
This web page was produced as an assignment for an undergraduate course at Davidson College.
My Favorite Yeast Proteins: SAE2p and YGL176Cp
Introduction
In my previous webpages, I have investigated the function of yeast genes SAE2 and YGL176C using published literature and various internet sites. First, I explored the ontology of these genes in My Favorite Yeast Gene webpage. Later, I examined their expression profiles in My Favorite Yeast Expression webpage. Indeed, I have mined existing resources on the web to confirm the annotation of SAE2 and employed similar techniques to figure out the previously unidentified function of the non-annotated gene, YGL176C. In this final assignment, I will analyze SAE2p and YGL176Cp using several protein databases and data sets. I will also develop ideas for future experiments to test my predictions.
SAE2
Recall: SAE2 is yeast gene located on the Crick-strand of Chromosome VII between coordinates 174328 and 173291. Although the gene does not have a known molecular function, it encodes the 345 amino acid protein SAE2p. SAE2p is a meiotic and mitotic DNA-repair enzyme localized to the nucleus and cytoplasm of yeast cells. Microarray expression data generally confirm the ontology of SAE2.
Unfortunately, the structure of SAE2p has not yet been crystallized and deposited into the Protein Data Bank. The lack of a crystallized structure of SAE2 is not surprising because merely 20,000-30,000 proteins have been crystallized to date (Campbell AM, 2005).
When I searched for SAE2, I found that it was not present on the 2D database (Fig.1). Indeed, SAE2 was not present in the extract characterized in by this yeast 2D gel in this database.

Figure 1. Search results for SAE2 in the 2D Database show no hits. (Sanchez JC, et al., 1996; <http://ca.expasy.org/cgi-bin/ch2d-search-ful?makeWild=&SEARCH=SAE2>).
| SAE2p is not a transmembrane protein. Because it did not resolve on the 2D gel, perhaps SAE2p is a small molecule or a fairly basic molecule. |
Using the Expasy Server, I found the theoretical isoelectric point and molecular weight of SAE2 to test my hypothesis. The molecular weight of SAE2 and isoelectric point of SAE2 are 40.09702 kDa and 6.17, respectively (Fig 2).
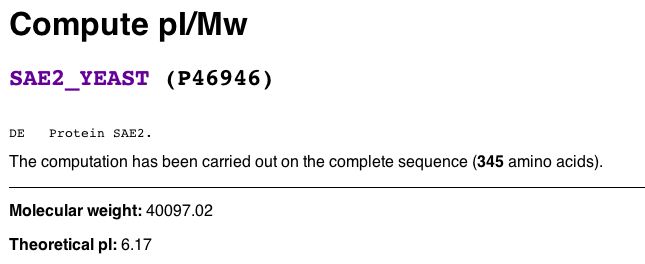

Figure 2. Calculated Molecular weight and theoretical isoelectric point of SAE2. (Gasteiger E, et al., 2003; <http://www.expasy.org/cgi-bin/pi_tool1?P46946@noft@>).
