This web page was produced as an assignment for an undergraduate course at Davidson College.
Summary
The paper in question investigates bacterial (Bacillus subtilis) transcriptome regulation in response to 104 environmental stimuli. The particular model organism was chosen due to the precense of relatively dramatic environmental changes in its natural subterranean habitat. The authors evaluted the response of a subset of approximately 3000 promoter regions in response to sigma factor signalling. They measured the transcription levels of mRNAs as a function of condition. They defined 3000 transcription units in the B. subtilis transcriptome. They found that the majority of these TUs were expressed at highly variable levels and that these expression levels were usually characterized by distinct upshifts or downshifts. However, not all transcripts were regulated preciscely. A group of 174 extended transcripts lacked termination sites and eventually produced asRNA overlap. Such overlap is present in 13% of the organism's genome. The authors hypothesize that these antisense RNAs could act as regulatory mechanisms for their respective sense RNAs.
This hypotheis was supported by eveidence that environmental stimuli trigger sigma factors which regulate the production of this asRNA. Also, two thirds of the observed variation in promoter expression was due to sigma factors which have been linked to essential environmental responses. Eighty percent of all observed asRNAs are caused by sigma factor driven transcription. Another supporting piece of evidence is that sigma factor binding sites appear to be much more highly conserved in protein-coding RNAs. This would imply that there has been some selection pressure on the maintainance of these sites, and therefore that they do in fact play a significant role in control of protein-expression by preventing translation of sense RNAs.
Figure Analysis
Figure 1*. (Nicolas et al. 2012)
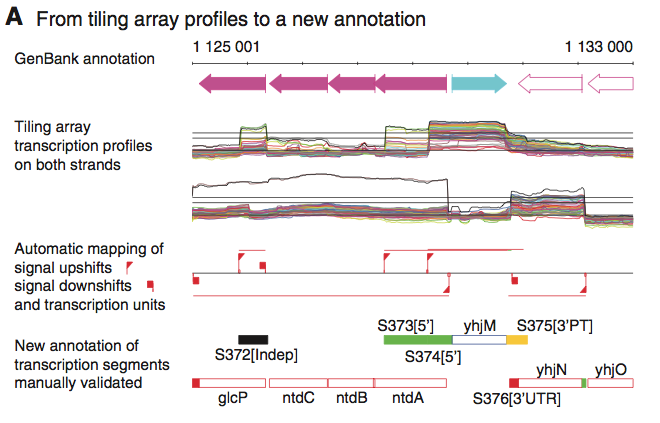
Panel A of figure 1 shows the authors' initial analysis and display of the raw data. The topmost section is an exerpt of a genetic map showing the location and direction of the transcripts in question. The second component, "Tiling array transcription profiles on both strands," display strand specific signals from 50 mRNA abundance profiles which the authors describe as representative. The lines against which the abundance profiles are displayed represnt the fold increase relative to background. From bottom to top, the lines represnt 1, 5, and 10 fold differences from background. The third section provides a simplified illustrations of the trends shown in the previous component. It shows the positions of marked and sudden changes in transcription levels which the authors descirbe as upshifts and downshifts. This panel is helpful to visualize a simplified concept of the trends shown. One possible shortcoming, however, is the depiction of an upshift on the bottom strand when it appears that only a small subset of mRNAs were transcribed at a significantly higher rate (>>10x) compared to the relatively lower transcription profiles of the other mRNAs displayed. The final panel shows the authors' annotation of new RNA features(Sxxx) against known transcripts.
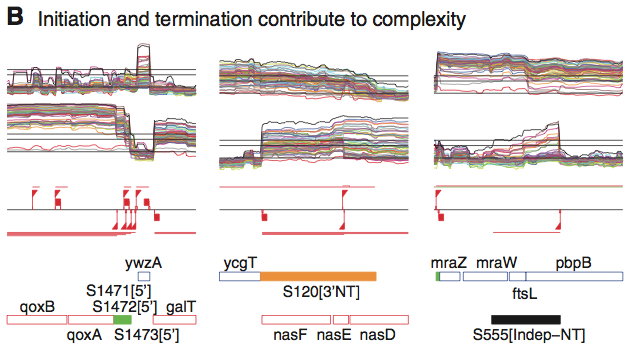
This figure shows transcription levels of several qoxAB promoters. Of particular importance are the extended transcripts S120 and S555. They are antisense to several of the other transcripts shown. These two transcripts lacked intrinsic terminators, and as such were transcribed well beyond their coding regions. The authors state that these transcripts extended up to 3.4kb beyond the normal location of the 3' termination site.
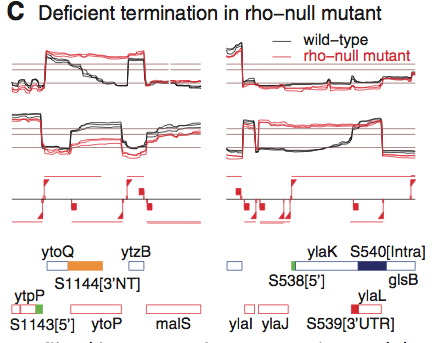
Panel C demonstrates the importance of the termination facor Rho. In the case of transcription of both ytoQ and ylaK, there is novisible downshift at the end of the coding sequence. However, this lack of termination occurs in different circumstances in each case. ytoQ has no intrinsic termination site, and the wt stain exhibits a gradual decrease in expression( due to what the authors describe as a "partially efficient terminator") whilst the rho-null strain produces an extended length stranscript until encountering the terminator at ytzB. In ylaK, on the other hand, the is a termination site, but the transcript doesn't terminate until the ylaL termination site. One point of concern in this part of the figure is the apparent termination and re-initiation of the ylaK transcript in the rho-null strain. This phenomenon is not mentioned in the paper and is puzzling to say the least. However, the figure does seem to support the authors' claim the rho is an inhibitor of antisense transcription.
