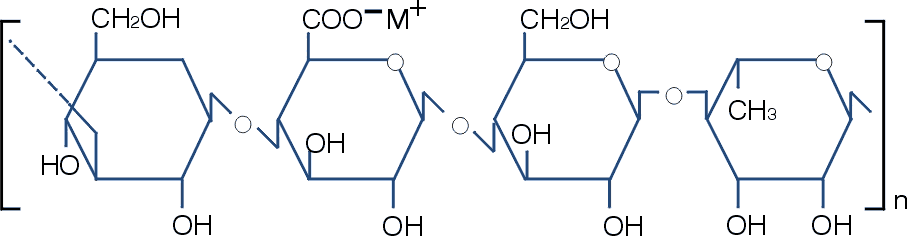
Structure of Gellan Gum. Above is the structure of gellan gum. The molecular structure of gellan gum consists of a linear repeating tetrasaccharide unit composed of glucose, rhamose and glucuronic acid. This image was taken from <www.kelco.com>
Reversible Gels for Electrophoresis and Isolation of DNA
Background
Gel electrophoresis is perhaps one of the most familar tools used by molecular
biologists. In gel electrophoresis, molecules of DNA or protein are
loaded into a gel (which is bathed in buffer), a current is applied, and
the negatively charged molecules migrate towards the positive electrode.
The porous gel through which the molecules migrate consists of different
size tunnels. Smaller molecules migrate faster than larger molecules
because they navigate more readily through the gel. More
info on gel electrophoresis
The most popular anti-convective materials for gel electrophoresis are polyacrylamide and agarose. The pore size of polyacrylamide makes it suitable for the separation of smaller nucleic acids (from a few base pairs to several thousand). Agarose, on the other hand, is best suited for the electrophoresis of large nucleic acids. Although these gels are relatively effective at separating different size molecules, they possess some undesireable properties. For instance, even purified agarose possesses varying amounts of negative charge. The presence of negative charges on the gel media results in a flow of buffer molecules toward the negative electrode due to electroosmosis (Cole, 1999). As a result of the opposing buffer flow, negatively charged molecules such as DNA are slowed in their progress.
Another difficulty concerns the success with which nucleic acids are recovered from isolated bands in agarose and acrylamide gels. In electro-elution, the band cut out of the gel is placed into a dialysis tube and an electric field is applied to move DNA or proteins out of the gel (Cole, 1999). This process often proves difficult and suffers from variable losses of material. Crush and soak techniques consist of cutting the band out, physically crushing the gel, and allowing the protein or nucleic acid to diffuse out of the gel into the buffer. Problems with these techniques include the slow diffusion process and variable recovery.
Reversible Gels
An alternative technique
for the recovery of DNA has recently been developed. This new technique
utilizes a carbohydrate polymer called gellan gum which forms strong gels
in the presence of divalent metal cations. However, the gel is reversible
in the sense that it can be converted back to a solution by the addition
of a chealting agent such as EDTA. Gellan electrophoresis gels can
also be formed using diamines. These gels are reversed by increasing
the pH, which results in the deprotonation of the diamine. Gellan
gum is a linear tetrasaccharide which is based on repeating glucose, rhamose
and glucuronic acid units. Gellan gum is produced by bacteria in
a fermentation process. Gellan gum undergoes deacylation and precipitation
steps which then produces a product that forms strong brittle gels at low
concentrations. The ability of gellan gum to form gels at low concentrations
combined with its reversibility have made it a promising candidate for
electrophoretic procedures.

Applications of Gellan Electrophoresis Gels
Gellan gum may be used to make electrophoresis gels in a range of polymer concentrations and buffer compositions. Once the gellan gum particles are in solution, the divalent cation is added. The divalent cation most commonly used to cast the gels is calcium, however, magnesium may also be used. The gels are characterized as mechanically strong yet brittle, which means that the gels will crack when not supported. Gellan electrophoresis gels as low as 0.03% may be constructed. These low concentration gels are preferable, because when converted back to solution, there are lower concentrations of gellan gum present in the resulting DNA solutions (Cole, 1999). Typical gellan electrophoresis gels have a concentration of 0.1%. These gels have sufficient mechanical strength and resolve DNA in the range of approximately 50-1kb (see figure 1 below). Furthermore, it has been found that higher concentrations of gellan gum do not improve the resolution of lower-molecular-weight DNA. Additionally, electrophoresis with higher concentrations has been found to be slower compared to 0.1% gellan electrophoresis gels.
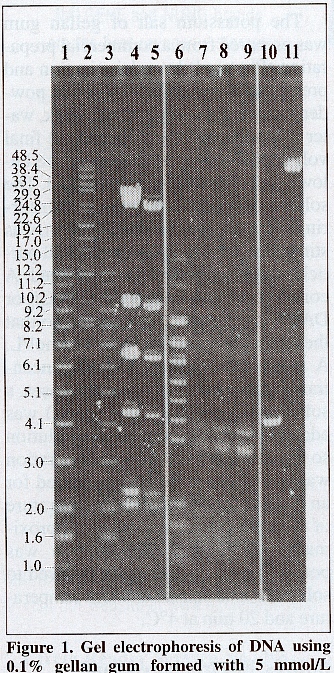
CaCl2. The buffer was TBC and electrophoresis
was run at 140 V for 16h. Lanes 1 and 3 are the kb ladder.
