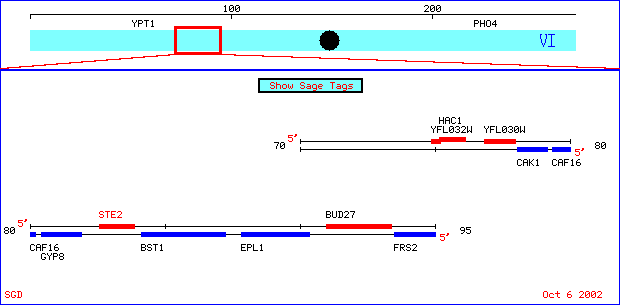
Cytogenetic position of STE2 (and YFL030W). Graphic courtesy SGD. <http://genome-www4.stanford.edu/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0001868>
Ste2 is a gene found in chromosome 6 (position coordinates 82578 to 83873) of the yeast Sacchromyces cerevisiae. It encodes a protein that a seven-transmembrane domain protein directly involved in the reproductive cycle of these yeast. S. cerevisiae are found in two mating types, a and alpha, and each makes a pheromone to induce transcription of genes necessary for mating. This pheromone, in turn, is an extracellular signal that binds to the alpha-factor pheromone receptor that the gene Ste2 encodes. Ste2, therefore, is found in type a cells. After binding alpha factor, Ste2 protein undergoes a conformational change and the associated G-alpha subunit initiates further components of the pheromone response pathway. The pheromone receptor encoded by the yeast gene is a class D G-protein coupled receptor (SGD, 2002, <http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=ste2>). The Sacchromyces Genome Database webpage about integral membrane proteins in yeast is helpful in understanding protein relationships in the yeast cell. Mutants without this gene are viable but sterile, which is further evidence pointing to this genes relationship to yeast reproduction. To see the complete DNA and amino acid sequences of Ste2, click here. Ste3 is similar to function as Ste2, but it is found in alpha cells and binds to a different pheromone (a factor).
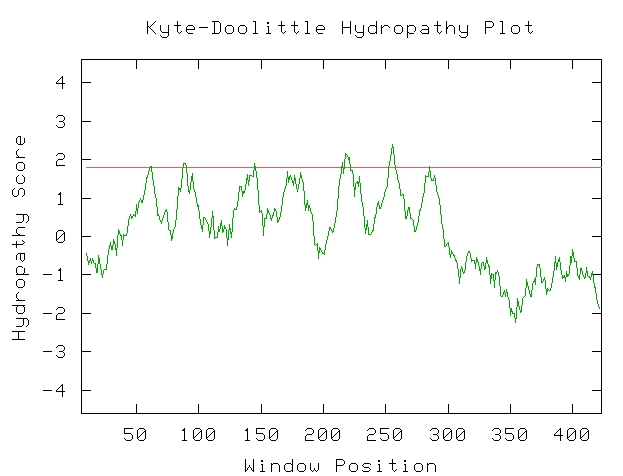
Screen shot from a search on the Kyte-Doolittle Page (KYTE DOOLITTLE HYDROPATHY PLOT, 2002)
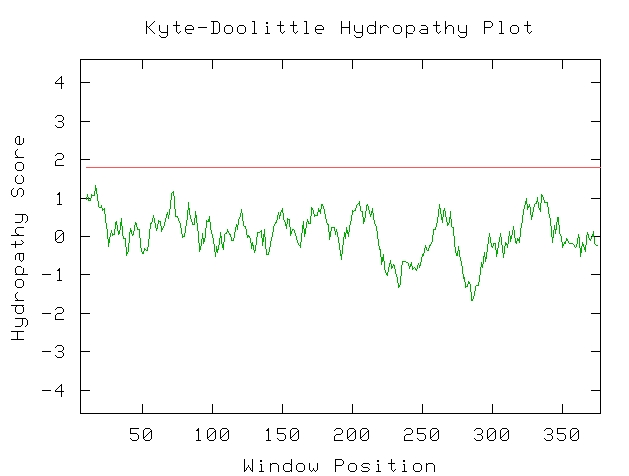
The Kyte-Doolittle hydropathy plot graphs a sequence's potential at having transmembrane regions. As is consistent with the literature, the STE2 gene does indeed have seven transmembrane regions, necessary for the protein to perform its function. Even though not all of the peaks cross the line that indicates transmembrane characteristics, the peaks are very regular and can be interpreted as accurately indicating the literature's claims.

Screen shot from a search on the PREDATOR (PREDATOR, 2002, <http://npsa-pbil.ibcp.fr/cgi-bin/secpred_preda.pl>)
Using the PREDATOR website, it is noted that the amino acids in the pheromone receptor have multiple alpha helices. This is to be expected, as Ste2 codes for an integral membrane protein which typically have multiple alpha helices.
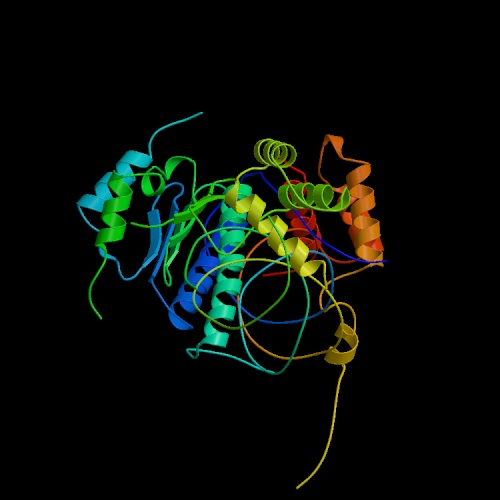
This picture is not of the product of Ste2, which is unavailble. This is the product of Ste6, which is similar in function. Courtesy PDB.
As would be expected, the literature regarding this gene is incredibly extensive, as this is many a researcher's area of expertise. The STE2 gene and the protein it encodes integrates itself nicely into different areas of yeast research, as multiple pathways are involved in producing the G-protein and many factors regulate these processes. For instance, research by Martin et al. shows us multiple recessive alleles found within the STE2 gene and their ability to retain pheromone receptor function:
A total of 157 amino acid positions in seven different mutagenic libraries corresponding to the seven predicted transmembrane segments were analyzed, yielding 390 alleles that retain at least 60 % of normal signaling function. These alleles contained a total of 576 unique amino acid substitutions, including 61 % of all the possible amino acid changes that can arise from single base substitutions. The receptor exhibits a surprising tolerance for amino acid substitutions. (Martin et al., 2002)
Also mentioned in the literature is a study conducted by Duran-Avelar, et al., whose purpose was to examine mutants that disrupted the Ste2 gene's coding of the alpha-pheromone. Again, the mutant they found, 8L4, showed reduction in growth, diminished activation of Fus1 gene, and impaired mating competence (Duran-Avelar, et al., 2002). There are many similar articles floating in the literature, as is found in a PubMed search under the simple heading "Ste2".