Abstract
Mutations in MSH6, a DNA repair gene, are associated with a predisposition to the development of colorectal cancer. The MSH6 gene, which forms a heterodimer with MSH2 in vivo, is quite well characterized, with multiple conserved domains across many species. Within the MSH6 gene, there are a large number of alpha helices, believed to play a role in DNA binding, and several upstream motifs that function in DNA binding as well, underscoring the importance of the ability of MSH6 to bind to aberrant DNA. The control of MSH6, however, is not as well understood. An assessment of yeast MSH6 and MSH2 transcription in response to MSN deletions revealed similar patterns of expression, suggesting the potential coregulation of MSH2 and MSH6. Further review of genes associated with transcription revealed that NRM1 acted similarly to MSH2 and MSH6 in response to MSN deletions and several other factors, suggesting that MSH2, MSH6, and NRM1 may all have some common coregulator. Within the MSH6 gene, multiple SNPs have been characterized, some of which are suspected to give rise to the colorectal cancer phenotype. One such mutation, found within an alpha helix of a suspected DNA binding region, is likely to impede upon the ability of the MSH2-MSH6 heterodimer to bind aberrant DNA. If mutated MSH6 is indeed unable to bind to aberrant DNA, then one potential treatment for persons predisposed to colorectal cancer would be to place a WT copy of MSH6 into all their cells, enabling formation of a functional MSH2-MSH6 heterodimer. While much progress has been made in the study of MSH6, one remaining mystery is why MSH6 mutations are associated with the development of colorectal cancer and not cancer in general. Analysis of colon-specific factors should help elucidate the mechanisms behind the specificity of MSH6 and colon cancer.
Background on MSH6 and Colorectal Cancer
The development of Colon Cancer [Figure 1], the fourth most common cancer in the United States, is known to have a strong genetic component [Colon Cancer; MedlinePlus: Colorectal Cancer]. There were an estimated 93,800 cases of Colon Cancer diagnosed in 2000—and 47,700 of those are likely to have resulted in death [Colon Cancer]. While colon cancer is quite treatable if caught early, people often have no symptoms until the disease has progressed quite far, underscoring the importance of routine screening [MedlinePlus: Colorectal Cancer].
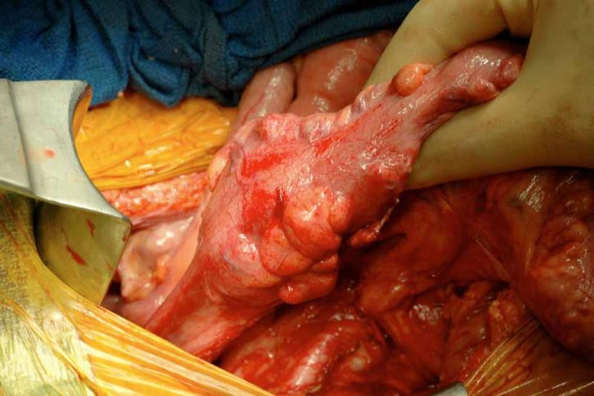
Figure 1 [http://www.jyi.org/articleimages/1070/originals/img0.jpg]
Mutations in the MSH6 gene have been implicated in the predisposition of an individual to develop colorectal cancer, though it is unclear why the colon seems to be preferentially affected by alterations in MSH6 activity [Colon Cancer]. MSH6 is located on chromosome 2 and associates with the protein of the MSH2 gene to form a heterodimer that plays a role in DNA repair during mitosis [Ensembl Genome Browser; Kariola 2002; OMIM]. Several possible mechanism for aberrant DNA repair in the presence of an MSH6 mutation have been suggested, including decreased stability of the MSH2-MSH6 heterodimer and an inability to bind to mismatched DNA [Kariola 2002; Bowers 1999].
References
1. Bowers J, Sokolsky T, Quach T, Alani E: A mutation in the MSH6 subunit of the Saccharomyces cerevisiae MSH2-MSH6 complex disrupts mismatch recognition. Journal of Biol Chem 1999, 274: 16115-16125.
2. Colon Cancer [http://www.ncbi.nlm.nih.gov/books/bv.fcgi?call=bv.View..ShowSection&rid=gnd.section.102]
3. Ensembl Genome Browser [http://www.ensembl.org/index.html]
4. Kariola R, Raevaara T, Lönnqvist KE, Nyström-Lahti M: Functional analysis of MSH6 mutations linked to kindreds with putative hereditary non-polyposis colorectal cancer syndrome. Human Mol Gen 2002, 11: 1303-1310.
5. MedlinePlus: Colorectal Cancer [ http://www.nlm.nih.gov/medlineplus/colorectalcancer.html]
6. Online Mendelian Inheritance in Men (OMIM) [http://www.ncbi.nlm.nih.gov/sites/entrez?db=OMIM]
The MSH6 Gene
The MSH6 gene, located on chromosome 2 in humans, is quite heterogeneous and has been identified in numerous and diverse species, from yeast to fruitfly to zebrafish to mouse to chimpanzee [BLAST; Ensembl]. Not only is the gene found in multiple species, it is also quite well conserved among them [Figure 1] [Ensembl; EBI Tools: ClustalW]. Human and yeast MSH6 show the lowest level of conservation at the amino acid level (28%) while human and chimpanzee show the highest conservation (99%) [EBI Tools: Clustal W].

