This web page was produced as an assignment for an undergraduate course at Davidson College
Microarray Analysis of my Favorite Yeast Genes
A Brief Introduction to Microarrays:
The use of microarrays is a brand new technique that allows a researcher to examine the response of an entire genome to changing conditions. A microarray is a slide onto which single-stranded copies of thousands of genes have been spotted. Experimentally treated cells are lysed and their RNA is allowed to hybridize with the DNA on the chip. By comparing the amount of bound RNA between experimental and control cells, changes in expression resulting from the experimental treatment can be seen.
Microarray analysis of the annotated gene TUB1/YML085C:
On the previous page I explained that TUB1 is the yeast gene that encodes for alpha tubulin, an important component of microtubules in the cell cytoskeleton and an integral player in chromosome movement during mitosis. To see this page click here. On this page I will analyze expression patterns of TUB1 under a variety of experimantal conditions using microarray data available at http://genome-www4.stanford.edu/cgi-bin/SGD/expression/expressionConnection.pl .
TUB1 expression when exposed to alpha factor hormone:
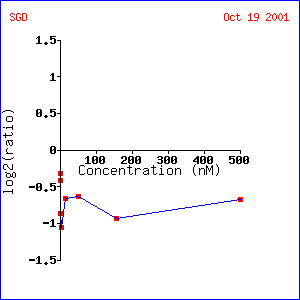
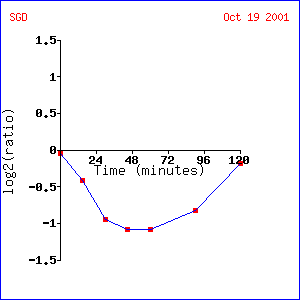
Yeast (Saccharomyces cerevisiae) are capable of reproducing either asexually (budding) or sexually. Alpha factor is a hormone that stimulates yeast to reproduce sexually (3). Exposure to alpha factor arrests cells in the G1 phase of the cell cycle and prompts the expression of genes involved in sexual reproduction (3). TUB1 is generally repressed when exposed to alpha factor. The repression is greatest (about a 1-fold repression) at an alpha factor concentration of about 200 nM. This makes sense because when a call is exposed to alpha factor it stops budding and therefore stops mitosis. As tubulin is involved in chromosome sorting during mitosis it makes sense that TUB1 is repressed. Not surprisingly, TUB2 and TUB3, other components of tubulin (see other page) showed very similar expression patterns to TUB1. In addition, the gene STU2, a gene encoding a protein involved in microtubule nucleation also showed a similar expression pattern. When TUB1 expression over time after exposure to alpha factor was examined, quick repression with a gradual rise back to control levels was observed. Repression was greatest after about 48 minutes and had returned to control levels by 120 minutes. In this experiment, TUB1 was seen to have an expression pattern similar to SPC34, another gene encoding a protein involved in microtubule nucleation.
 |
 |
|
Expression of TUB1 under varying concentrations of
alpha factor (1)
|
Expression of TUB1 over time after exposure to alpha
factor (1)
|
|
Below: Genes that show similar expression patterns
to TUB1 under varying concentrations of alpha factor (1)
|
|
Scale : (fold repression/induction)
![]()
Click on a color strip to see data for that gene.
