
My Favorite Yeast Proteins
AAC3 (aka: YBR085W) Protein:
code:
MSSDAKQQET NFAINFLMGG VSAAIAKTAA SPIERVKILI QNQDEMIKQG TLDKKYSGIV DCFKRTAKQE GLISFWRGNT ANVIRYFPTQ ALNFAFKDKI KLMFGFKKEE GYGKWFAGNL ASGGAAGALS LLFVYSLDFA RTRLAADAKS SKKGGARQFN GLTDVYKKTL KSDGIAGLYR GFMPSVVGIV VYRGLYFGMF DSLKPLVLTG SLDGSFLASF LLGWVVTTGA STCSYPLDTV RRRMMMTSGQ AVKYNGAIDC LKKIVASEGV GSLFKGCGAN ILRSVAGAGV ISMYDQLQMI LFGKKFK
SGD http://genome-www4.stanford.edu/cgi-bin/SGD/YPD/ypd.pl?locus=aac3
New information on AAC3 from FunctionJunction :
pI = 9.95 (isoelectric point occurs at a pH of 9.95), other names for the protein include ANC3, YBR0753
cellular role: energy generation and small molecule transport.
Information from the Yeast PathCalling Home Page (CuraGen & University of Washington):
http://genome-www.stanford.edu/cgi-bin/SGD/functionJunction - Picture of AAC3 interactions with other proteins -CCT4 and SAC6.

On the right side of the screen picture above, it can be seen that the AAC3 protein interacts with both CCT4 and SAC6. CCT4 codes for a chaperonine protein. The SAC6 protein is involved in endocytosis, osmotic responses, and polar budding. Because AAC3 is an antiporter, it may create osmotic differences across the membrane it lies in. If this is true, it could interact with the SAC6 protein to resolve such osmotic problems. Furthermore, because the AAC3 protein is involved in moving ATP, the currency of the cell, and endocytosis requires energy, the SAC6 protein and the AAC3 protein may interact in order to allow the AAC3 protein to give the SAC6 protein the energy it needs to carry out its function.
http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=PubMed
YDL246C Interactions:
Yeast Path Calling Map:
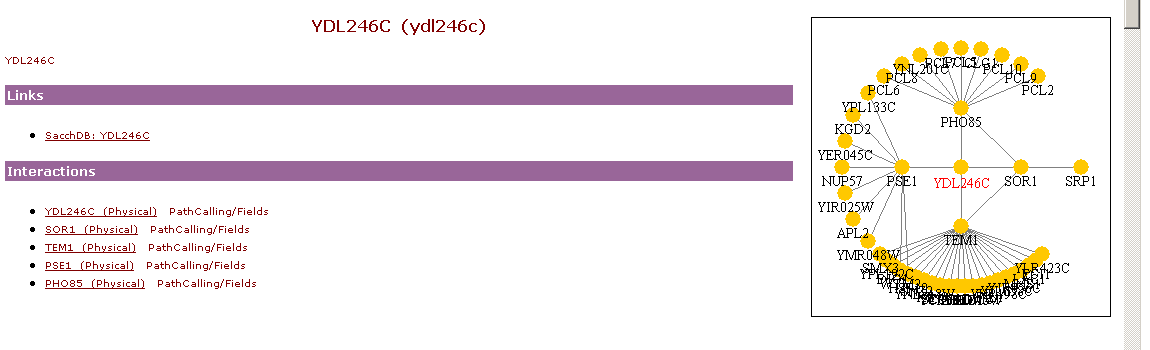
http://genome-www.stanford.edu/cgi-bin/SGD/functionJunction
The right side of the above picture shows the interaction between SOR1, TEM1, PSE1, PHO85, and YJR037W.
Interaction Map A
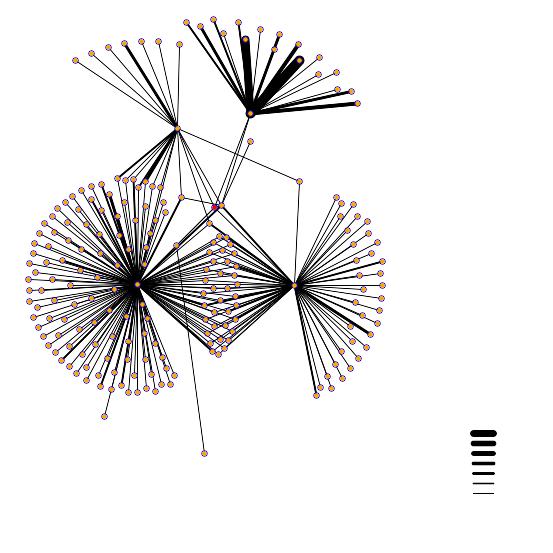
In the above interaction map, the red dot in the middle of the picture is the YDL246C protein. There are four distinct lines coming out of the YDL246C protein. They connect to to the following proteins and are located in the following positions: YJR037W - is located between the, large cluster in bottom left and the central red dot. TEM1 in bottom right - GTP binding protein. PHO85, upper right Protein Kinase. PSE1, upper left. SOR1 - the dot located directly on the right side of the red dot.
My unannotated YDL246C protein is likely an alcohol dehydrogenase because it posses long-chain alcohol dehydrogenase homology.
The functions of the proteins that it interacts with are listed below:
PHO85: kinase-related transforming protein; protein kinase homology.
TEM1: kinase-related transforming protein; protein kinase homology.