Figure 1
Yet, in spite of the relatively low conservation between the human and yeast MSH6 genes, their molecular function, cellular component, and biological process are identical [NCBI; SGD]:
- Molecular Function
- ATP Binding
- ATPase Activity
- Guanine/Thymine mispair binding
- Single Base insertion/deletion binding
- ATP Binding
- Cellular Component
- Nucleus
- Biological Process
- Mismatch repair
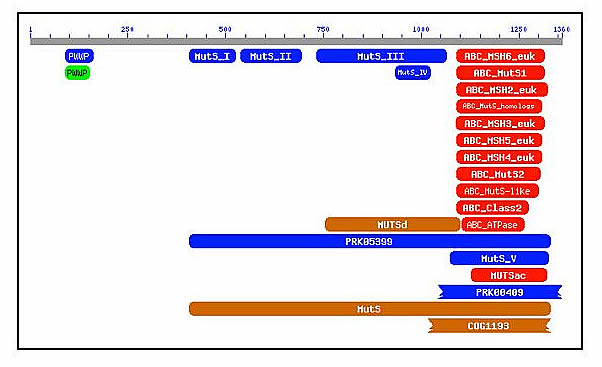
The high conservation of the biological properties of MSH6 among so many species suggests that its function is particularly crucial in survival. While the entire MSH6 gene is fairly well preserved in most species, there are several regions within the gene that are particularly similar across species. The region from approximately amino acid 1000 to amino acid 1360 is very well conserved [Figure 2], with orthologs in multiple species as well as high conservation with other MSH sequences (ie MSH2, MSH3, etc) with e-values from 2 e-84 to 9 e-9 [NCBI Conserved Domains]. Since this region of the MSH6 gene is very well conserved, it suggests that it is particularly important in the proper functioning of the gene in DNA repair.
Figure 2 [Click to Enlarge]
Nevertheless, there are several other regions of the MSH6 gene that are quite interesting. In the 500 base pairs upstream from the start of MSH6 transcription, motif finding with JASPAR revealed several motifs at a threshold of 90% [JASPAR]:
- SPI1
- Functions in DNA binding
- ZNF42_1-4
- Consensus Sequence for DNA binding in zinc fingers
- GATA3
- Consensus Sequence for DNA binding
The importance of the ability of the MSH6 gene to bind to DNA is underscored by three consensus sequences, all upstream of the MSH6 gene, that play a role in DNA binding. Interestingly, in order to identify a potential TATA box, the threshold had to be lowered to 65%, suggesting that MSH6 may use some other mechanism besides a TATA box to initiate transcription [JASPAR].
Secondary Structure
While the MSH6 gene has been fairly well characterized, much less is known about its secondary structure. Nevertheless, an analysis of the MSH6 structure reveals some interesting characteristics of the gene. Using PREDATOR, three types of expression within the MSH6 protein were found [PREDATOR]:
- 32.13% Alpha Helix
- 13.6% Extended Strand
- 54.26% random coil
The pattern of extended strand, alpha helices, and random coil revealed no immediately obvious pattern [Figure 3]. Nevertheless, an interesting pattern was found. Within the highly conserved region of the MSH6 protein (amino acids 1000-1360), 41.67% of the secondary structure was composed of alpha helices while just 28.7% of the less highly conserved region was [PREDATOR]. Alpha helices are suspected to play a role in DNA binding, suggesting that the more conserved portion of the MSH6 gene and protein may play a particularly crucial role in the DNA binding that is necessary for the repair of mismatched DNA [DNA-Binding Motifs]. Kyte-Doolittle Hydrophobicity plots revealed no transmembrane domains, suggesting that MSH6 exists freely within the cell, likely enabling MSH6 to have access to aberrant DNA [Hydropathicity Plots].
Figure 3 [Click to Enlarge]
Single Nucleotide Polymorphisms
Within human MSH6, many single nucleotide polymorphisms have been identified, some of which are potentially implicated in the development of colorectal cancer. 72 total SNPs of a variety of different types have been identified within the MSH6 gene [ENSEMBL]:
- Upstream 1
- Non-Synoymous coding 2
- Synonymous coding 19
- Intronic 21
- Splice Site intronic 3
- Frameshift coding 3
- Splice Site, 3Prime UTR 1
- Downstream 1
Total 72
(Note: these data include both substitutions and indels)
Presumably, intronic mutations and synonymous coding mutations should not affect the ability of the MSH6 gene to function. However, any non-synonymous coding mutation could have a detrimental impact on the ability of the protein to function. The importance of the MSH6 protein’s ability to bind to DNA for proper functioning suggests that any mutation that affects the ability of MSH6 to bind DNA could have a particularly detrimental impact on protein function. Because alpha helices are suspected to play a role in DNA binding [DNA-Binding Motifs], a mutation in an alpha-helix could have an especially negative impact on the function of MSH6.