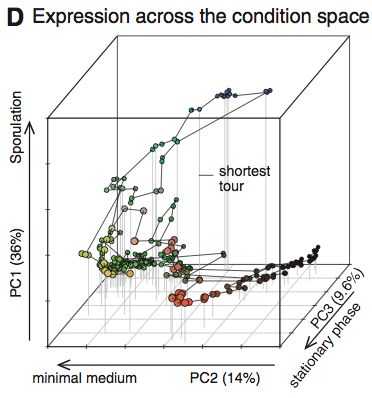
Panel D is a three-dimensional principal component analysis. It is a visualization of the effect of external environmental stimuli on the expression of the 269 transcriptomes. The authors state that these stimuli and conditions are responsible for approximately 60 percent of observed variation. They also use this figure to demonstrate the high correlation in transcription level changes in related genes.
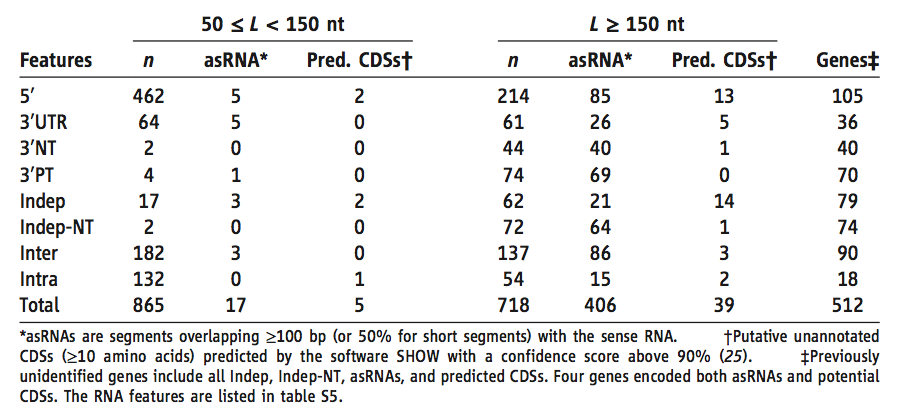
Table 1*.(Nicolas et al. 2012)
This table provides an overview of the characteristics of the previously unidentified RNA features. They are first organized by size (left vs. right column groups). They are then quantified based on the characteristics listed relative to annotated genes and if they are either antisense or previously unannotated they are counted in the appropriate columns.
Figure 2*.(Nicolas et al. 2012)
Panel A of this shoes a tree showing correlations between promoters. The corelations were determined by measuring downstream RNA signlas, and DNA sequence information was not considered. Panel B shows promoters clustered by sigma factor binding sites. There appears to be a high correlation between this grouping and that predicted by the tree in panel A. This panel also shows the high number of promoters associated with sigma factor bonding sites. Those which could not be associated are shown in grey. Panel C graphs the modeled transcription levels of the indicated clusters from panel B. The authors estimated the activity of a cluster by the average transcription signal for each individual promoter. The x-axis represents "shortest tour" conditions from the supplemental material. It is definitely a flaw in this figure that one must consult the supplemental material to fully analyse it.
Figure 3*. (Nicolas et al. 2012)
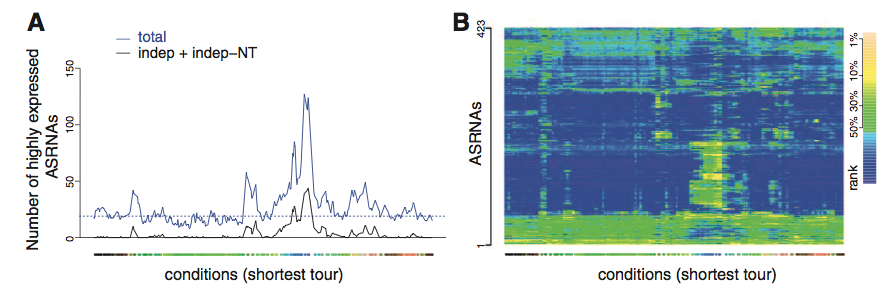
Figure 3 depicts the variation in transcription in response to conditions. Panel A shows the total number of certain asRNAs expressed at a given condition. Only the top 30% are shown, but one might assume that the others follow the same trends. Panel B is a heat map of all asRNAs characterized in table S11. Again, one cannot fully appreciate the figure without consulting supplemental material.
Reactions
Though interesting and thought-provoking, there are several aspects of the paper which deter from the overal effect. The claim by the authors that the data in figure 1c suggest that rho is an inhibitor of antisense transcription seems somewhat of a stretch of the data. Whilst the data certainly don't appear to be in contradiction, they don't immediately support the authors claim. It is true that through-transcribed regions often lead to antisense transcripts. However, this is not always the case (as shown repeatedly in the paper). While it seems certain that rho is involved in termination, it may not be a direct signalling element for asRNAs.
Perhaps the biggest difficulty with the paper is its heavy reliance on supplemental material. Granted, the authors were space constrained, but it is difficult to read a figure well if one must consult supplemental material. An ideal paper is able to stand alone to support its conclusions. It should only be necessary to consult supplemental material when one wishes to understand the article on a deeper level than necessary for basic understanding. The refferences to supplemental material in figures 2 and 3 detract significantly from their effieciency.
*Permisions Pending
References