Sequencing of 50 Human Exomes Reveals Adaptation to High Altitude
Yi et al. (2010)
This web page was produced as an assignment for an undergraduate course at Davidson College.
Summary:
Evaluation:
I liked this paper because I thought that the authors of this paper used a very interesting methodology to try and identify genes that, through natural selection, increased a population’s fitness to live in a high altitude environment. By identifying SNPs with very elevated allele frequencies in one population compared to the other, they identified SNPs that had been acted upon by natural selection and were able to establish in which population natural selection was acting by comparing both groups to a third more distantly related population. I believe that this methodology has important implications for identifying ways in which natural selection can affect a population and determining evolutionary relationships.
I think it was good that the authors focused on the one gene who’s SNPs were the most significant because they were not trying to imply that their findings were overly significant by saying that they had identified tons of genes responsible for natural selection. However, figure 1 did not clearly convey how they were proving EPAS1 to be an outlier, and as a result a gene likely acted on by natural selection.
Introduction:
One very recent example of how environmental pressures can cause changes in allele frequencies of a population is found in residents of the Tibetan Plateau. Natural selection has been acting on the people of the Tibetan Plateau to provide them with advantageous alleles that allow them to survive in high altitudes with oxygen levels about 40% lower than at sea level. These adaptations can be seen in Tibetans’ average birth weight, hemoglobin levels, and oxygen saturation of blood in infants and adults after exercise. Proof of these altered allele frequencies comes from this paper, whose authors sequenced 50 exomes of Tibetans and found changes in genes that represent strong candidates for altitude adaptation.
Method:
The exomes of 50 unrelated Tibetans from two villages in the Tibet Autonomous Region of China were sequenced using Illumina Genome Analyzer II platform. The exome sequences of interest were obtained with the NimbleGen 2.1 exon capture array. By using this they could target 34 Mb of sequence from exons and flanking regions in almost 20,000 genes. Sequence reads were aligned using SOAP to the reference human genome. Since exomes were only sequenced to a mean depth of 18 times, which does not guarantee confident inference of different genotypes, they statistically estimated the probability of each possible genotype with a Bayesian algorithm. This algorithm also estimated SNP probabilities and population allele frequencies at each site.
The exome data was compared to 40 genomes from Han individuals, which were sequenced around fourfold coverage per individual. The samples were appropriate comparisons because of the low genetic differentiation between the two populations. Genes with strong allele frequency differences between populations are potential targets for natural selection, but to determine what group was affected by selection exome sequences had to be compared to a more distantly related population. They used 200 Danish samples, analyzed in the same way as the Tibetan samples, and by comparing the FST values between all the samples they determined the frequency change occurred in the Tibetan population. FST values measure population genetic differentiation between samples.
Results:
Genes with extreme Tibetan population branch statistic values (PBS), which are also a measure of divergence, represent strong candidates for the genetic basis of altitude adaptation. 34 genes in the data set that had functions related to “response to hypoxia” had significantly higher PBS values than the genome average. The gene with the strongest signal of selection was the EPAS1 gene and based on frequency differences it was inferred to have a very long Tibetan branch relative to other genes. To confirm natural selection, PBS values were compared against neutral simulations under the estimated demographic model and out of one million simulations none surpassed the PBS value for EPAS1. The result also remained statistically significant after accounting for the number of genes tested. Other genes had p-values under .005, but after correction for multiple tests none of them were statistically significant even though the authors still believe that some of the genes may contribute to altitude adaption.
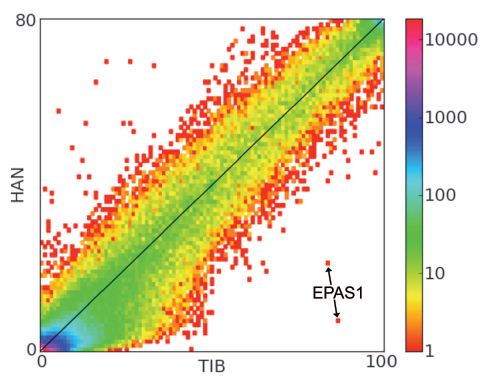
Figure 1: A two-dimensional unfolded site frequency spectrum for SNPs in Tibetan (x axis) and Han (y axis) population samples. This chart shows the SNP frequency distribution found in the populations. The caption of the paper says that, “the number of SNPs detected is color-coded according to the logarithmic scale plotted on the right”, but I found this explanation to be a bit confusing. To me it seems as if the x-axis and y-axis are plotting the frequency of the SNPs in the populations and can be read as, the SNPs in the lower left are not common in either population and the SNPs at the upper right are very common in both populations. This interpretation of the chart correlates with their proposed result of SNPs in the EPAS1 gene being much more frequent in the Tibetan population than the Han population, but doesn’t take the color scale into consideration. The color scale is very confusing to read. The authors have chosen to make both one and 10000 very similar shades of red, which make it difficult to tell if the points on the graph are representative of one or 10000. Additionally, if the color scale is measuring the number of SNPs detected, how can it reach 10000 SNPs if only 200 people were included in the study? The figure also has colored the genes in the lower left corner with a higher SNP frequency color than those found higher and farther to the right. This contradicts the x-axis and y-axis.I feel that the authors should have excluded the color scale to make the figure more legible. However, the main point of the figure can still be determined, which is that EPAS1 is represented by the two intronic SNPs being pointed out by the arrows. Since the SNPs are such outliers to the rest of the group they show strongly elevated derived (non-ancestral) allele frequencies in the Tibetan sample compared to the Han sample, which gives evidence for natural selection acting on that gene.