| SAE2p cannot be considered a basic protein because its isoelectric point is 6.17 (it would have to have an pI>> 7 to be very basic). SAE2 is also not a very small molecule, given its MW is approximately 40 kDa. Thus, SAE2p did not resolve in the 2D gel for some other reason. Regardless, find the approximate region in Fig 3 below where you think SAE2p would have appeared on the 2D gel. If you guess correctly using Internet Explorer web browser, a blue box will appear. |
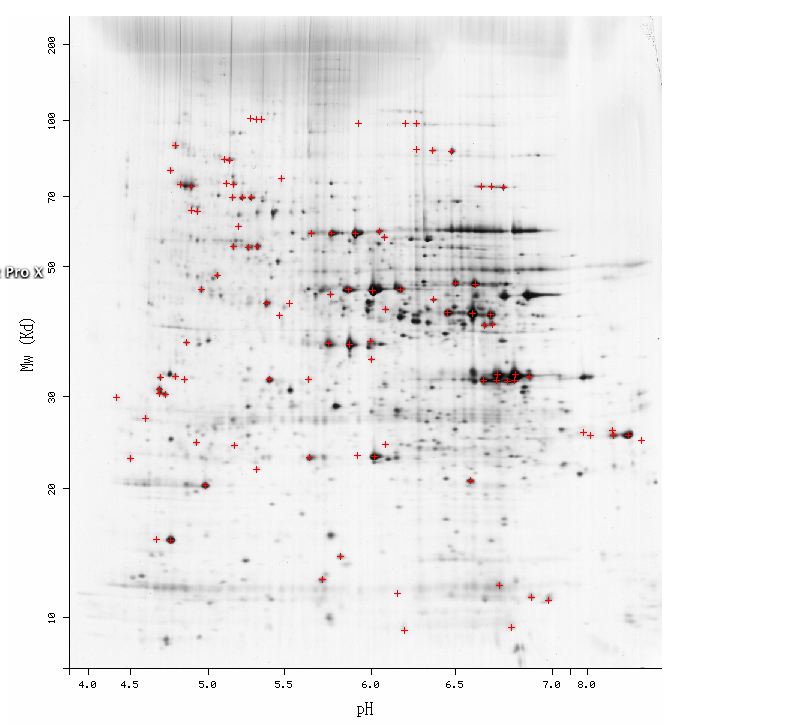
Figure 3. Screen shot of the yeast 2D gel to analyze proteins. Note the location where SAE2 would have appeared can be approximated given the calculated MW and theoretical pI. (Sanchez JC, et al., 1996; <http://ca.expasy.org/cgi-bin/map2/def?YEAST>).
A team from Yale University created single gene yeast knockouts with specially engineered transposons. By mining their TRIPLES database, one can investigate the expression, transcription location of the knocked out genes, and the resulting phenotype of the knockout mutants. I found that SAE2 is knocked-out using a transposon that inserts at codon 303 (Fig 4).
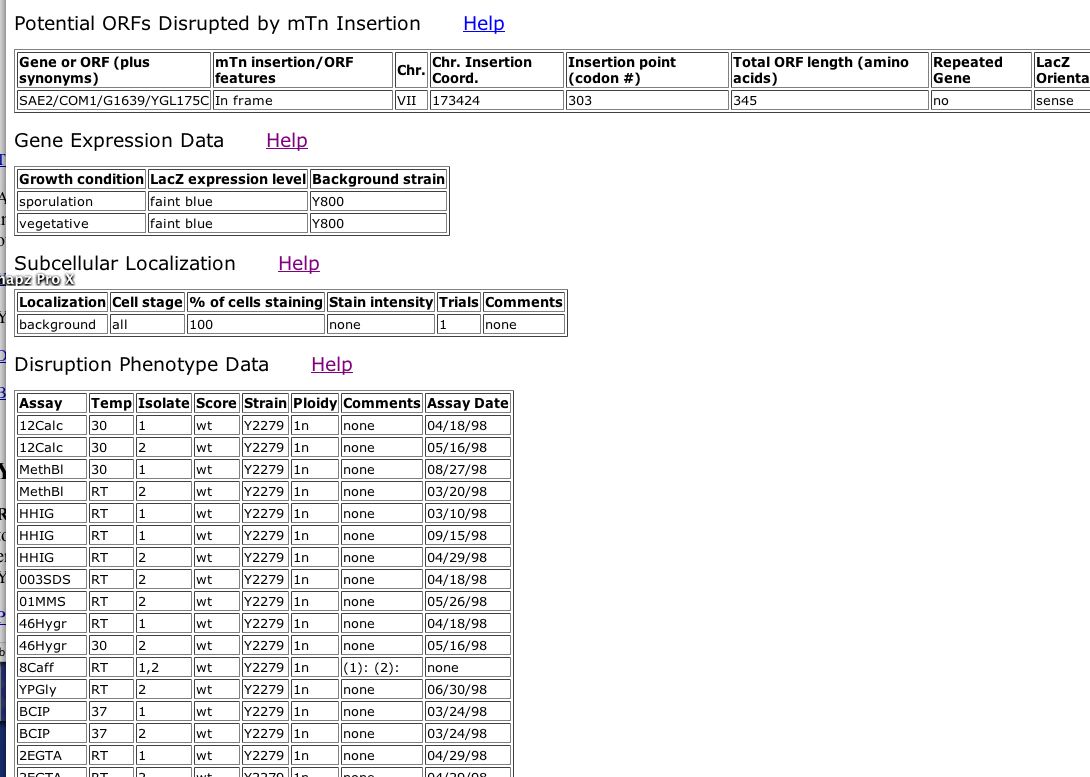
Figure 4. Screen Shot of data from TRIPLES database for SAE2. (Kumar A, et al., 2000; <http://ygac.med.yale.edu/triples/get_clone_info.asp?cloneid=V46G1>).