Lane 2 is the high-molecular-weight ladder. Lanes 4 and 5 depict
the HindIII restriction digest of lambda DNA. Land 7 is the
BstEII
restriction digest of lambda DNA. Lane 10 is pBR322 DNA restriction
digestion by HindIII. The last lane, lane 11 is the lambda
DNA (48.5 kbp). Numbers on the side indicate the size of DNA standards
from lanes 1 and 2 in kbp (Cole, 1999).
The electric field required for the gellan electrophoresis gel below was 3.5-fold higher than the agarose gel. This difference reflects the high electroosmosis in the gellan gum. A comparison of standard gellan and agarose electrophoresis gels reveals that staining with ethidium bromide is 4-fold less sensitive with gellan gum compared to staining in agarose (see figure 2 below). The gellan electrophoresis gels have low background fluorescence allowing long exposures, but the fluorescence intensity of the DNA was lower compared to agarose. When DNA is purposely overloaded (lanes 4 and 5 in figure 2), the capacity of gellan electrophoresis gels appears good, as judged by the lack of smearing between bands. Thus one can see that there are clear benefits to both gels.
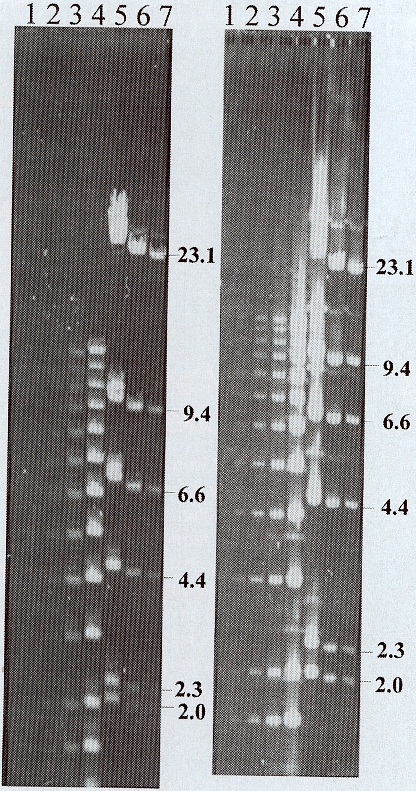
Figure 2. Comparison of loading and staining intensity of
gellan gum (first column) and agarose (second column) electrophoresis gels.
The gellan gel (0.1%) was electroporesed at 140V for 16h using TBC.
The agarose gel (0.5%) was electrophoresed at 40V for 16h using the TB
with EDTA. Lanes 1-4 are the kb ladder at 0.04, 0.16, 0.8, and 4.0
ug total DNA. Lanes 5-7 contain lambda DNA digested with HindIII
at 5.5, 1.1, and0.6 ug total DNA. The gellan gum gel was exposed
for 8s, and the agarose gel was exposed for 1s. Numbers on the side
are the size of the HindIII fragments of lambda DNA in kb (Cole,
1999).
The overall effectiveness of gellan electrophoresis gels can be improved with the addition of linear polymers before casting. For instance, the addition of linear polymers such as hydroxethyl cellulose (HEC) increases the resolution of lower-molecular-mass DNA and allows the gels to be run in a few hours. In addition, the electroosmosis is reduced. The increased resolution with the addition of HEC is probably due to several factors. Besides increasing the sieving of DNA, the viscous HEC solution decreases the diffusion of low-molecular-weight DNA. The decreased time for electrophoresis would also result in the decreased diffusion of low-molecular-weight DNA.
Restriction digestion, Ligation, and Transformation Using DNA Isolated from Gellan Electrophoresis Gels
Perhaps the best measure of the effectiveness of an electrophoretic gel is the readiness with which functional molecular techniques may be applied to the isolated DNA. Cole has acheived successful restriction digestion (using EcoR1, HindIII, and BstEII) of DNA isolated from gellan electrophoresis (1999). It has further been found by Cole that dissolved gellan gum solutions do not inhibit the self-ligation of DNA fragments (1999). Finally, the influence of gellam gum on transformation efficency has been shown to be negligible (Cole, 1999).
Gellan gum offers an alternative for the separation and isolation of DNA. When additional polymers are added to a gellan solution, they influence the separation of the bands on the gel by lowering the electroosmotic effect. The chemical structure of the polymer, the molecular weight, and the concentration all influence the separation (Cole, 1999).
Sources
1. Cole, K.D. 1999. Reversible gels for electrophoresis and isolation of DNA. BioTechniques 26:748-756 (April).
For additional info on gellan gum visit these sites:
www.kelco.com (Describes the properties and structure of gellan gum)
http://bio3.ist.utl.pt/bsrg/ResTopics/Biotechgellan.html (Biotechnology of gellan gum)
http://www.acs.org/vc2/1rp/rp1_orb.html (A less successful application of gellan gums)
Send comments, questions, and suggestions to: jubussone@davidson.edu