One SNP that is located in an alpha helix is rs35717727, a non-synonymous coding mutation [PREDATOR, ENSEMBL]. In 98-99% of the population, glutamic acid is found at amino acid 1234 while 1-2% of the population has the amino acid glutamine instead [ENSEMBL]. This mutation could have a negative impact on the function of MSH6 because glutamic acid and glutamine have different properties—glutamic acid is charged while glutamine is not [Amino Acid Abbreviations]. An alteration in the charge of the MSH6 protein in a presumed DNA binding region has the potential to affect the ability of MSH6 to bind mismatched DNA to initiate repair which could then possibly lead to the development of colorectal cancer.
Conclusion
In conclusion, analysis of the MSH6 gene reveals several important aspects of the gene:
- -MSH6 is very well conserved among a variety of species, suggesting it plays an important role in the organisms
- -The ability of MSH6 to bind to DNA is paramount, a finding that is supported by its upstream motifs, large percentage of alpha helices, and high conservation of DNA binding regions
- -Any inability of MSH6 to bind to mismatched DNA has the potential to lead to the development of colon cancer
References:
1. Amino Acid Abbreviations [http://bio.davidson.edu/Courses/Molbio/aatable.html]
2. BLAST: Basic Local Alignment and Search Tool [http://www.ncbi.nlm.nih.gov/blast/Blast.cgi]
3. DNA-Binding Motifs. [http://arapaho.nsuok.edu/~biology/Tutorials/DNAbinding.htm]
4. EBI Tools: Clustal W [http://www.ebi.ac.uk/Tools/clustalw/index.html]
5. Ensembl Genome Browser [http://www.ensembl.org/index.html]
6. Hydropathicity Plots [http://www.vivo.colostate.edu/molkit/hydropathy/index.html]
7. The JASPAR Database [http://jaspar.genereg.net/]
8. National Center for Biotechnology Information (NCBI) [http://www.ncbi.nlm.nih.gov/]
9. NCBI Conserved Domain Database (CDD) [http://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml]
10. Predator [http://bioweb.pasteur.fr/seqanal/interfaces/predator-simple.html]
11. Saccharomyces Genome Database (SGD) [http://www.yeastgenome.org/]
MSH6 Expression in Yeast
While much information can be learned from directly studying human MSH6, the importance of a good model organism to study the gene is vital as well. The high conservation of MSH6 in multiple species provides a wealth of potential model organisms for the study of MSH6. However, the large amount of information available about genome function and regulation in yeast make them an especially helpful model organism. While the human and yeast amino acid sequence of MSH6 has only 28% conservation, the similarity in their molecular function, cellular component, and biological process (below) make the MSH6 yeast gene a good model for the study of human MSH6 [NCBI; SGD]:
- Molecular Function
- ATP Binding
- ATPase Activity
- Guanine/Thymine mispair binding
- Single Base insertion/deletion binding
- Cellular Component
- Nucleus
- Nucleus
- Biological Process
- Mismatch repair
Response of MSH6 to MSN2/MSN4 Deletions
The Hoopes Lab at Washington University in St. Louis generously provided us with microarray data that looked at the effect of several deletions and growth states on genomic expression in yeast. Some of these microarrays looked at the effects of a deletion in MSN2/MSN4. MSN2 and MSN4 are both related to the activation of transcription. They bind elements of some DNA during stressful conditions and induce the expression of those genes [NCBI]. In addition to assessing the effects of the MSN deletions, the Hoopes lab looked at the effects of stationary versus log phase growth.
Since MSH6 is involved in DNA repair in mitosis after DNA replication, it is reasonable to think that a deletion in MSN, which plays a role in expression of some genes, might have an impact on the expression of MSH6 [NCBI].
Thus, 3 microarray experiments were analyzed:
- wild type stationary phase growth vs. log phase growth (WTst/WTlog)
- msn2/msn4 deletion mutant stationary phase growth vs. wild type log phase growth (Δ MSNst/WTlog)
- msn2/msn4 deletion mutant stationary phase growth vs. log phase growth (Δ MSNst/ Δ MSNlog)
Indeed, the transcription of MSH6 [red line below] was affected by MSN deletions [Figure 1]. When MSN4/6 were deleted, there was a drop in the expression of MSH6, suggesting that MSN2/4 normally play a role in the induction of expression of MSH6. Further, the change in MSH6 expression was most pronounced in stationary growth phase, suggesting that MSH6 may have differential activity in stationary versus log phase growth.