| It appears that SAE2 is transcribed a small amount in both the sporulation and vegetative phase. It makes sense that SAE2p would be transcribed both during reproductive and normal growth/stationary phase because it is meiotic and mitotic DNA repair enzyme. The location for SAE2p was not determined by this group because localization could not be distinguished from the background. Finally, under most conditions, disrupted SAE2 did not confer a distinct phenotype. However, SAE2 knockout exhibits a "strong" change in growth under the condition of YPD + 0.003% SDS. SDS is a detergent, which when present probably denatures proteins (as in SDS-PAGE) (Fig 4). Indeed, SDS probably disrupts overall functioning of yeast cells because proteins would be denatured. If SAE2 also has a role in protein folding, SDS treatment would be particularly detrimental to SAE2 knockouts. |
![]()
Figure 4. Screen shot illustrating the strong phenotype that occurs in SAE2 knockout yeast under YPD + 0.003% SDS treatment. (Kumar A, et al., 2000; <http://ygac.med.yale.edu/triples/get_clone_info.asp?cloneid=V46G1>).
By the Yeast Two-Hybrid Method, SAE2 is shown to interact with SAE2 and YCR086W (Fig 5).

Figure 5. Screen shot illustrating the tabel of interactions of SAE2 with SAE2 and YCR086. The left column represents the "Bait," while the right column represents the "prey" in the Y2H experiment. (Uetz P, et al., 2000; <http://depts.washington.edu/%7Eyeastrc/th_11.htm>).
| The Y2H database provides interesting information if not a great quantity. It confirms the function of SAE2p. SAE2 interacts with YCR086W, which according to the MIPS, is involved in "nuclear migration" and DNA processing. SAE2p is a DNA-repair enzyme so it makes sense that it would interact with another protein implicated in DNA processing. Interestingly, SAE2p appears to interact with SAE2p (itself). Perhaps SAE2p forms a dimer in vivo. |
In the DIP Database compiled by David Eisenberg and his colleagues at UCLA, SAE2p appears to interact directly with three proteins: Srp1p, Kap95p, and Csm1p (Fig 6).
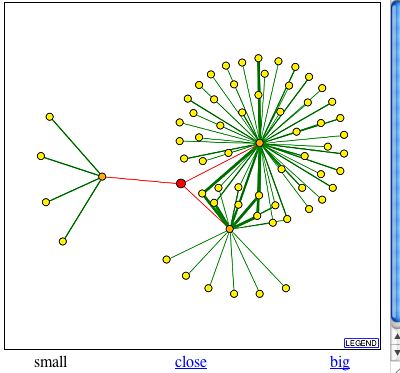
Figure 6. Screen shot of SAE2p protein interactions as in DIP database. SAE2p appears as the red dot in the center of the figure and proteins that directly interact with SAE2p appear as orange dots. (Eisenberg D, 1999-2004; <http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=1603>).
| The proteins with which SAE2p interacts according to the DIP database have related ontologies to SAE2p. For example, Srp1p is localized to the cytoplasm and nucleus, just like SAE2p. Srp1p is involved with nucleocytoplasmic transport and protein carrier activity. Perhaps Srp1p helps transport SAE2p to DNA break repairs sites or interacts with it in some other way. Likewise, Kap95p is localized to the cytoplasm and the nuclear pore. This protein plays a role in protein-nucleus import and protein carrier activity, as well. Finally, Csm1p is understandably very directly involved with SAE2p because it is the same protein, YCR086w, detected in Y2H experiments and explained above. |
SAE2p was again shown to interact with YCR086Wp in the circuit diagram produced by Schwikowski B, et al. (2000) (Fig 7).
