolic signaling domain of TLR2 is depicted below in Figure 2 .
Figure 2 depicts the structure of the cytosolic TIR signaling domain of human TLR2 as rendered by Jmol. Protein structure acquired from the Protein Data Bank. |
Research has indicated that TLRs and TLR2 in particular are capable of recognizing a number of endogenous ligands. Necrotic cells and their protein byproducts such as extracellular matrix breakdown peptides initiate production of NFκB, which is important for regulating expression of inflammatory and tissue-repair genes as well as dendritic cell maturation; research has shown that TLR2 expression is required for NFκB activation in necrotic cells. Furthermore, heat shock proteins HSP60, HSP70, and GP96 have been shown to activate TLR2/4 pathways, leading to the activation of NFκB and ultimately resulting in cytokine production and dendritic cell maturation, both of which are critical for an inflammatory response. The ability of both microbial and endogenous ligands to function through the TLR pathway may be advantageous in inducing inflammatory and tissue-repair responses; however it remains to be seen whether triggering of the TLR pathway by endogenous ligands results in formation of an adaptive immune response (Beg, 2002).
Table 1 provides a summary of the ligands (and their origin) which may be bound by TLR2, TLR2/TLR1 heterodimers, or TLR2/TLR6 heterodimers. Table 1 adapted from (Akira and Sato, 2003). |
The TLR2 molecule signals through a well-understood pathway that is found in most multicellular organisms. Activation of the TLR pathway by recognition of its ligand results in the activation of the transcription factor NF-κB and ultimately results in the increased expression of co-stimulatory molecules and inflammatory cytokines (Janeway et al., 2005). Some research indicates that the TLRs and TLR2 in particular might not always recognize their peptide directly, but rather interact with other pattern recognition receptors in lipid rafts or protein complexes. In this case, it has been hypothesized that extracellular TLR domains act as stabilizing signaling complexes (Sabroe et al., 2003). The signaling pathways used by TLRs share many of the same proteins as the IL-1R pathway. Activation of TLR leads to recruitment of the MyD88, which mediates the interaction between IL-1R associated kinases 1 and 4 (IRAK1 and IRAK4) (Akira and Sato, 2003). MyD88, formally known as myeloid differentiation factor 88, is a promiscuous adaptor protein for many receptors with a cytosolic TIR domain (Fitzgerald and O'Neill, 2000). Phosphorylation of IRAK4 in turn triggers the phosphorylation of IRAK1. IRAK1 then recruits tumor necrosis factor receptor-associated factor 6 (TRAF6), and the resulting change in conformation of the IRAK4/IRAK1/TRAF6 complex causes it to the leave the receptor complex. This complex then interacts with TAK1 binding protein 2 (TAB2) and the resulting TAB2/TAK1/TAB1/TRAF6 complex moves to the cytosol and leaves the IRAKs behind. In the cytoplasm TAK1 is activated and in turn activates MAP kinases and IKK. IKK is then responsible for activating NFκB by degrading IκB. NFκB subsequently induces gene transcription (Akira and Sato, 2003).
The pathway described above is known as the MyD88-dependent pathway, and all TLR signaling with the exception of TLR3 depends on this adaptor protein. Furthermore, research indicates that another TIR domain-containing protein known as TIRAP/Mal is essential for TLR2 and TLR4-dependent cytokine production by functioning as an adaptor in the MyD88-dependent pathway (Akira and Sato, 2003). TIRAP/Mal has a C-terminal TIR domain; mice deficient in TIRAP/Mal have been shown to have difficulty producing inflammatory cytokines when exposed to LPS and lipopeptides (Takeda and Akira, 2003). Additionally, TLR2 is known to interact with CD14 (Sabroe, et al., 2003). TLR2 requires the co-receptor CD14 in order to initiate signaling upon recognition of ligands (Wetzler, 2000). The regulation and turning-off of any pathway is equally important to its ability to become activated. Evidence indicates that an adaptor protein known as TOLLIP is responsible for regulation of the TLR2 signaling pathway by interacting with the TIR domain of TLR2 and inhibiting IRAK activation (Wetzler, 2000). Furthermore, one of the genes activated by NFκB is IκB, which has a inhibitory effect on NFkB and thus turns the pathway off (Janeway et al., 2005).