Figure 1 [Click to Enlarge]
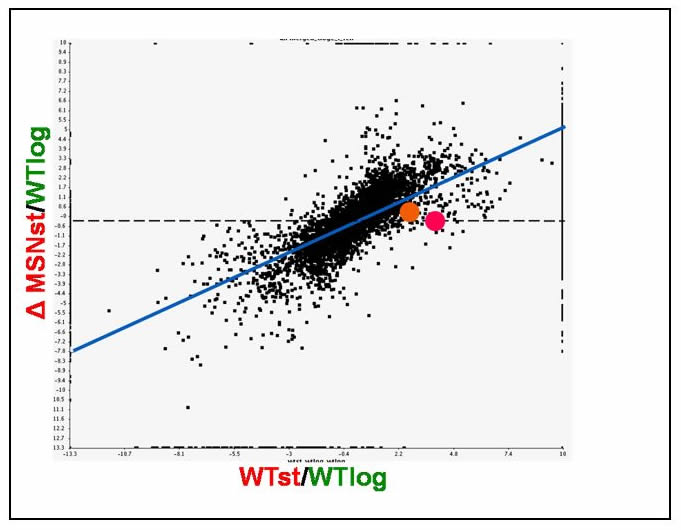
In order for MSH6 to function in DNA repair in vivo, it must form a heterodimer with the protein of the MSH2 gene [Acharya 1996]. As a result, the function of MSH2 in response to an MSN4/6 deletion was also assessed to determine whether or not MSH2 and MSH6 are potentially regulated by some common factor. In the presence of deletions in MSN4 and MSN6, MSH2 and MSH6 responded in a similar manner, suggesting that MSH2 [pink dot] and MSH6 [orange dot] are controlled by some common factor [Figure 2].
Figure 2 [Click to Enlarge]
MSH2/MSH6 Activity and Control
To assess whether or not MSH2 and MSH6 act in a similar manner in other conditions, microarray data from several other studies were also examined [Expression Connection]. In the cell cycle, both MSH2 and MSH6 are expressed in a very dynamic manner, with their expression rapidly changing within a short period of time. The dynamic expression of MSH2 and MSH6 in the cell cycle was significant at a Pearson Correlation Coefficient of >0.8 and suggests that they are potentially regulated by some common 3rd factor [Spellman 1998].
In order to determine a possible factor that could be regulating the expression of MSH2 and MSH6, 9 genes involved in transcription that expressed similar patterns of expression as MSH2 and MSH6 in response to the deletions in MSN4 and MSN6 were assessed. One transcription factor’s expression emerged as being particularly interesting.
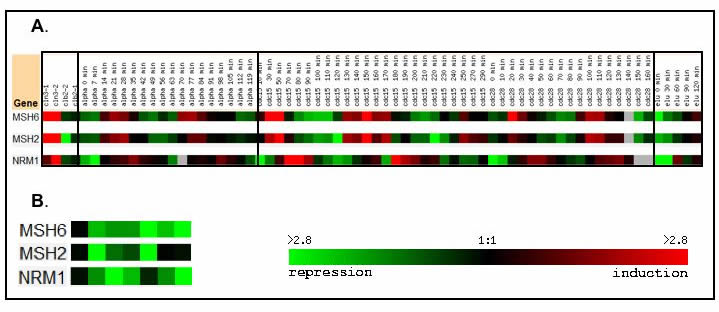
The NRM1 gene represses transcription when cells exit the G1 phase [Entrez Gene]. When the activity of MSH6, MSH2, and NRM1 was studied in the cell cycle, an intriguing pattern was seen. While MSH2 and MSH6 act in nearly simultaneous manners, NRM1 acts in a very similar manner, but its expression is slightly delayed relative to that of MSH2 and MSH6 [Figure 3A]. The activity of MSH6, MSH2 and NRM1 was not significant at a Pearson Correlation of >0.8, but visual inspection reveals that the expression of the three genes does appear to be related [Spellman 1998].
Figure 3 [Click to Enlarge]
Great similarity of expression was also seen in the activities of MSH2, MSH6, and NRM1 in response to alpha-factor, though their expression was not significant at a Pearson Correlation of >0.8 [Roberts 2000]. Alpha-factor is secreted by yeast cells during mating and activates Protein Kinase C which in turn sets off a signaling cascade in the yeast cells that triggers a variety of changes in the transcription of the genome [Roberts 2000].
In the presence of alpha-factor, the expression of MSH2, MSH6, and NRM1 were all significantly repressed [Figure 3B] [Roberts 2000]. Though the marked delay in NRM1 expression was not as visible, the strong similarity in the expression patterns of MSH2, MSH6, and NRM1 in response to alpha-factor suggests that there could be some common factor regulating their expression or that one of these three genes could be regulating the expression of the other two.
Future Experiments
To asses whether MSH2, MSH6, or NRM1 expression is regulating expression of the other two genes, more bench experiments will need to be performed. To determine potential experiments, strains of yeast from the TRIPLES database were analyzed for their usefulness in elucidating the control mechanisms of MSH2, MSH6, and NRM1. There were two strains of yeast that would be particularly useful in answering this question, one that had an insertion in MSH2 that interrupted its function and one that had an insertion in MSH6 that interrupted its function [TRIPLES].