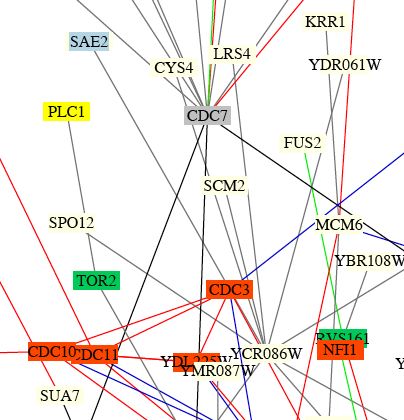
Figure 7. Screen shot of small portion of integrated circuit diagram showing protein-protein interaction in yeast. Note SAE2 near the top of the figure in a blue box, indicating that the protein SAE2p is involved in membrane fusion. SAE2 is connected by a gray line to YCR086C, indicating that the two proteins encoded by their gene interact and that the localization or cellular roles are unknown for one or both proteins. (Schwikowski B, et al, 2000; <http://occawlonline.pearsoned.com/bookbind/pubbooks/ bc_mcampbell_genomics_1/medialib/seq/Benno/NB_Figure1color.pdf>).
| It is not surprising that SAE2p interacts with YCR086w because we have seen it come up before in the Y2H data and the DIP database. This figure must be dated, because the function and location of both proteins SAE2p and YCR086p have been determined. It seems strange to me that SAE2 is categorized as involved in membrane fusion, as it encodes a DNA-repair enzyme. Perhaps SAE2p has alternative functions and/or the function "membrane fusion" fits it best in this case (as opposed to chromatin structure, cell structure, lipid metabolism, cytokinesis, or unknown role). |
I could not locate SAE2 (aka YGL175C) in the KEGG Pathway website (Fig 8). I cannot ascertain information about pathways in which SAE2 is involved.

Figure 8. Screen shot demonstrating result when the KEGG database is searched for SAE2. (Kanehisa M, et al., 2002; <http://www.genome.jp/dbget-bin/www _bfind_sub?dbkey=pathway&keywords=SAE2&mode=bfind&max_hit=1000>).
| Conclusions about SAE2p: The proteomics databases generally confirm the annotaion of SAE2p. |
YGL176C
Recall: YGL176C is the non-annotated genomic "neighbor" of SAE2. It is also located on the Crick-strand of Chromosome VII, but it spans coordinates 173085 to 171421.YGL176C currently has no known molecular function, biological process, or cellular component. In the first assignment, I predicted that this gene encoded the 554 amino acid protein YGL176Cp that is receptor and/or involved in protein folding. Using the microarray expression data in the second assignment, however, I hypothesized that YGL176Cp is involved in maintaining cellular homeostasis by transporting molecule across membranes.
Unfortunately, YGL176Cp has not yet been crystallized, so there is not an image to represent the 3-D structure of YGL176Cp.
YGL176C did not appear in the yeast 2D gel. This is as to be expected because YGL176C has several transmembrane domains and therefore would be difficult to solubilize. However, using the Expasy Server, I found the theoretical isoelectric point and molecular weight of YGL176C to be 63.27600 kDa and 8.60, respectively (Fig 9).
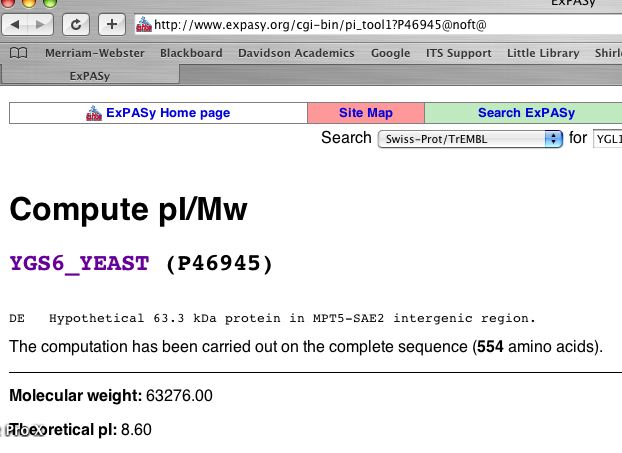
Figure 9. Calculated Molecular weight and theoretical isoelectric point of YGL176Cp. (Gasteiger E, et al., 2003; <http://www.expasy.org/cgi-bin/ pi_tool1 ?P46945@noft@>).