Figure 3 depicts the MyD88-dependent signaling pathway used by both IL-1R/AcP and the Toll-like Receptors. Figure 2 adapted from (Akira and Sato, 2003). |
The activation of the Toll pathway has several important effects in inducing innate immunity. These effects include the production of cytokines and chemokines, including IL-1, IL-6, CXCL8, IL-12 and TNF-α. One important medical condition related to the Toll pathway is septic shock. Systemic release of LPS and subsequent activation of the Toll pathway induces septic shock as a result of the systemic release of TNF-α. Binding of bacterial ligands such as lipoproteins to TLR2 on macrphages results in the production of nitric oxide (NO) which has toxic effects on bacteria, and also results in the secretion of cytokines such as IL-12, which stimulate production of IFN-γ by T cells when the T cells interact with macrophages (Janeway et al., 2005). A diagram depicting the interaction between bacteria, macrophages, and T cells can be found in Figure 4.
 |
Figure 4 depicts the ligation of TLR2 on macrophages by mycobacteria and the subsequent production of nitric oxide and IL-12. The macrophage then interacts with and stimulates the production of IFN-γ by T cells, resulting in the recruitment of more macrophages the killing of the intracellular bacteria. Fig. 11.15 from Janeway et al., 2005. |
Additionally, activation of the Toll pathway leads to increased expression of co-stimulatory molecules such as B7.1 (CD80) and B7.2 (CD96), which have important role in inducing adaptive immune responses to pathogens. These co-stimulatory molecules are expressed by macrophages and tissue dendritic cells; when presented with MHCII:peptide complexes these molecules activate naive CD4 T cells. The cytokines produced by activated NFkB help mediate migration of dendritic cells from the infected tissue to lymph nodes, where they may encounter and activate leukocytes involved in the adaptive immune response (Janeway et al., 2005). There are important health-related consequences that result from TLR2 deficiency or mutation. TLR2 deficient mice are susceptable to infection by Gram-positive bacteria as well as meningitis from Streptococcus pneumoniae. Furthermore, TLR2-deficient mice fail to fight infections from spirochetes, and both TLR2 and TLR4 are responsible for inducing innate immune responses to Mycobacterium tuberculosis (Takeda and Akira, 2003). While these results have been produced under laboratory conditions, it is sufficient to say that TLR2 is critical to real-world pathogen recognition and defense in mammalian immune systems, and that we as human beings benefit tremendously from this ancient pathway.
Key Functions of Toll-like Receptors
Future Questions and Research
Much remains to be seen about how TLRs recognize pathogen peptides en vivo The characterization of which TLRs respond to which pathogens has been conducted under controlled environments, and little is known about how the TLRs are able to recognize the pathogen peptides in the body. Additionally, research suggests that there might be a relationship between TLRs and Fc receptors, complement receptors, and lectins. Furthermore, TLRs might play a role in detection of viruses via the MyD88-independent pathway (Takeda and Akira, 2003).
The activation of TLRs by adjuvant has raised the possibility of using adjuvants separated from their toxic effects in order to promote new vaccine therapies. However, it is unlikely that development will occur due to the inability of TLRs to discriminate between the beneficial and harmful natures of different adjuvants, and they will respond with both an effective immune response and a damaging inflammatory response (Janeway et al., 2005).
Please click the ![]() or the link to view online abstract.
or the link to view online abstract.
![]() Akira, S, S Sato. (2003). Toll-like receptors and their signaling mechanisms. Scandinavian Journal of Infectious Diseases. 35: 555-62.
Akira, S, S Sato. (2003). Toll-like receptors and their signaling mechanisms. Scandinavian Journal of Infectious Diseases. 35: 555-62.
![]() Beg, AA. (2003). Endogenous ligands of Toll-like receptors: implications for regulating inflammatory and immune responses. Trends in Immunology. 23: 509-12.
Beg, AA. (2003). Endogenous ligands of Toll-like receptors: implications for regulating inflammatory and immune responses. Trends in Immunology. 23: 509-12.
![]() Fitzgerald, KA, L AJ O'Neill. (2000). The role of the interleukin-1/Toll-like receptor superfamily in inflammation and host defence. Microbes and Infection. 2: 933-43.
Fitzgerald, KA, L AJ O'Neill. (2000). The role of the interleukin-1/Toll-like receptor superfamily in inflammation and host defence. Microbes and Infection. 2: 933-43.
Janeway, CA, P Travers, M Walport, MJ Shlomchik. Immunobiology 6. New York: Garland Publishing, 2005.
![]() Kiyoshi, T, S Akira. (2003). Toll receptors and pathogen resistance. Cellular Microbiology. 5: 143-53.
Kiyoshi, T, S Akira. (2003). Toll receptors and pathogen resistance. Cellular Microbiology. 5: 143-53.
Sabroe, I, RC Read, M KB Whyte, DH Dockrell, SN Vogel, SK Dower. (2003). Toll-like receptors in health and disease: complex questions remain. The Journal of Immunology. 171: 1630-35.
![]() Wetzler, LM. (2003). The role of Toll-like receptor 2 in microbial disease and immunity. Vaccine. 21: S2/55-60.
Wetzler, LM. (2003). The role of Toll-like receptor 2 in microbial disease and immunity. Vaccine. 21: S2/55-60.
This page created by: Kyle Kinsell
Return to the Immunology Homepage
Return to Kyle Kinsell's Immunology Homepage