To identify if MSH2 or MSH6 is controlling the expression the other two genes, the two insertion strains should each be grown in normal growth conditions with their cell cycles synchronized. Synchronization of the cell cycles is vital given the dynamic expression patterns of MSH2, MSH6, and NRM1 [Spellman 1998]. Then, two microarray studies should be performed:
- ΔMSH6/WT
- ΔMSH2/WT
The microarray chip would simply need spots of cDNA and positive and negative control spots to ensure that the microarray was completed correctly. Finally, the expression patterns of NRM1 and MSH2/6 should be assessed. If a deletion in MSH2 or MSH6 controls the expression of the other two genes, some resultant change in their transcription levels should be seen. Therefore, this study should allow determination of whether or not MSH2 or MSH6 is controlling the expression patterns of the other 2 genes.
Conclusion
In conclusion, a study of microarray data revealed several interesting aspects of the activity and potential control of MSH6:
- MSN2/MSN4 appear to induce the expression of MSH6, particularly in stationary growth phase
- Similar expression patterns of MSH2 and MSH6 suggest potential coregulation
- NRM1 may also be coregulated with MSH2/MSH6 or it could be regulated by MSH6 or MSH2
- Future study should help elucidate if MSH2 or MSH6 is controlling expression of the other genes studied
References:
- Acharya S, Wilson T, Gradia S, Kane MF, Guerrette S, Marsischky GT, Kolodner R, Fishel R: hMSH2 forms specific mispair-binding complexes with hMSH3 and hMSH6. Proc Natl Acad Sci USA 1996, 93: 13629-13634.
- Entrez Gene [http://www.ncbi.nlm.nih.gov/sites/entrez?db=gene]
- Expression Connection [ http://db.yeastgenome.org/cgi-bin/expression/expressionConnection.pl]
- National Center for Biotechnology Information (NCBI) [http://www.ncbi.nlm.nih.gov/]
- Roberts CJ, Nelson B, Marton MJ, Stoughton R, Meyer MR, Bennett HA, He YD, Dai H, Walker WL, Hughes TR, Tyers M, Boone C, Friend SH: Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 2000, 287:873-880.
- Saccharomyces Genome Database (SGD) [http://www.yeastgenome.org/]
- Spellman PT, Sherlock G, Zhang MQ, Iyer VR, Anders K, Eisen MB, Brown PO, Botstein D, Futcher B: Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 1998, 9:3273-97
- TRIPLES Database [http://ygac.med.yale.edu/triples/default.htm]
Acknowledgements:
The Hoopes Lab at Washington University, St. Louis for the microarray data.
