This web page was produced as an assignment for an undergraduate course at Davidson College.
The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution
Summary/Figure explanations
This article deals with the ATCC11842 strain of the organism Lactobacillus bulgaricus, which is responsible for the commercial production of yogurt, and details the analysis of its genomic sequence. Through genomic analysis techniques and gene annotation, the researchers came up with the conclusion that L. bulgaricus is in the process of evolving into a more specialized environmental niche, namely, milk.
The researchers began by examining the properties and characteristics of the primary DNA sequence in full. Using shotgun sequencing, the genome of L. bulgaricus was found to be 1,864,998 base pairs long. Table 1 in the article outlines the basic facts regarding the genomic sequence, and gives, in addition to the genome size, the GC content of the whole genome (49.7%), the GC content of coding sequences (51.6%), the GC content of coding sequences at codon position 3 (65.0%), number of coding sequences (1562), the percentage of the genome that are coding sequences (73%), the number of coding sequences with unknown function (598), and the number of pseudogenes (270).
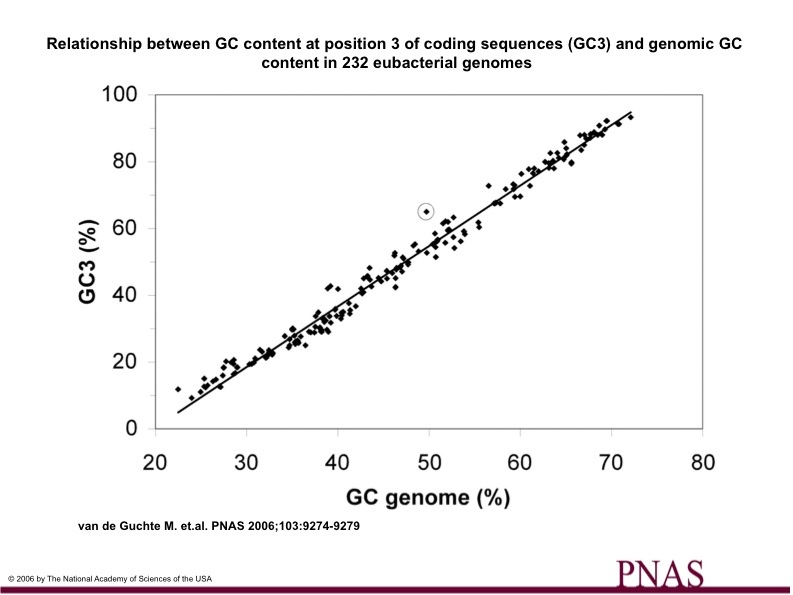
These numbers given in Table 1 reveal some striking differences in L. bulgaricus compared to its closest genetic relatives. For one, the overall GC content is much higher than those of L. bulgaricus's relatives. This is attributed to the abnormally high 65.0% GC content of codon position 3. This is illustrated in Figure 1, which plots the relationship of GC content at codon position 3 in relation to genomic GC content for several eubacteria (232). As can be seen, L. bulgaricus (the circled data point) is one of the furthest points away from the best fit line, showing how the GC content of 65.0% at codon position 3 differs from the expected value.

(Figure 1)
The article then goes on to state that evolution at codon postion 3 is generally quicker than at positions 1 or 2, and the fact that L. bulgaricus has a much higher GC content at position three than expected suggests that L. bulgaricus is in the midst of evolutionary change. However, the authors of the article do not go into detail, and do not explain why codon position 3 evolves faster and therefore do not offer any explanation of the information on which they based their observations.
In addition to the GC content for codon position 3, L. bulgaricus also has an unsuspectedly high number of pseudogenes. The high number of pseudogenes seems to suggest that genes in the genome of L. bulgaricus are actively being taken out of the genome by way of selective pressures. Many of the pseudogenes seem to have formerly coded for proteins that can no longer be found in any of the genome, which suggests L. bulgaricus is in the process of specialization. Many functions that are completely lost to the organisms due to pseudogenes are involved in metabolism and DNA transcription. Transport systems for lactose, fructose, and glycerol are present, but many other sugar transport systems and pathways are either not present or incomplete, implying that L. bulgaricus was once fit to live in other, non-milk environments prior to these changes. There is also an absence of enzymes involved in amino acid synthesis, which also suggests an adaptation to living in milk, though the authors do not clearly state why this is so.
