This web page was produced as an assignment for an undergraduate course at Davidson College.
Yeast Genes: MSR2 vs YOR331C
Preface: This web page will investigate one annotated gene and one non-annotated gene from Saccharomyces cerevisiae (yeast). The two yeast genes I chose for this page were MSR2 and YOR331C.
MSR2
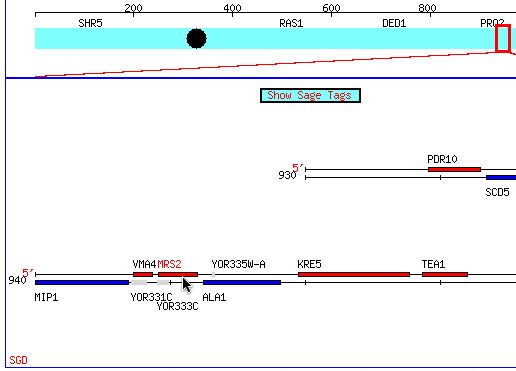
Figure 1: The cursor points to the location of the MSR2 gene on Chromosome XV. Permission obtained from SGD(citation). (http://db.yeastgenome.org/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0005861)
Biological Process:
MSR2 is attributed with two biological processes, Group II intron splicing and mitochondrial magnesium ion transport.
 Definition
from SGD, 2002; http://db.yeastgenome.org/cgi-bin/SGD/GO/go.pl?goid=373
Definition
from SGD, 2002; http://db.yeastgenome.org/cgi-bin/SGD/GO/go.pl?goid=373
Mitochondrial magnesium ion transport is the movement of magnesium ions into or out of the membrane of a mitochondria organelle.
Biological Function:
MSR2 is responsible for the magnesium ion tranporter activity within yeast cells. The molecular function that MSR2 is involved in is the "directed movement" of magnesium ions through the cell membrane of yeast and even between individual yeast cells (SGD, 2002; http://db.yeastgenome.org/cgi-bin/SGD/GO/go.pl?goid=15095).
Cellular Component:
The MSR2 gene codes for a magnesium ion transporter protein in both mitochondria and the cell membrane. For this reason, the location of the protein is within the cell membrane and the membrane of mitochondria.
Exonic and Chromosomal Coordinates(ChrXV)
Exon: 1 - 1413
Chromsome: 944589 - 946001
Information from SGD.
Sequence Information
Protein Information
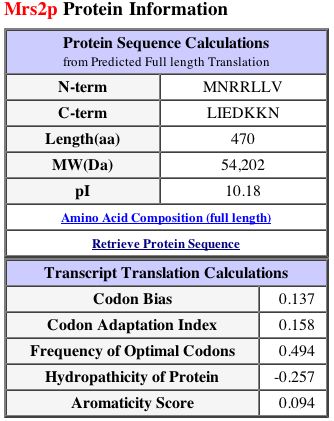
Figure 3: This figure displays protein information and transcript translation calculations for MRS2. Image from SGD, 2002; http://db.yeastgenome.org/cgi-bin/SGD/protein/protein?sgdid=S0005861. Permission pending.
Location of the Magnesium Ion Transporter
To verify that the MSR2 gene codes for a transmembrane protein, this web site will use a Kyte-Doolittle Hydropathy plot of the amino acid sequence.
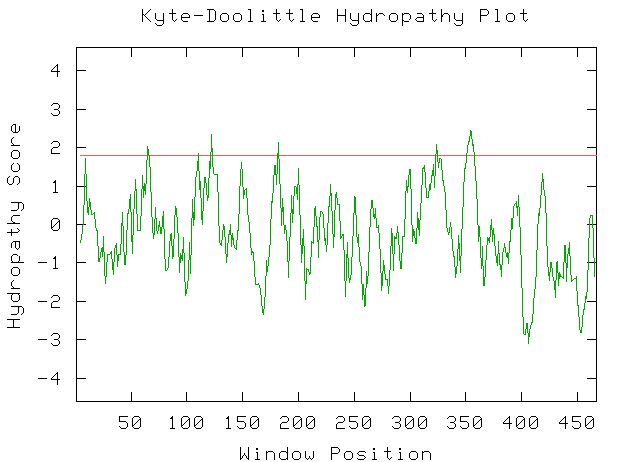
Figure 3: This is the Kyte-Doolittle Hydropathy plot for the amino acid sequence of MSR2. Every peak above the red line suggests a transmembrane region of the protein. Image from http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/activities/kd/kyte-doolittle.htm. Permission pending.