The MSH6 Gene Circuit
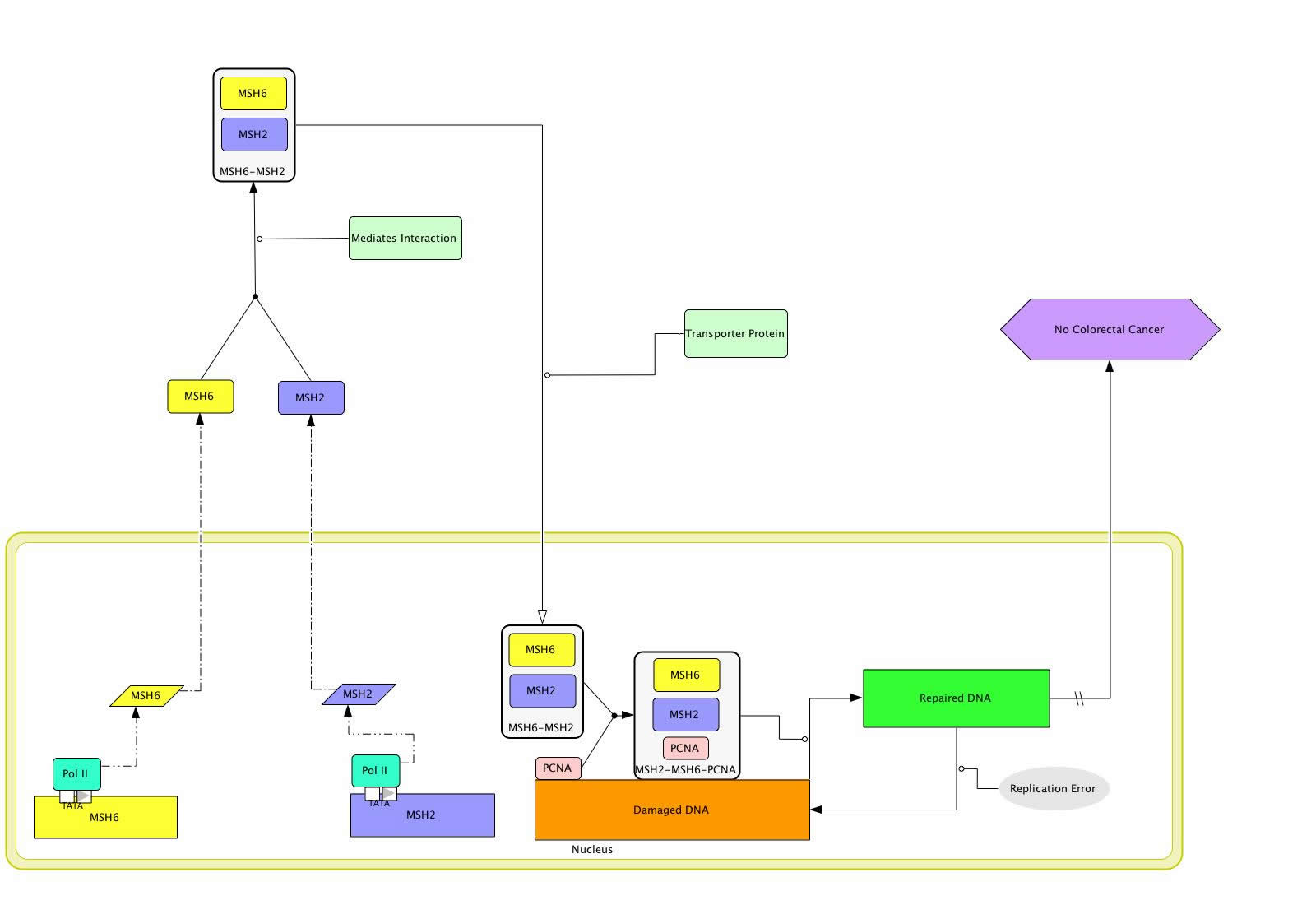
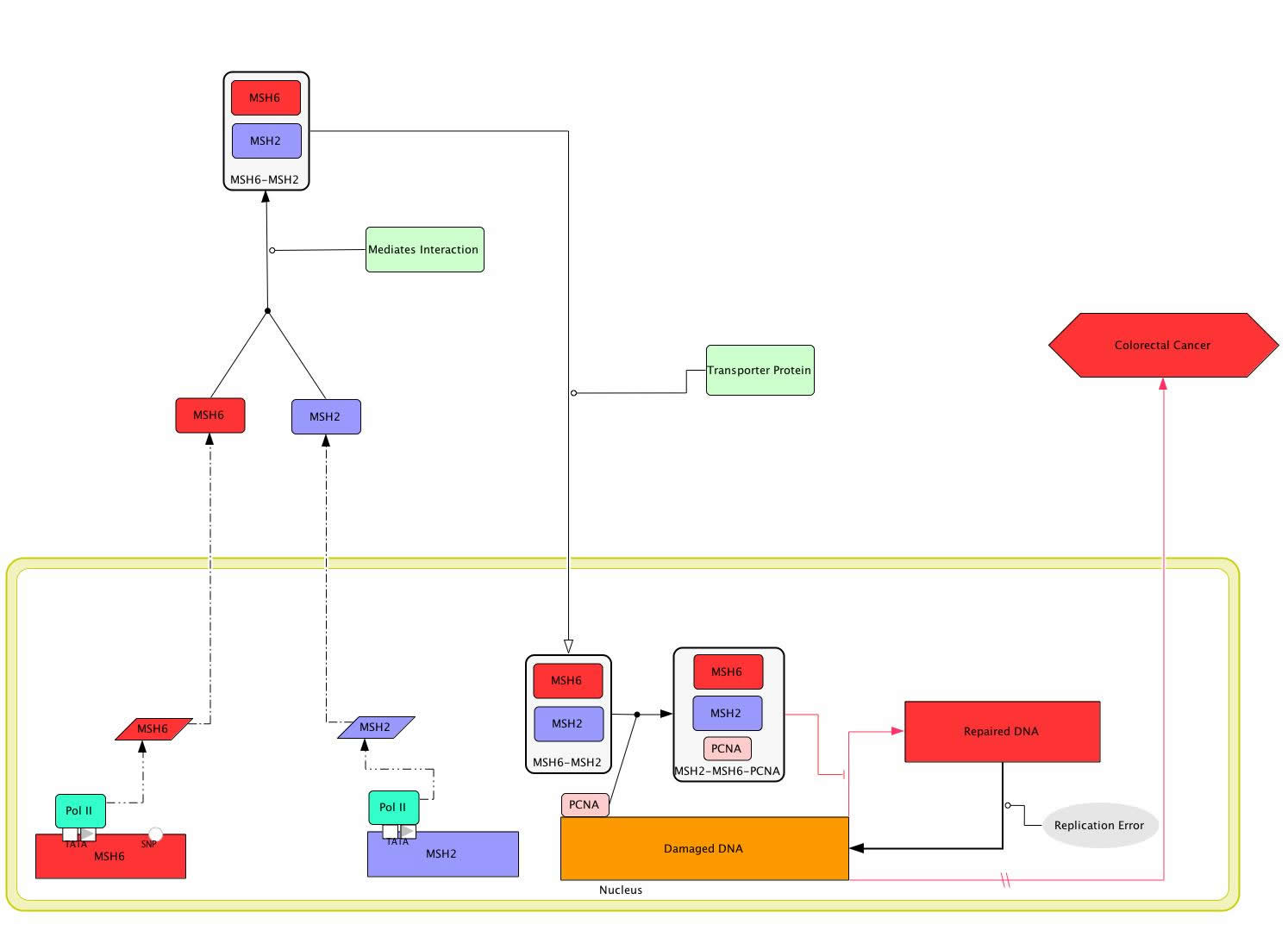
The MSH6 protein is not functional on its own but instead forms a heterodimer with MSH2 that is then functional in DNA repair [Figure 1] [Acharya 1996]. However, in order for the MSH2-MSH6 heterodimer to recognize mismatched DNA, it must also associate with the protein PCNA [Clark 2000; Flores-Rozas 2000; Lau 2003]. PCNA, or Proliferating Cell Nuclear Antigen, is suspected to tag newly replicated DNA and play a role in directing the MSH2-MSH6 heterodimer to new DNA that potentially contains mismatched bases [Flores-Rozas 2002; Lau 2003]. The ability of the MSH6-MSH2-PCNA complex to bind DNA appears to be interrupted when there is mismatched DNA. As a result, it is believed that when the MSH6-MSH2-PCNA complex finds mismatched DNA, the MSH2-MSH6 heterodimer dissociates from PCNA and binds to the mismatched DNA, enabling repair of the aberrant DNA and, through some unknown mechanism, preventing the development of colon cancer [Colon Cancer; Lau 2003; OMIM].
Figure 1 [Click to Enlarge]
The Disease MSH6 Gene Circuit
However, some persons have mutations in their MSH6 that cause their gene to vary from the rest of the population. One known mutation, or SNP, in the MSH6 gene is a non-synonymous coding mutation found within an alpha helix at amino acid 1234 [ENSEMBL; PREDATOR]. 98-99% of the population has Glutamic Acid at amino acid 1234 while 1-2% has the amino acid glutamine instead [ENSEMBL]. This mutation could have a negative impact on the function of MSH6 because glutamic acid and glutamine have different properties—Glutamic acid is charged while glutamine is not [Amino Acid Abbreviations]. An alteration in the charge of the MSH6 protein has the potential to have negative affects. If the mutation in MSH6 is in a DNA binding region, the MSH2-MSH6 heterodimer should be able to form as should the MSH2-MSH6-PCNA complex [Figure 2]. However, once the MSH2-MSH6-PCNA complex locates mismatched DNA, MSH2-MSH6 would be unable to bind to the mismatched DNA and initiate its repair. As a result, the mismatched DNA would remain incorrect rather than being repaired. As a result of mismatched DNA, the person is at a higher risk of developing colon cancer [Lau 2003; Online Mendelian Inheritance in Man].
Figure 2 [Click to Enlarge]
Other Interactions