Figure 2 shows some aspects of the L. bulgaricus genome graphically. The outermost circle displays the pseudogenes and on which strand they appear (red or blue). The second circle shows insertion sequence elements (transposases and hypothetical genes). Gray represents elements with less than 4 copies. Elements with more than 5 copies are represented by red (ISL7), purple (ISL4), blue (ISL5), and green (ISL4-5). Another circle of note is the third circle (from the outside), which depicts the coding sequences of the genome (not including pseudogenes) and on which DNA strand they are found (red or blue). The fourth circle shows rRNA and tRNA genes in red and green, respectively. Circles 5 and 6 depict GC with a window size of 2000 (less than -.1 in cyan and more than +.1 in red) and AT with a window size of 500 (less than .3 in cyan and more than .7 in red), respectively. The innermost circle shows genomic position.
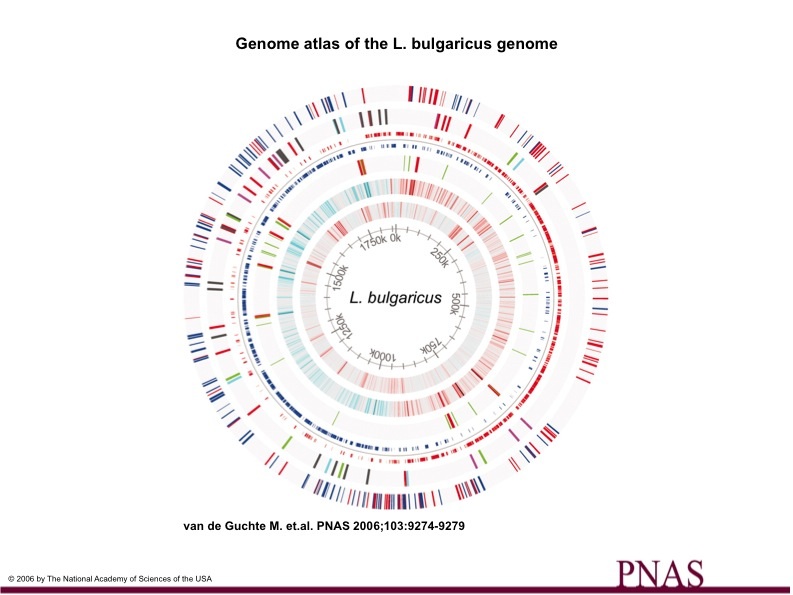
(Figure 2)
The number of tRNA and rRNA genes in the L. bulgaricus genome is also interesting to note. There is typically a correlation between number of tRNA and rRNA genes and genome size, which is shown in Figure 3 (below). Chart A compares the number of 16s rRNA genes versus the genome size of 54 genomes, while Chart B shows the number of tRNA genes compared to genome size of 54 genomes. Again, L. bulgaricus (shown by the circled data points) is an outlier in both graphs. The article states that both tRNA and rRNA numbers are 50% higher than the average and 20-30% higher than the next highest values from other organisms. The authors explain that the elevated number of tRNA and rRNA genes correspong to numbers similar to organisms with 3-5 million base pair genomes, which may may imply that L. bulgaricus's genome has recently reduced in size. However, it is important to remember that this may not be the only explanation. There are several exceptions to the "rule" in genomics, and L. bulgaricus having high numbers of tRNA and rRNA genes may be completely unrelated to the matter of whether or not the organism has undergone recent evolutionary changes.