Function of Magnesium Ion Transporter
According to SGD, the function of the magnesium ion transporter is for the splicing of group II introns (SGD 2002, http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=YOR334W). The activity of the magnesium ion is apparently critical for group II intron splicing.
In reality, Gregan et al explains that the MRS2 gene product, Mrs2p actually has an indirent effect on group II intron RNA splicing by providing mitochondria with a "suitable Mg2+ concentration." Interestingly, if yeast mitochondria are devoid of Mrs2p proteins and are still provided with normal concentrations of magnesium, normal mRNA splicing will occur. Therefore, the Mrs2p proteins are essential for group II intron splicing because they provide sufficient levels of magnesium to the mitochondria. Without the indirect effect of Mrs2p, mitochondria would be blocked from group II intron splicing because of a lack of magnesium (Gregan J et al. 2001; http://www.genesdev.org/cgi/content/full/15/17/2229).
Structure
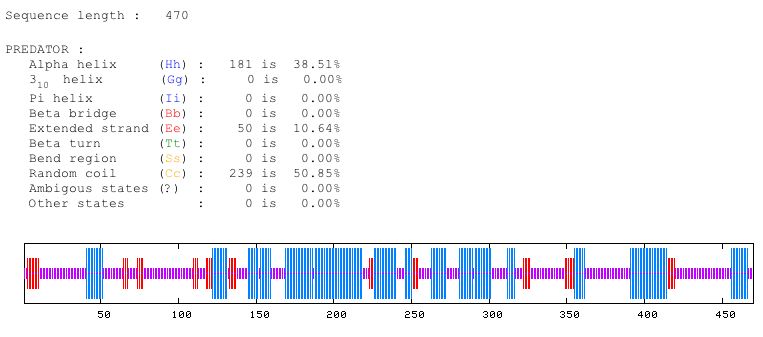
Figure 4: Predator predicts that approximately 38 percent of the protein is alpha helices, 10 percent is extended strand, and 50 percent is random coil. Image from Predator. Permission pending.
Phenotypes and Mutant Alleles
SGD explains that with a systematic deletion of the MRS2 gene from the genome, the resulting yeast cells are viable. Yeast cells do experience one "growth defect on a non-fermentable (respiratory) carbon source." Steinmetz explains that mitochondrial gene deletions can impair mitonchondrial respiration (Steinmetz et al 2003). Gregan noticed that the MRS2 gene can act as a suppressor to mutant yeast cells that create intron splicing defects (Gregan J et al. 2001). SGD also states that one mutation can cause a null mutant that is viable and has a pet- phenotype associated with a "block" in all group II mRNA intron splicing (2002).
YOR331C
SGD explains that YOR331C is a dubious ORF located just downstream from MSR2.
Exonic and Chromosomal Coordinates(ChrXV)
Exon: 1 - 558
Chromsome: 944115 - 943558
Information from SGD.
Sequence Information
Cellular Location of YOR331C
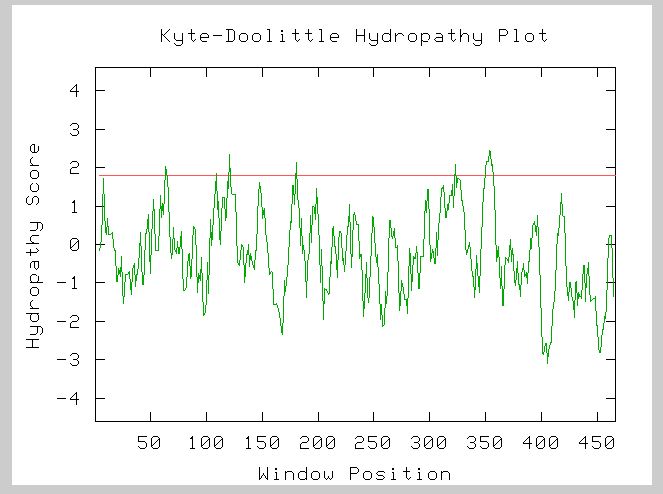
Figure 5: This is the Kyte-Doolittle Hydropathy Plot for YOR331C's amino acid sequence. There are 5 clear transmembrane regions for YOR331C. Image from Kyte-Doolittle.