| Given the isoelectric point is 8.6, YGL176Cp is mildly basic. The basic pI provides another explanation as to why the protein did not resolve on the yeast 2D gel. Regardless, using the calculated molecular weight and theoretical pI, find the approximate region in Fig 10 where you think YGL176Cp would have appeared on the 2D gel. If you guess correctly using Internet Explorer webbrower, a blue box will appear. |

Figure 10. Screen shot of the yeast 2D gel to analyze proteins. Note that YGL176C would appear in the region of the blue box if it had been isolated in the experiment. (Sanchez JC, et al., 1996; <http://ca.expasy.org/cgi-bin/map2/def?YEAST>).
| We did not gain much information about YGL176C using 2D gel data. YGL176C did not appear on the gel. Even if it did, we cannot assume homology between proteins that share the same pI and MW. Two proteins could have similar pI and MW, but not share any common function. |
YGL176C appears in the PROWL database. This site could potentially provide new information about YGL176C, but the links are not functional at this time for some reason (Fig. 11).

Figure 11. Screen shots of hits when searching for YGL176C on the PROWL database. (ProteoMetrics, LLC, 1997-2000; <http://129.85.19.192/prowl-cgi/ReadSequence.exe?name=db|nr-Saccharomyces-cerevisiae|gi|6321262|ref|NP_011339.1>).
It appears that a transposon inserted into YGL176C at codon 498, creating a knockout for this gene. According to the data in Fig 12 below, YGL176C is expressed both during sporulative and vegetative conditions. Thus YGL176C is expressed during normal growth and reproductive conditions. YGL176C is not specifically expressed under one condition. Howev,er the location of transcription of YGL176C could not be determined by this dataset (the localization could not be distinguished from the background). Finally, the YGL176C knockout yeast did not show any abnormal phenotype under any conditions that the team at Yale surveyed. Other proteins probably compensate for the lack of YGL176Cp because it is encoded by a non-essential gene.
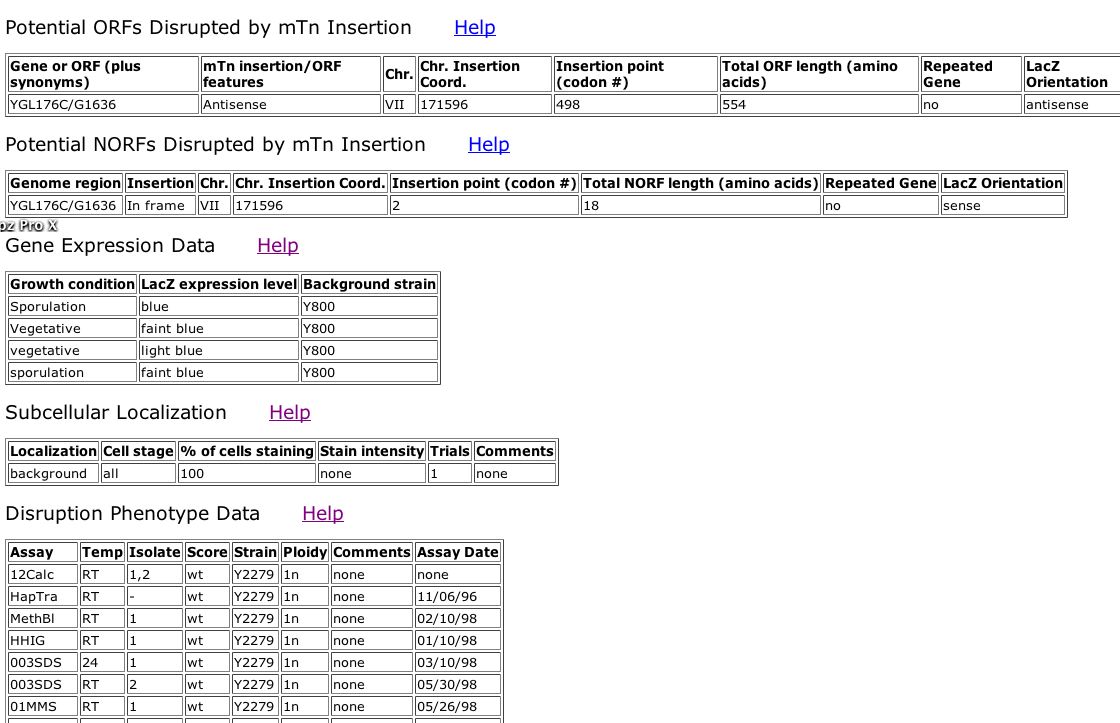
Figure 12. Screen Shot of data from TRIPLES database for YGL176C. (Kumar A, et al., 2000; <http://ygac.med.yale.edu/triples/get_clone_ info.asp?cloneid= V15D11>).