In addition to interacting with MSH2 and PCNA, MSH6 also interacts with a group of proteins that are all components of a large complex called BASC, or BRCA-1-associated genome surveillance complex [Figure 3] [Human Interactome Map; Wang 2000]. This large group of proteins is believed to play a role in the identification and repair of mismatched DNA. It is suggested that BASC plays a role in sensing the presence of malformed DNA and then directs the necessary components for repair, one of which is the MSH2-MSH6 heterodimer, to the location of the erroneous DNA [Wang 2000]. This paper [Wang 2000] underscores the fact that the MSH2-MSH6 heterodimer is only one part of a much larger group of proteins that play a role in the repair of DNA.
Figure 3 [Click to Enlarge]
Why Colorectal Cancer?
If, as Wang [2000] suggests, MSH2-MSH6 is part of a complex that plays a role in DNA repair throughout the body, it is intriguing to consider why aberrations in the MSH6 gene are associated with a predisposition to the development of colon cancer, rather than to cancer in general [Kariola 2002]. Thus far, no clear answer to the question has been found [Colon Cancer]. It is possible that there is some colon-specific component (i.e. an enzyme or other protein) that affects MSH6 in the colon in a different manner than in the rest of the body. Alternatively, it is possible that the conditions of the colon make MSH6 more sensitive so that small errors in the structure of MSH6 are more likely to cause problems in the colon than in other parts of the body.
In order to attempt to determine if there is some specific factor in the colon that associates with MSH6 that isn’t present in other parts of the body, potential protein-protein interactions that are specific to the colon would need to be determined. There are several different methods that can be used to identify protein interactions. Yeast two-hybrids have been a particularly successful method for determining protein-protein interactions. It is likely that Yeast two-hybrids could be particularly useful for determining potential interactions of MSH6 and transcription factors within the colon. The Yeast two-hybrid method requires that the protein be tagged to the nucleus, making MSH6 a good candidate since it has to be inside the nucleus for DNA repair. By performing Yeast two-hybrids on transcription factors known to exist within the colon, potential new interactions that might affect MSH6 expression could be found.