|
Orf |
|
Gene |
|
|
|
Process |
|
Function |
|
Component |
|
|
TUB1 |
|
|
mitotic chromosome
segregation* |
|
structural protein of
cytoskeleton |
|
spindle pole body* |
||
|
|
ALG5 |
|
|
not yet annotated |
|
dolichyl-phosphate
beta-glucosyltransferase |
|
not yet annotated |
||
|
|
CLB1 |
|
|
regulation of CDK activity* |
|
G2/M-specific cyclin |
|
cellular_component unknown |
||
|
|
CDC11 |
|
|
establishment of cell
polarity (sensu Saccharomyces)* |
|
structural protein of
cytoskeleton |
|
septin ring (sensu
Saccharomyces)* |
||
|
|
TUB2 |
|
|
mitotic chromosome
segregation* |
|
structural protein of
cytoskeleton |
|
spindle pole body* |
||
|
|
KIP2 |
|
|
mitotic anaphase B* |
|
microtubule motor |
|
cytoplasmic microtubule* |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
TUB3 |
|
|
mitotic chromosome
segregation* |
|
structural protein of
cytoskeleton |
|
spindle pole body* |
||
|
|
FEN1 |
|
|
fatty acid biosynthesis* |
|
molecular_function unknown |
|
endoplasmic reticulum
membrane |
||
|
|
SNU56 |
|
|
mRNA splicing |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
HOG1 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
||
|
|
HOS3 |
|
|
biological_process unknown |
|
molecular_function unknown* |
|
not yet annotated |
||
|
|
NDD1 |
|
|
not yet annotated |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
CDC5 |
|
|
protein amino acid
phosphorylation* |
|
protein serine/threonine
kinase |
|
cytoplasm* |
||
|
|
CDC1 |
|
|
mating (sensu Saccharomyces)* |
|
molecular_function unknown |
|
cell |
||
|
|
HTA2 |
|
|
not yet annotated |
|
not yet annotated |
|
nucleosome |
||
|
|
HST3 |
|
|
not yet annotated |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
STU2 |
|
|
microtubule nucleation |
|
structural protein of
cytoskeleton |
|
spindle pole body |
||
|
|
FIN1 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
||
|
|
APG13 |
|
|
autophagy* |
|
molecular_function unknown |
|
cellular_component unknown |
||
|
|
|
|
|
|
|
|
|
|
|
|
|
Orf |
|
Gene |
|
|
|
Process |
|
Function |
|
Component |
|
|
TUB1 |
|
|
mitotic chromosome
segregation* |
|
structural protein of
cytoskeleton |
|
spindle pole body* |
||
|
|
SWI6 |
|
|
cell cycle |
|
transcription factor |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
PEX5 |
|
|
protein-peroxisome
targeting* |
|
peroxisome targeting
sequence binding* |
|
cytosol* |
||
|
|
SPC34 |
|
|
microtubule nucleation |
|
structural protein of
cytoskeleton |
|
spindle pole body |
||
|
|
ERG3 |
|
|
ergosterol biosynthesis |
|
C-5 sterol desaturase |
|
endoplasmic reticulum |
||
|
|
CSM2 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
DAM1 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
cellular_component unknown |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
AOR1 |
|
|
not yet annotated |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
HHO1 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
PXL1 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
BUB2 |
|
|
mitotic spindle checkpoint |
|
molecular_function unknown |
|
spindle pole body |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
MNN2 |
|
|
protein amino acid
glycosylation |
|
not yet annotated |
|
not yet annotated |
||
|
|
RHK1 |
|
|
protein amino acid
glycosylation |
|
not yet annotated |
|
not yet annotated |
||
|
|
AXL2 |
|
|
axial budding* |
|
molecular_function unknown |
|
integral plasma membrane
protein* |
||
|
|
RSR1 |
|
|
polar budding* |
|
signal transducer* |
|
plasma membrane* |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
cellular_component unknown |
||
|
|
DOT6 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
TUB1 expression in response to changing environmental conditions:
In an experiment that measured the expression of yeast genes under a wide range of environmental changes including heat and cold chock and exposure to menadione and DDT, TUB1 was found to cluster with some of the same genes that it clustered with in the alpha factor experiments. Not surprisingly, these included TUB2 and TUB3. Also included was CIS3, a gene that encodes for a protein involved in cell wall structure. It is not surprising that many of the structural protein genes cluster together in many of these experiments.
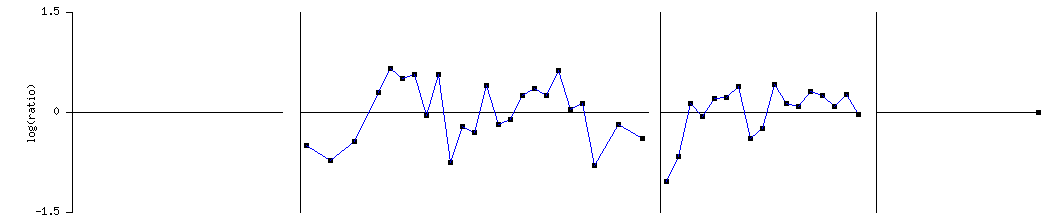
TUB1 expression through the cell cycle:
Levels of TUB1 expression were found to vary throughout the cell cycle. This makes sense as differing amounts of microtubules and consequently differing levels of alpha tubulin are necessary at differing points in the cell cycle. Again, TUB1 was found to cluster with TUB2 and CIS3 in this experiment.