SOR1: L-iditol 2-dehydrogenase, YJR037W: hypothetical protein.
PSE1: transmembrane protein.
SPR1: is an integral membrane protein in the nucleus involved in the elongation and sliding of microtubules in mitosis.
Interaction Map B:
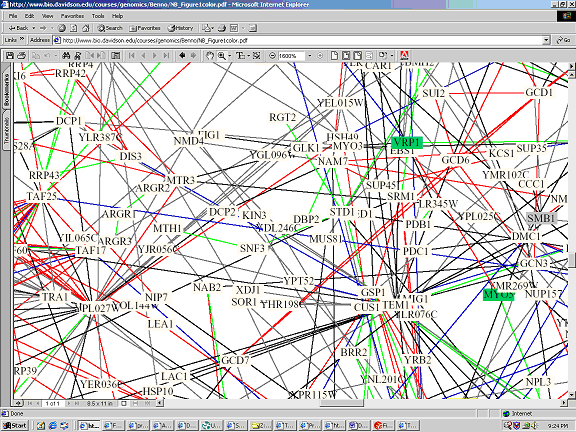
Full Image: http://bio.davidson.edu/courses/genomics/Benno/NB_Figure1color.pdf
Key: Red - cellular role and subcellular localization of interacting proteins are identical. Blue - localizations are identical but functions differ. Green - cellular roles are identical but localizations differ. Black - cellular role and localization are different or unknown.
It is interesting to note that SRP1 is not connected to YDL246C on the above interaction map (interaction map B), however the text section of the Dipdoe website states that there is an interaction. On the first interaction map (the Yeast Path Calling Map) there appears to be an interaction between SRP1 and YDL246C, but this interaction occurs through the intermediary SOR1. These discrepancies may be due to the different conditions in which the interactions were studied, or simply the fact that different proteins may behave differently at different times.
http://genome-www.stanford.edu/cgi-bin/SGD/functionJunction
According to a SGD worm-yeast protein comparison, the protein again appears to be a sorbitol or alcohol dehydrogenase (as was previously thought). It shows outstanding similarity with the yeast SOR1 L-iditol 2-dehydrogenase gene. In interaction map A, SOR1 shares every connection with YDL246C except one. This one connection goes to a protein called NUP1. NUP1 is a nuclear pore protein that is required, if not substituted for by NUP2, for proper growth. Its function is to move proteins between the nucleus of the cell and the cytoplasm.
http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=sor1;
To better understand the function of the YDL246C ORF, a better understanding of the proteins that it interacts with is needed. It seems fairly clear, through homology, that YDL246C is in fact a dehydrogenase. As a dehydrogenase, it is probably involved in an oxidation-reduction reaction somewhere in the cell. It is highly likely that it removes a hydrogen from one substrate and transfers it to another. The protein is alleged to interact with PHO85, PSE1, SPR1, TEM1, YJR037W, and SOR1. All of these proteins make excellent possibilities for designing experiments. However, it might be possible to design an experiment that illuminates the basic function of the protein and its location in the cell.
Experiments:
For the YDL246C protein, I would first like to know exactly where it is located. I would use the DNA sequence to create an abundance of the protein. I would then inject the protein into an animal such as a rabbit in order to create an antibody. After isolating this antibody, I would label it and then let it loose in a yeast cell. Hopefully, the labeled antibody would then lead me to the location of the YDL246C protein. From the protein's location, a lot could be learned. I expect to find the protein in a mitochondrial membrane or in the cell wall membrane as part of some oxidation-reduction reaction. I would also create antibodies to the above listed proteins that YDL246C is alleged to interact with. In separate cell samples, I would use these antibodies to test for the locations of these other proteins. If they to were located in the same area/membrane as YDL246C, then I would expect them to be part of the same reaction chain that YDL246C is involved in.
After discovering the location, it would be easier to understand what other proteins it interacts with, and perhaps, to some extent, why. This would be especially true if the protein was found in one of the mitochondrial membranes. If it were located there, it could be some as of yet unexplored part of the energy creating mechanisms of the cell. The protein is alleged to be an alcohol dehydrogenase. To test this, I would first insert extra copies of the gene or its transcription factor into a yeast cell in order to cause it to be over expressed. I would grow the yeast cells (in an anaerobic environment) in the presence of a large quantity of ethanol that has had deuterium substituted for its hydrogens. This heavy alcohol, if dehydrogenated (or perhaps I should say dedeuterianated) would leave deuterium molecules in the cell. If the YDL246C protein is part of the metabolic pathway for the alcohol, then the deuterium atoms might end up being added to the final metabolic water product. If not the final water product, then the deuterium would be incorporated somewhere else. By comparing the weight of a given volume of water or protein from the YDL246C-overexpressed cell against the same amount of water or protein from two control cells (one in which non-deuterinated ethanol was the food source, and another in which deuterinated ethanol was used in conjunction with a cell that was not overexpressing YDL246C), I could possibly determine where the deuterium atoms went and whether or not the overexpression of the YDL246C protein lead to an increase in deuterination of one of the products. This experiment, if successful, in conjunction with my first experiment, would lead directly to the cellular function and location of YDL246C.
A final experiment, would be to study the interaction of YDL246C with the 5 above listed proteins (especially with the questionable SRP1 interaction) under various partial pressures of oxygen. A protein-protein interaction macroarray, in which my protein was plated, and the other proteins were used as probes, would show the effects of differing partial pressures of Oxygen on the interactions of the proteins.