Table 1: This table identifies the top 30 genes with the highest PBS values, which represent the strongest frequency changes found in the Tibetan population. These PBS values are based on the Fst value, which ranges from 0 to 1. Zero represents a fully interbreeding population and a one represents distinct populations. P-values are also given to show significance, but after correction none of these are still statistically significant. Oxygen related genes within 100 kb of the identified loci are given to show how these loci could be involved in altitude adaptation.
Figure 2: Both are representations of population-specific allele frequency change. (A) Each dot on this graph represents one gene and its placement across the x-axis tells us the number of SNPs found in the gene. The y-axis is showing the Fst based PBS value for the Tibetan branch of the exome comparison. This means it is measuring how genetically different the Tibetan genes are when compared to the equivalent gene in the other populations. The three outlier genes are good candidates for natural selection, since they are much different than the average variance between populations. (B) The branches on the left represent the relatedness between the populations based on the genomic average Fst-based branch lengths. The branches on the right represent the relatedness between the populations based on Fst-based branch lengths for the EPAS1 gene. These differences indicate the significant differentiation by the Tibetan population caused by natural selection, identified because of the rapid change in allele frequencies.
EPAS1:
The EPAS1 SNP with the greatest frequency difference was intronic and had a frequency of 9% in Han population and 87% in the Tibetan sample. No nonsynonymous SNP had a population frequency difference greater than 6%. EPAS1 is also known as hypoxia-inducible factor 2α (HIF-2α). The narrow expression profile of EPAS1 includes adult and fetal lung, placenta, and vascular endothelial cells. Also a protein stabilizing mutation in EPAS1 is associated with erythrocytosis, which causes and excess of red blood cells. This suggests a link between EPAS1 and red blood cell production. The SNP with the most extreme frequency difference in EPAS1 correlated with significant associations in lower erythrocyte quantities and correspondingly lower hemoglobin levels. I find it strange that a SNP that is supposed to confer greater adaptation to altitude would cause a drop in the cells and proteins capable of carrying oxygen throughout the body. The paper does present an interesting point that suggests this allele may provide carriers with the ability to maintain sufficient oxygenation at a high altitude without increasing erythrocyte levels. This could be important for preventing erythrocytosis, which can be caused by hypoxic stress on the body. Even though I feel like it was significant that the genes with strong PBS values had known roles in oxygen transport and regulation, more testing needs to be done on the alleles of this gene to see if and how they actually confer greater fitness in higher altitudes.
Some other genes were also identified that have roles in oxygen control, but based on another SNP variation study done on Andean highlanders, of all the genes identified in this paper that could play a role in altitude adaptation, only the EGLN1 gene was identified as a common gene between the populations. This result meant that Andean highlanders have adapted different genes to give them an advantage in high altitudes. This is finding is consistent with the idea that the Tibetan and Andean populations have taken very different evolutionary paths to adapt to the high altitude environments they live in, indicated by large quantitative differences in numerous physiological traits comprising the oxygen delivery process (Beall 2007).
Given that the estimate of divergence between the Han and Tibetan populations occurred 2750 years ago, the SNP at the EPAS1 gene may represent the strongest instance of natural selection documented in human populations, which would suggest that the EPAS1 gene has an important role for human survival and/or reproduction in the Tibetan region. This discovery gives us an idea about how quickly humans can adapt to survive in new climates and shows that evolution does not, like many people believe, occur at a continuous pace.
References:
Cynthia M. Beall. 2007. Two routes to functional adaptation: Tibetan and Andean high-altitude natives. PNAS. 104: 8655-8660
http://www.pnas.org/content/104/suppl.1/8655.full
Xin Yi, et al. 2010. Sequencing of 50 human exomes reveals adaptation to high altitude. Science, 329(5987): 75-78.
Scientific Article on Adaptation to High Altitude
Genomics Page
Biology Home Page
Email Questions or Comments to ermcguire@davidson.edu.
© Copyright 2011 Department of Biology, Davidson College, Davidson, NC 28035