Alternatively, there could be other proteins besides transcription factors that affect the expression or function of MSH6 within the colon. To assess other protein-protein interactions, NMR would first need to be utilized to identify all the proteins that are present within a sample of the colon. Then, a microarray hybrid experiment would need to be done. The proteins that were identified as being present within the colon would be spotted on the chip and MSH6 would be the probe. Positive and negative controls would be necessary to ensure that the microarray hybridization was functional. Finally, scanning of the microarray for MSH6-protein interactions would allow assessment of proteins known to exist within the colon that MSH6 might be associating with.
Performing a similar experiment on control cells (IE cells from an organ not affected by an MSH6 mutation) should enable determination of the interactions of MSH6 and other proteins not in the colon. If interactions are seen in one group and not the other, that interaction should be further assessed to see how the protein and MSH6 interact. Because the alteration of MSH6 activity in just the colon could be due to a loss of interaction or to a gain in some interaction, interactions in both the colon and in control cells would need to be assessed.
Potential Treatment for Persons with an MSH6 Mutation
If, as proposed above, one reason that the MSH6 protein is unable to repair DNA is because it cannot bind to the aberrant DNA and fix it, then having functional MSH6 present would likely be useful. Thus, one potentially helpful treatment for persons with an MSH6 mutation that predisposes them to develop colon cancer would be to insert wild type MSH6 into their cells that can then form a heterodimer with MSH2 and bind to aberrant DNA to initiate repair.
Since transcription and repair of transcription have to occur in all cells, having WT MSH6 protein in all cells should not be problematic. As a result, using a protein-transduction domain (PTD) on the WT MSH6 would enable the MSH6 protein to cross cell membranes and enter into all cells in the body. Normally, MSH6 is tagged for the nucleus, so the WT MSH6 should enter the nucleus and find MSH2 to form a heterodimer with. Ideally, using PTD-mediated delivery of MSH6 would not illicit an immune response in the host, so there would be no potential rejection of the WT MSH6.
However, an immune response to the MSH6 is a possibility that needs to be considered. Additionally, there might be some concern about the availability of MSH2 to bind with the WT MSH6. If the mutated MSH6 protein is still able to form a heterodimer with MSH2, then it is possible that most of the MSH2 will be bound to the mutated MSH6 and unable to bind to the newly inserted WT MSH6. Alternatively, it is possible that the mutated MSH6 is degraded, so the WT MSH6 protein would be able to bind to MSH2. An assessment of the state of MSH6 proteins that are unable to bind DNA would likely be useful in helping to determine the likelihood of there being enough MSH2 present to bind the WT MSH6.
Conclusion
In conclusion, a study of the circuit diagram of MSH6 revealed the following information:
- One potential reason that MSH6 is unable to function is due to an inability to bind aberrant DNA
- It is unclear why MSH6 mutations predispose an individual specifically to colon cancer—analysis of factors within the colon might help to elucidate this
- Insertion of WT MSH6 protein into persons with MSH6 mutations has the potential to curb the effects of the MSH6 mutation
References:
- Acharya S, Wilson T, Gradia S, Kane MF, Guerrette S, Marsischky GT, Kolodner R, Fishel R: hMSH2 forms specific mispair-binding complexes with hMSH3 and hMSH6. Proc Natl Acad Sci USA 1996, 93: 13629-13634.
- Amino Acid Abbreviations [http://bio.davidson.edu/Courses/Molbio/aatable.html]
- Clark AB, Valle F, Drotschmann K, Gary RK, Kunkel TA: Functional interaction of proliferating cell nuclear antigen with MSH2-MSH6 and MSH2-MSH3 complexes. The Journal of Biol Chem 2000, 275: 36498-36501.
- Colon Cancer [http://www.ncbi.nlm.nih.gov/books/bv.fcgi?call=bv.View..ShowSection&rid=gnd.section.102]
- Ensembl Genome Browser [http://www.ensembl.org/index.html]
- Flores-Rozas H, Clark D, Kolodner RD: Proliferating cell nuclear antigen and Msh2p-Msh6p interact to form an active mispair recognition complex. Nature America 2000, 26: 375-378.
- Human Interactome Map. [http://www.himap.org/]
- Kariola R, Raevaara T, Lönnqvist K, Nyström-Lahti M: Functional analysis of MSH6 mutations linked to kindreds with putative hereditary non-polyposis colorectal cancer syndrome. Human Molecular Genetics 2002, 11: 1303-1310.
- Lau PJ, Kolodner RD: Transfer of the MSH2·MSH6 complex from proliferating cell nuclear antigen to mispaired bases in DNA. The Journal of Biol Chem 2003, 278: 14-17.
- Online Mendelian Inheritance in Men (OMIM) [http://www.ncbi.nlm.nih.gov/sites/entrez?db=OMIM]
- Predator [http://bioweb.pasteur.fr/seqanal/interfaces/predator-simple.html]
- Wang Y, Cortex D, Yazdi P, Neff N, Elledge SJ, Qin J: BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes and Dev 2000, 14: 927-939.