| We did not gain any novel information regarding YGL176Cp from the TRIPLES database. However, perhaps YGL176Cp does not play an essential role in protein folding because there is no disruption phenotype shown when the YGL176C knockout yeast are grown in YDP + 003% SDS, as in the case of knockout SAE2 yeast. |
Y2H, Additional Y2H Results, or Full Database
YGL176Cp does not appear in the Y2H data set. We cannot determine with what proteins YGL176Cp interacts using this information.
YGL176Cp appears in the DIP database, but there is not an accessible graph about its interactions (Fig 13). We cannot gain any new information about YGL176Cp with the DIP database.
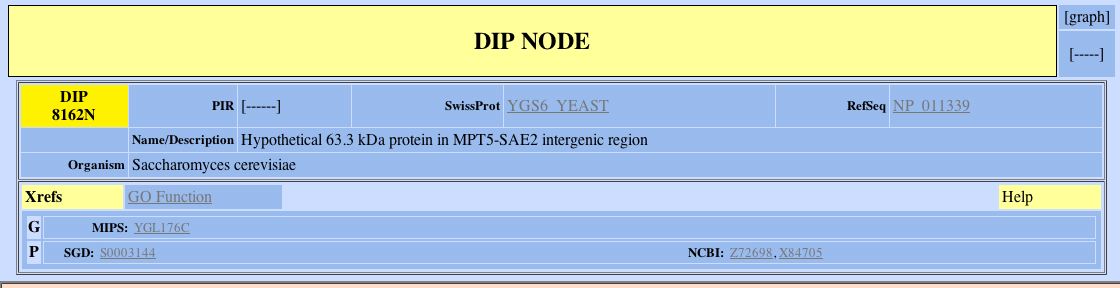

Figure 13. Information about YGL176Cp from the DIP database. Note that the the word [graph] is not underlined in the top right corner. This means there is no link to a graph depicting the protein-protein interactions of YGL176Cp. (Eisenberg D, 1999-2004; <http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=8162>).
Benno Figure 1, Benno Aging pdf, Benno Membrane pdf, Benno Degradation pdf
YGL176C does not appear in the Schwikowski B, et al. (2000) data sets. We cannot gain any information regarding its interactions with other proteins by this route.
YGL176Cp does not appear in the KEGG pathway database. We cannot gain any insight about which pathways this protein is involved via this site.