 |
|
Expression of TUB1 through the cell cycle (1)
|
In Conclusion:
The microarray data presented here support the fact that TUB1 encodes alpha tubulin, part of tubulin that makes up microtubules. TUB1 consistently clusters with TUB2 and TUB3, the genes that encode the other subunits of tubilin. In addition, it clusters with other genes involved in microtubule formation such as CIS3 and SPC34.
Microarray Analysis of the Unannotated Yeast Gene LEE1/YPL054W:
LEE1 is an unannotated yeast gene whose function has yet to be scientifically investigated. In the previous page BLAST searches discovered tow zinc finger regions that lead me to hypothesize that LEE1 may encode a transcription factor. To see these data, click here. Now I will analyze microarray data using microarray data available at http://genome-www4.stanford.edu/cgi-bin/SGD/expression/expressionConnection.pl to see if I can make any further hypotheses on the function of LEE1.
LEE1 expression when exposed to alpha factor hormone:
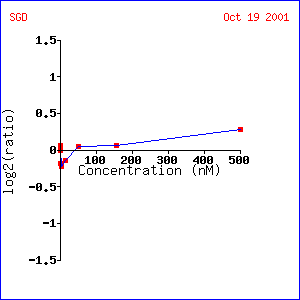
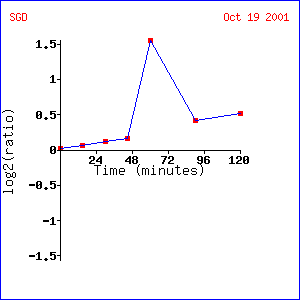
LEE1 showed almost no change when exposed to varying concentrations of alpha hormone. The lack of change in this experiment is explained when the response to alpha factor over time is examined. LEE1 shows almost no change in expression for the first 48 minutes after being exposed to alpha factor. Then there is a sharp, almost 1.5 fold induction that gradually falls afterwards. These data suggest that LEE1 is somehow involved in yeast mating. Probably, since it takes so long to be induced, LEE1 could be involved in the replication or exchange of DNA after two cells have mated. Unfortunately, though, almost all of the genes that show expression patterns similar to LEE1 are unannotated so it is hard to predict exactly what role LEE1 might play in this process.
 |
 |
|
Expression of LEE1 under varying concentrations of
alpha factor (1)
|
Expression of LEE1 over time after exposure to alpha
factor (1)
|
|
Below: Genes that show similar expression patterns
to LEE1 over time after exposure to alpha factor (1)
|
|
|
Orf |
|
Gene |
|
|
|
Process |
|
Function |
|
Component |
|
|
LEE1 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
RCN1 |
|
|
not yet annotated |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
FOX2 |
|
|
fatty acid beta-oxidation |
|
3-hydroxyacyl-CoA
dehydrogenase* |
|
peroxisomal matrix |
||
|
|
BOP2 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
AUT1 |
|
|
autophagy* |
|
molecular_function unknown |
|
cellular_component unknown |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
RNY1 |
|
|
biological_process unknown |
|
not yet annotated |
|
not yet annotated |
||
|
|
YPS6 |
|
|
not yet annotated |
|
aspartic-type endopeptidase |
|
not yet annotated |
||
|
|
INO1 |
|
|
not yet annotated |
|
inositol-3-phosphate
synthase |
|
not yet annotated |
||
|
|
PCL8 |
|
|
regulation of glycogen
biosynthesis |
|
cyclin-dependent protein
kinase regulator |
|
not yet annotated |
||
|
|
TES1 |
|
|
fatty acid metabolism |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
UBP11 |
|
|
not yet annotated |
|
ubiquitin-specific protease |
|
cellular_component unknown |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
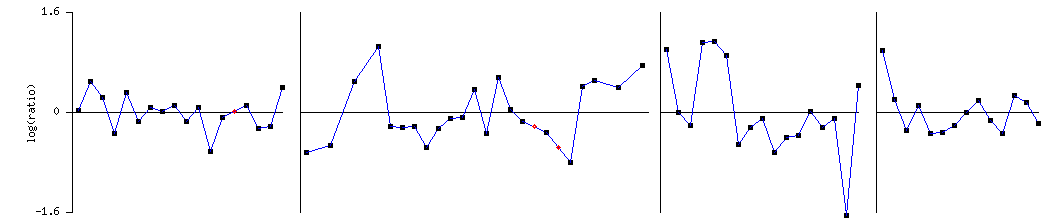
LEE1 expression through the cell cycle:
LEE1 shows varying expression throughout the cell cycle. It does show a marked peak though during S phase. S phase is when replication occurs (2), indicating that LEE1 may be involved in DNA replication. Unfortunately, again, almost no genes with known function cluster with LEE1 in this experiment.