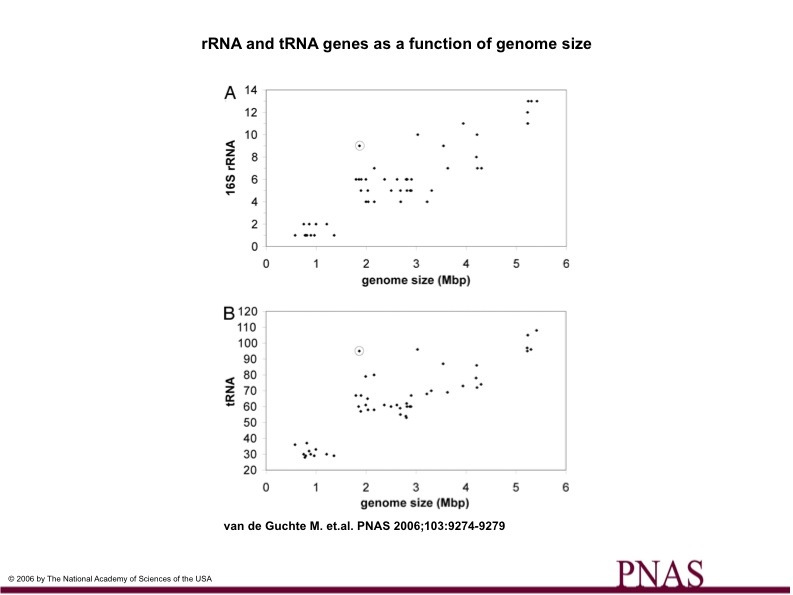
(Figure 3)
The researchers also compared the genome of L. bulgaricus to other closely related organisms. Figure 4 shows a genome comparison between L. bulgaricus and L. acidophilus. It can easily be seen that the two organisms are highly syntenic, and are obviously closely related. The colors of the data points indicate similarity between the genes, with red, orange, and yellow being the most similar. The most notable differences between L. bulgaricus and its relatives are the presence of pathways for the synthesis of folate, saturated fatty acids, purines, and pyrimidines in L. bulgaricus, which are not present in the others, as well as the lack of genes involved in sugar transport and metabolism in L. bulgaricus that are found in the other organisms.
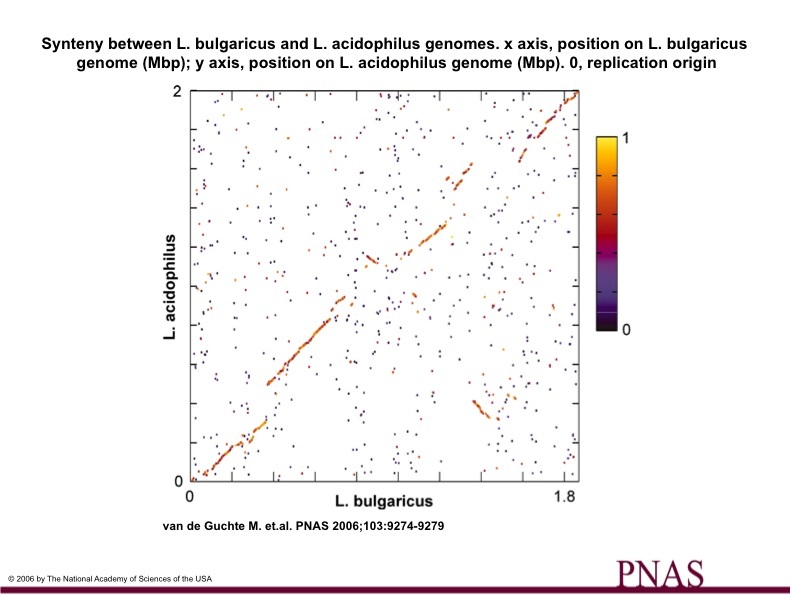
(Figure 4)
Opinion/ Lessons learned
I believe the authors of this article did a great job of conveying their information and explaining their hypothesis and conclusion. The data they present supports their conclusions nicely, and, for the most part, their logic is clear and easy to follow. However, I do have a few qualms in regard to the article. The authors seem to present their information and show how it supports their conclusions, while at the same time either neglecting to mention or putting very little emphasis on other possibilities. I believe its important to at least briefly acknowledge that there may be other explanations and to give examples of those other possibilities. Also, in a few of their descriptions and subsequent explanations, the authors chose not to incluse the data their writing was based from. I think all relevant data should be made known, and allow readers and peers to make their own conclusions regarding it. This is especially true if the author is going to use that excluded data to help make their case. The authors could have also given more information as to which organisms they compared L. bulgaricus to in their figures, as well as why those organisms were chosen.
This article illustrated different genome analysis techniques and explained what inferences could be drawn from them quite nicely. I especially liked Figures 1 and 3, and the data they compared. They are comparisons I myself would not have thought to make, and they clearly convey their information and support the authors' conclusions. It was also useful for them to compare the genome of L. bulgaricus with its closest relatives and to make clear the differentiation. This provided an insight into how L. bulgaricus could be diverging from its genetic relatives. Reading this article also showed me the importance of clearly presenting data and providing different possible explanations for the data.
References
Guchte et al. 2006. The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. PNAS: Proceedings of the National Academy of Sciences of the United States of America. <http://www.pnas.org/content/103/24/9274.full?sid=b48b2cc8-db0d-454e-89a2-a9ff030c85a4>. Accessed 2008 Oct 9.
Return To Biology Course Materials
Davidson College Biology Department