This web page was produced as an assignment for an undergraduate course at Davidson College
Comparing a popular publication and
scientific journal's article about ASPM
Background:
On January 13, 2004, Patrick Evans, Jeffrey Anderson, Eric Vallender, Sandra Gilbert, Christine Malcom, Steve Dorus, and Bruce Lahn published their paper in Human Molecular Genetics entitled "Adaptive Evolution of ASPM, a major determinant of cerebral cortical size in humans". The New York Times reported on January 14, 2004, that Dr. Bruce Lahn and colleagues at the University of Chicago decoded the ASPM gene, a gene thought to be one of several implicated in brain size.
According to Evans' et al (2004, Adaptive), five loci had been linked to primary microcephaly, a congenital condition in which there are reductions in the cerebral cortex size of the brain, and the head circumference is below average size. A person with microcephaly also may have problems with motor skills, speech, and hyperactivity, have convulsions, mental retardation, and facial anomalies associated with small head size. One of the five loci was identified through linkage as ASPM (Abnormal spindle-like microcephaly associated). ASPM has also been identified in a number of other organisms: several apes and other primates, dogs, cats, sheep, cows, mice, rats, and flies. Although the biochemical function of the ASPM protein is not known in mammals, there is data on the homologous gene in Drosophila, asp. The asp protein has been shown to be a player in the organization of mitotic and meiotic spindle structures of flies, so it is thought that the ASPM protein has a similar role in mammals. Because disruption of ASPM causes primary microcephaly, Evans et al. believes that alterations to the amino acid sequence of ASPM could affect the regulation of mitotic spindle orientation in brain cells, and that this alters the size of the cerebral cortex.
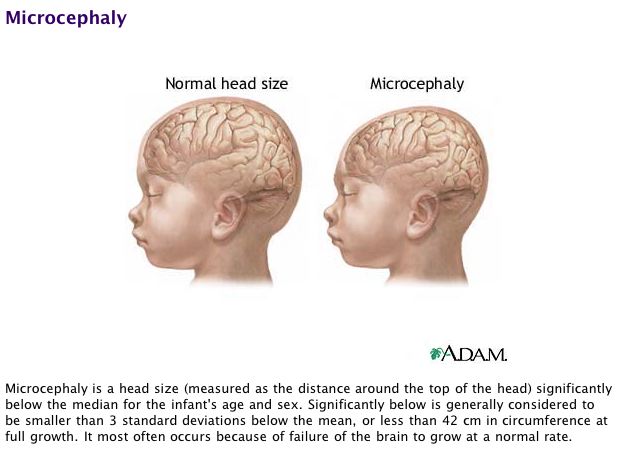
MedlinePlus
The Popular Article:
The New York Times reported that the ASPM gene has been under selective pressure. ASPM, according to the article, was identified because of its role in causing microcephaly, a disorder in which people are born with abnormally small cerebral cortexes. Over the last 18 million years, there has been a progressive change in ASPM - 15 changes to be exact - which has been correlated to a steady increase in the size of the cerebral cortex through the evolution of the great apes, a group of organisms that includes orangutans, gorillas, chimps, and humans. Conversely, there seems to have been no selective pressure on the same gene in monkeys, dogs, cats, and cows.
Dr. Bruce Lahn, a geneticist involved with the study, was contacted and quoted in the article: " 'There has been a sweep every 300,000 to 400,000 years, with the last sweep occurring between 200,000 and 500,000 years ago' ... referring to a genetic change so advantageous that is sweeps through a population endowing everyone with the same improved version of a gene" (Wade 2004). According to Dr. Jianzhi Zhang, an evolutionary biologist not involved with Evans' study, says that the gene seems to have been kept stable since the last sweep, resisting any changes in the protein product, a process called "purifying selection"(Wade 2004). Dr. Zhang also reports that the brains of humans with microcephaly are about four and a half times smaller in mass than normal brain masses, a size comparable to early hominids that lived three million years ago, suggesting that microcephaly negates about three million years of human development.
The Times also contacted Dr. Christopher Walsh, a neurologist that identified ASPM gene mutations as a cause of microcephaly. He is quoted saying: "The ASPM gene is 'almost unique' ... because in all known animal genomes, it has resisted the usual duplication events and been maintained as a single copy" (Wade 2004).
Five other genes, have also been found to cause microcephaly when mutated, the article continues, so ASPM is not the only one implicated in brain size. It states that the function of the gene is to determine the number of cells in the cerebral cortex of developing fetuses.
The Scientific Article:
Evans' et al. paper focuses on the evolution of ASPM in the primate lineage. In the introduction, it states that "genes involved in the development of the brain are likely to also play a role in the evolution of the brain. However, this proposition is yet to be validated by empirical data" (Evans et al. 2004, Adaptive). From the beginning, Evans et al. admits that ASPM can only be implicated in the evolution of brain size, and that the research findings are "consistent with a possible role of ASPM in the evolutionary enlargement of the human cerebral cortex" (Evans et al. 2004, Adaptive). In the discussion, Evans et al. speculates the function of the ASPM protein, because there is no data about the biological function. Using the Drosophila homologous gene, the function of ASPM is thought to regulate mitotic spindle orientation in dividing cells in the cerebral cortex, thus controlling or altering the size of the cortex.