 |
|
Expression of LEE1 through the cell cycle (1)
|
LEE1 expression during sporulation:
This experiment found that LEE1 is initially induced and then repressed during sporulation. This would be consistent with the hypothesis that LEE1 is involved in replication of DNA. In this case, DNA would be replicated first and then spore formation would ensue. Although this may seem similar to alpha factor response, the initial lag before induction of LEE1 in response to alpha factor could be explained be the time it takes for a yeast to shmoo to another yeast cell before DNA replication begins.
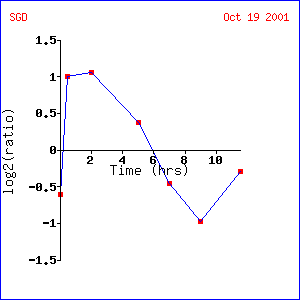 |
|
|
Expression of LEE1 during sporulation (1)
|
|
|
Below: Genes that show similar expression patterns
to LEE1 during sporulation (1)
|
|
Orf |
|
Gene |
|
|
|
Process |
|
Function |
|
Component |
|
|
LEE1 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
SCD6 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
YRR1 |
|
|
transport |
|
not yet annotated |
|
not yet annotated |
||
|
|
MNT3 |
|
|
O-linked glycosylation |
|
alpha-1,3-mannosyltransferase |
|
cellular_component unknown |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
PAN5 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
cellular_component unknown |
||
|
|
SNP1 |
|
|
mRNA splicing |
|
not yet annotated |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
BUD8 |
|
|
pseudohyphal growth* |
|
molecular_function unknown |
|
cell |
||
|
|
MON1 |
|
|
not yet annotated |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
SSL1 |
|
|
transcription initiation,
from Pol II promoter* |
|
general RNA polymerase II
transcription factor |
|
transcription factor TFIIH |
||
|
|
WTM1 |
|
|
meiosis |
|
transcription factor |
|
not yet annotated |
||
|
|
SFH1 |
|
|
chromatin modeling |
|
molecular_function unknown |
|
nucleosome remodeling
complex |
||
|
|
PRY3 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
YSA1 |
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
molecular_function unknown |
|
not yet annotated |
||
|
|
|
|
|
biological_process unknown |
|
not yet annotated |
|
not yet annotated |
||
|
|
CAF4 |
|
|
not yet annotated |
|
not yet annotated |
|
not yet annotated |
In Conclusion:
The gene clustering data for LEE1 from these experiments was not very informative. In most cases the vast majority of clustered genes were of unknown function and those that were known did not show any clear trends. From expression patterns, though, it seems at least possible that LEE1 may be involved in DNA replication. Coupled with the data from the previous page that suggest that LEE1 is a transcription factor, my best guess would be that LEE1 is a transcription factor that regulates genes involved in DNA replication.
Sources:
1) SGD, Expression connection. 2001.http://genome-www4.stanford.edu/cgi-bin/SGD/expression/expressionConnection.pl
2) "The Cell Cycle" http://pc65.frontier.osrhe.edu/hs/science/bcell2.htm
3) Zymo Research. 2001. Alpha-factor mating pheromone. http://pc65.frontier.osrhe.edu/hs/science/bcell2.htm
please email me at jdwillson@davison.edu with comments or questions
Davidson | J.D.'s Genomics Home | Genomics Home