| Conclusions about YGL176C: We did not gain any new information about YGL176Cp from proteomics databases and data sets. Generally, YGL176Cp did not appear in the databases. When it did, however, the databases provided minimal information about its function or they were malfunctioning. Thus, I must return to the hypothesis I made after the last assignment. I predict that YGL176Cp is a transmembrane transport molecule that may have a role in protein folding. |
| Future Experiments:
Indeed, many more experiments need to be conducted to narrow down or determine the function of YGL176Cp. First of all, the protein-protein interaction of YGL176Cp should be examined more definately. Using the Y2H method, specifically with YGL176Cp as the bait, one could begin to estimate the function of this protein by identifying proteins act as its "prey". If YGL176Cp has a role in protein folding, it should widely interact with one or more proteins. If it is a receptor, it should have a ligand or multiple ligands. Finally, if YGL176Cp is a transport molecule it should interact with molecule(s) that opens and/or closes the channel (perhaps one particular protein or ATP). Other experiments I would perform would involve determining the cellular location of YGL176Cp. The TRIPLES database using the transposon/epitope tag method with immunoflorescence did not provide any useful information about where YGL176C is expressed. It would be useful to determine where and when YGL176Cp is made. Therefore, I would attempt to find the YGL176Cp by repeating the transposon/epitope tag method and hopefully create a different insertion in YGL176C that would lead to more fruitful expression studies. Alternatively, I would determine the location of YGL176Cp by inserting the gene for GFP after YGL176C in a the DNA of a yeast cell. I could follow YGL176C using the attached glowing protein as a guide. If YGL176C encoded a a transmembrane molecule, it would likely be expressed in the cytoplasm and transported to a membrane. Eventually the GFP protein attached to a transmembrane protein would cause the membrane to illuminate in a characteristic pattern. Furthermore, although it would be difficult if it were a transmembrane protein, I would attempt to crystallized YGL176Cp. It is important to understand the function of YGL176Cp by examining the actual 3D structure of the protein. Using a chime image of the 3D structure of YGL176Cp, I could explore whether my proposed functions for the protein are complimentary to the actual shape of the molecule. Alternatively, perhaps chunks of the membrane could be crystallized and scanned for proteins, namely YGL176Cp. Finally, I would create a knockout of YGL176C and test it under high throughput, more stringent and/or different conditions than the TRIPLES group in order to gain a disruption phenotype. There must be an essential function of YGL176C that helps it under some condition, otherwise it would have been eliminated by genetic drift and evolution. Thus, the YGL176C knockout should show distinct phenotype under some condition. By figuring out which treatments affect YGL176C knockouts, I would be able to have a better idea as to the function of YGL176Cp. |
References
Campbell, AM. 2005. 2005. "Standards and Assignments for Student Web Pages in Genomics, Proteomics and Systems Biology." Genomics, Proteomics, and Systems Biology Website Fall 2005. <http://bio.davidson.edu/courses/genomics/GPBwebstandards.html#web4>. Accessed 2005 Nov 17.
Eisenberg D. 1999-2004. Database of Interacting Proteins. <http://dip.doe-mbi.ucla.edu/dip/Search.cgi?SM=3>. Accessed 2005 Nov 17.
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A. 2003. ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31:3784-3788. <http://www.expasy.org>. Accessed 2005 Nov 17.
Kanehisa M, Goto S, Kawashima S, Nakaya A. 2002. The KEGG databases at GenomeNet. Nucleic Acids Res. 30, 42-46. <http://www.genome.jp/kegg>. Accessed 2005 Nov 17.
Kumar A, Cheung KH, Ross-Macdonald P, Coelho PSR, Miller P, Snyder M. 2000. TRIPLES: a Database of Gene Function in S. cerevisiae. Nucleic Acids Res. 28, 81-84. <http://ygac.med.yale.edu/triples/triples.htm>. Accessed 2005 Nov 17.
ProteoMetrics, LLC. 1997-2000. ProteinInfo. <http://prowl.rockefeller.edu/prowl/proteininfo.html>. Accessed 2005 Nov 17.
Sanchez JC, Golaz O, Frutiger S, Schaller D, Appel RD, Bairoch A, Hughes GJ, Hochstrasser DF. 1996. The yeast SWISS-2DPAGE database. Electrophoresis 17: 556-565. <http://www.expasy.org/ch2d/>. Accessed 2005 Nov 17.
Schwikowski B, Uetz P, Fields S. 2000. A network of protein-protein interactions in yeast. Nature Biotechnology 18: 1257-1261. <http://occawlonline.pearsoned.com/bookbind/pubbooks/ bc_mcampbell_genomics_1/medialib/seq/Benno/NB_Figure1color.pdf>. Accessed 2005 Nov 17.
Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, Knight JR, Lockshon D, Narayan V, Srinivasan M, Pochart P, Qureshi-Emili A, Li Y, Godwin B, Conover D, Kalbfleisch T, Vijayadamodar G, Yang M, Johnston M, Fields S, Rothberg JM. 2000. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403: 623-627. <http://depts.washington.edu/%7Eyeastrc/th_11.htm>. Accessed 2005 Nov 17.
Return to Jackie's Genomics Main Page