The evolution of ASPM was calculated using the 10.4kb coding region of ASPM in primates that represented a phylogenetic tree of evolution, including New World Monkeys, Old World Monkeys, Lesser Apes, and Greater Apes. The ASPM Protein Sequence in humans can be found here. The coding region of each species used was compared to its neighbor on the tree using a ratio of non-synonymous amino acid substitution rates to synonymous amino acid substitution rates, taking into account the natural mutation rate. Higher ratios (greater than one) indicate a positive selection for the mutated gene because non-synonymous substitution rates are higher than synonymous substitution rates. The lineage moving towards humans has the highest ratio and the highest mutation rate, indicating that there was positive selection on ASPM as evolution progressed toward humans. The ASPM mutation rate was also compared to other organisms, such as domestic cats, domestic dogs, cows, sheep, mice, rats, and worms. Evans et al. noted that "the evolutionary rate of the lineages leading to humans is significantly higher" than compared to the other species, suggesting that the "ASPM protein evolution is a distinct feature of the ape lineages leading to humans and is not found in either non-hominoid primates or other non-primate mammalian orders" (Evans et al. 2004, Adaptive). A Tree of Organisms Used in the research is available.
Comparison:
The popular press article is very accurate in what it choose to include about Evans' et al. article. Although simplified, the information provided gives a general overview of the results of the research project. Significantly, none of the results reported in the Times article are exaggerated any more than the reported results from the scientific paper. Unlike some other mainstream reports of "discovered genes," this popular press article notes that ASPM is only one of five other unidentified genes that determines human brain size, and not the single gene factor in this trait. However, the Times failed to mention that the function of the ASPM protein is not exactly known in humans, and that its suggested function is derived from the homologous gene found in Drosophila.
As mentioned before, the scientific article focuses on the research done on ASPM, and the evolutionary results that are inferred from the data. The popular press article, on the other hand, focuses more on microcephaly and brain size as it has evolved over the last 18 million years. Most of the information about brain size and microcephaly as it relates to ASPM was taken from sources other than Evans et al. These sources appear to be legitimate, as Dr. Zhang's input was taken from a report published the previous month in Genetics, and Dr. Walsh was contacted via e-mail.
A major criticism of the popular press article is that it failed to mention any of the methods, statistical tests, or even specific data of the research. This omission may have occurred because of the complex nature of the statistical tests and results. However, the methods used to measure the changes of ASPM may be just as important as the results themselves. The ratios used to measure the changes, and the McDonald-Kreitman statistical test which measures positive selection, can be applied to other common genes in human's evolutionary lineage.
Another criticism is that the Times' quote from Dr. Lahn was not contained in the paper. The scientific paper states: "roughly 15 of the 19 non-synonymous substitutions that occured between the last human/chimpanzee ancestor and humans may have been adaptive and were driven to fixation by positive selection. This translates to about one adaptive amino acid change in ASPM for every 300 000-400 000 years since the human lineage diverged from chimanzees" (Evans et al. 2004 Adaptive). This excerpt has a different meaning than the Times quote, which suggests that a gene change swept through every individual, endowing them with a new, advantageous mutation. The Times did not specify if Dr. Lahn was contacted, or if they were quoting the paper.
Looking Ahead:
Evans and colleagues were at it again, publishing another paper in Human Molecular Genetics just two and a half months later. They identified microcephalin, one of the other six genes now known to be associated with primary microcephaly. Microcephalin also experienced selective pressures, but earlier in the human lineage of humans (Evans et al. 2004, Reconstructing). More research on microcephalin suggests that the protein regulates BRCA1 (a gene known to be associated with hereditary breast cancer) and/or Chk1, which are involved in DNA damage-induced cellular responses (Xu et al. 2004).
References:
Evans PD, Anderson JR, Vallender EJ, Choi SS, and Lahn BT. 2004. Reconstruction the evolutionary history of microcephalin, a gene controlling human brain size. Human Molecular Genetics 13: 1139-1145.
Evans PD, Anderson JR, Vallender EJ, Gilbert SL, Malcom CM, Dorus S, and Lahn BT. 2004. Adaptive evolution of ASPM, a major determinant of cerebral cortical size in humans. Human Molecular Genetics 13: 489-94.
Kua E, Reder M, Grossel MJ. 2004. Science in the news: a study of reporting genomics. Public Understanding of Science. 13 (3). <http://www.sagepublications.com>. Accessed 2004 Aug 27.
MedlinePlus. 2004 May 8. Medical Encyclopedia, Microcephaly. <http://www.nlm.nih.gov/medlineplus/ency/imagepages/17256.htm> Accessed 2004 Sep 14.
National Institute of Neurological Disorders and Stroke [NINDS]. 2001 Jul 01. NINDS Microcephaly Information Page. <http://www.ninds.nih.gov/health_and_medical/disorders/microcephaly.htm> Accessed 2004 Sep 14.
Wade, Nicholas. 2004 Jan 14. Evolution of Gene Related to Brain's Growth Is Detailed. New York Times. <http://gateway.proquest.com/openurl?url_ver=39.88-2004&res_dat=xri:pqd&rft_val_fmt=info:ofi/fmt:kev:mtx:journl&genre=article&rft_dat=xri:pqd:did=000> Accessed 2004 Aug 30.
Xu X, Lee J, and Stern DF. 2004 Aug 13. Microcephalin is a DNA damage response protein involved in regulation of CHK1 and BRCA1 [abstract]. In J Biol Chem 279(33):34091-4. PubMed database <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=Abstract&list_uids=15220350>. Accessed 2004 Sep 15.
Megan's Home
Genomics Home
Davidson Home
Contact Megan at mecastle@davidson.edu