YOR331C is definitely an integral membrane protein based on the Kyte-Doolittle results.
Structure
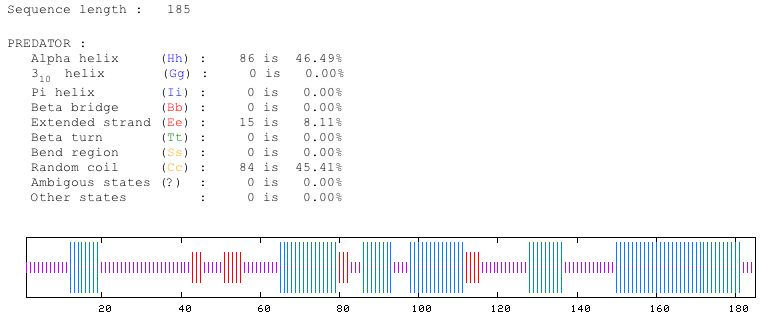
Figure 6: This shows the predicted structure of the protein sequence of YOR331C. Predator predicts that about 47 percent is alpha helices, 8 percent is extended strand and 45 percent is random coil. Image from Predator. Permission pending.
Conservation in other Sequences
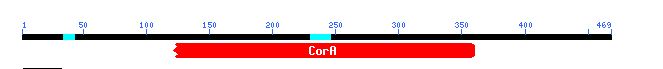
Figure 7: This is a figure from Conserved Domain that shows that some of the sequence from YOR331C is conserved in CorA, a bacterial protein. Image from CD.
Conserved Domain informs that 78.5 percent of the sequence of YOR331C is identical to CorA, a bacteria protein that transports magnesium ions in bacteria like E. coli.
I ran PSI-Blast and found that the YOR331C has homology to the polyproteins and orflabs of the SARS coronavirus, which seems very interesting(PSI-BLAST).
Conclusions
It seems very evident that YOR331C encodes a protein product whose function operates in the cell membrane of yeast. The fact that the YOR331C sequence shows similarity to CorA, a Mg+2 transporter seems very fitting since YOR331C is just slightly downstream from MSR2, a known magnesium transporter. Also, the two Predator structure predictions for the MSR2 protein sequence and the YOR331C protein sequence look very similar. Both MSR2 and YOR331C have attributed nearly the same percentages of their structure to alpha helices (38 compared to 47), extended strand (10 compared to 8 percent) and random coil (50 and 45 percent). Therefore I will conclude that YOR331C is an integral membrane protein whose function involves ion transport. More specifically, YOR331C is most likely some type of Mg+2 transporter because of the sequence similarity to CorA and the structural similarity from Predator.
References
Cherry JM, Ball C, Weng S, Juvik G, Schmidt R, Adler C, Dunn B, Dwight S, Riles L, Mortimer RK, Botstein D Nature 1997 387(6632 Suppl):67-73 Genetic and physical maps of Saccharomyces cerevisiae.
Dolinski, K., Balakrishnan, R., Christie, K. R., Costanzo, M. C., Dwight, S. S., Engel, S. R., Fisk, D. G., Hirschman, J. E., Hong, E. L., Issel-Tarver, L., Sethuraman, A., Theesfeld, C. L., Binkley, G., Lane, C., Schroeder, M., Dong, S., Weng, S., Andrada, R., Botstein, D., and Cherry, J. M. "Saccharomyces Genome Database" http://www.yeastgenome.org/ 2002http://www.yeastgenome.org 2003 Oct 7.
Juraj Gregan,1 Martin Kolisek, and Rudolf J. Schweyen. "Mitochondrial
Mg2+ homeostasis is critical for group II intron splicing in vivo." Vienna
Biocenter, Department of Microbiology and Genetics, University of Vienna,
A-1030 Vienna, Austria Vol. 15, No. 17, pp. 2229-2237, September 1, 2001
Steinmetz LM, Scharfe C, Deutschbauer AM, Mokranjac D, Herman ZS, Jones T,
Chu AM, Giaever G, Prokisch H, Oefner PJ, Davis RW. "Systematic screen
for human disease genes in yeast." Department of Genetics, Stanford University
School of Medicine, Stanford, California 94305, USA. larsms@stanford.edu Nat
Genet. 2002 Aug;31(4):400-4. Epub 2002 Jul 22.