Cell, Vol. 66, 637-647, August 23, 1991, Copyright © 1991 by Cell Press
Whitehead Institute for Biomedical Research
Howard Hughes Medical Institute
and Department of Biology
Massachusetts Institute of Technology
Cambridge, Massachusetts 02142
Summary
Segmental pattern in the Drosophila embryo is established by two maternal factors localized to the anterior and posterior poles of the egg cell. Here we provide molecular evidence that the localized posterior factor is the RNA of the nanos (nos ) gene. nos RNA is localized to the posterior pole of early embryos, and nos protein acts at a distance to direct abdomen formation. Synthetic nos RNA has biological activity identical to that of the posterior pole plasm. Injection of nos RNA rescues the segmentation defect of embryos derived from females mutant for all nine known posterior group genes. Injection of nos RNA into the anterior is able to direct formation of ectopic posterior structures. Our results demonstrate that a localized source of nos RNA is sufficient to specify abdominal segmentation and imply that other posterior group genes are required for localization, stabilization, or distribution of the nos gene product.
Introduction
Establishment of polarity in the Drosophila embryo requires maternal information that is provided to the egg cell during its maturation. Information for the establishment of the anterior-posterior axis consists of three independent morphogenetic systems that control the spatially restricted expression of target genes in the embryo (Nüsslein-Volhard et al., 1987). The anterior system controls the development of head and thorax, the posterior system, the abdominal region, and the terminal system, the most anterior and posterior larval structures (Frohnhöfer and Nüsslein-Volhard, 1986; Lehmann and Nüsslein-Volhard, 1991; Klingler et al., 1988). The anterior and posterior systems act through factors localized to the respective poles (Frohnhöfer and Nüsslein-Volhard, 1986; Lehmann and Nüsslein-Volhard, 1991). At the anterior end, the determining factor is the product of the bicoid (bcd) gene (Frohnhöfer and Nüsslein-Volhard, 1986). Localization of bad to the anterior pole requires other anterior group genes (Frohnhöfer and Nüsslein-Volhard, 1987). bcd RNA localization establishes a concentration gradient of bcd protein over the anterior half of the embryo (St Johnston et al., 1989; Driever and Nüsslein-Volhard, 1988a) that elicits proper expression of zygotic target genes such as the gap gene hunchback (hb ) (Driever and Nüsslein-Volhard, 1988b, 1989; Struhl et al., 1989).
For the posterior system, at least nine genes (nanos [nos ] pumilio [pum ] oskar [osk ], vasa [vas ] tudor [tud ], staufen [stau ] valois [vls ], cappuccino [capu ], and spire [spir ]) are required for abdomen formation (Boswell and Mahowald, 1985; Schüpbach and Wieschaus, 1986; Lehmann and Nüsslein-Volhard, 1986, 1987, 1991; Manseau and Schüpbach, 1989). Embryos derived from females mutant for any of these genes lack abdominal segments. This defect can be rescued by transplantation of cytoplasm from the posterior pole of wild-type embryos into the mutant abdomen (Lehmann and Nüsslein-Volhard, 1986, 1987, 1991). This result led to the hypothesis that the posterior pole plasm serves as the source of a signal whose function is required at a distance, in the prospective abdominal region. Synthesis of this rescuing activity during oogenesis is normal in all mutants, with the notable exception of nos mutant females (Lehmann and Nüsslein-Volhard, 1991). However, all posterior group mutant embryos, except pum , lack this activity at the posterior pole (Lehmann and Nüsslein-Volhard, 1986, 1987, 1991). These results suggested that the posterior group mutants affect a common, nos-dependent activity, synthesized during oogenesis and localized to the posterior pole of the embryo.
Although the pathway for establishment of the posterior pattern predicted the existence of a maternally provided "determinant" localized to the posterior pole, the molecular nature of such a determinant remained unclear. In this article, we describe the isolation and molecular characterization of the nos gene, and we present evidence that nos RNA is functionally equivalent to the morphogenetic activity found in the posterior pole plasm and that nos protein acts at a distance to direct abdomen development.
Results
The nos Gene
nos maps cytologically to band 91F13 on the right arm of the
third chromosome (Lehmann and Nüsslein-Volhard, 1991). The nos genomic
region was cloned by chromosomal walking (Figure 1A). Two deficiencies,
Df(3R)DI-A143, which uncovers the nos gene, and Df(3R)DI-HD28, which
complements all nos mutants (Vässin and Campos-Ortoga, 1987;
Alton et al., 1988; Lehmann and Nüsslein-Volhard, 1991), define a 20
kb region containing an essential part of nos gene function within
a 120 kb chromosomal walk. Since genetic evidence indicated that nos
function is only required maternally, the 20 kb region was analyzed
for maternally expressed transcripts. This analysis revealed a 2.5 kb SalI-XhoI
genomic DNA fragment, which hybridizes strongly to a single RNA species
of 2.4 kb present predominantly in ovaries and 0- to 2-hr-old embryos and
which hybridizes weakly in 2- to 8-hr-old embryos. No transcript is detected
in older embryos, larvae, or pupae (Figure 1 B). The developmental profile
of this transcript is consistent with the maternal mode of nos action
and parallels that of another maternal-effect gene, bcd (Berleth
et al., 1988; Figure 1B). Low levels of transcript are also detected in
males. At present there is no genetic evidence of a role for nos in
males. P element-mediated transformation of nos mutants with a 7.5
kb genomic DNA fragment (Figure 1A) that includes the 2.5 kb SalI-XhoI fragment
rescues the abdominal phenotype of nos mutants (E. Gavis and R. L.,
unpublished data). We will thus refer to the 2.4 kb transcript as nos
RNA.

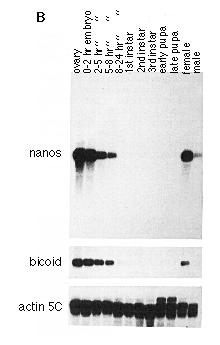
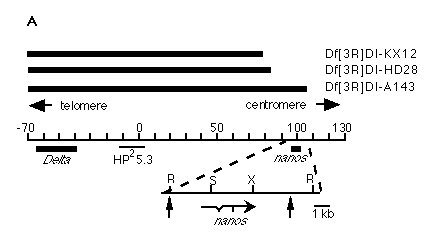
Figure 1. Molecular Cloning of nos
(A) Physical map of the nos region . Black bars at the top indicate
DNA that is absent in the indicated deficiency chromosomes; the Delta-nos
genomic region is represented by the central line; the location of the starting
clone (HP25.3) (Vässin et al., 1987) of the chromosomal walk, as well
as the locations of the Delta and nos transcribed regions,
are shown. Distances are given in kilobases with numbering starting at the
beginning of the walk. The nos region is shown in greater detail
below. Heavy line indicates 2.5 kb genomic fragment that hybridizes to nos
transcript. The 7.5 kb DNA fragment used for P element-mediated transformation
spans from an EcoRI to an SphI site as indicated by the vertical arrows.
Arrow at bottom indicates structure and 5' to 3' orientation of nos primary
transcript (restriction sites for the following enzymes are shown: R, EcoRI,
S, SalI, X, XhoI; scale as indicated).
(B) Northern blot analysis. Main panel shows nos transcript. nos
is a single species present predominantly in ovaries, early embryos,
and females. In this exposure, a weak signal is also detectable in males.
For comparison, the same blot was reprobed for bcd , a maternal transcript
that serves as a comparison for the profile of a maternal transcript (Frigerio
et al., 1986), and actin 5C, a constitutively expressed transcript (Fyrberg
et al., 1983) to show equal amounts loaded in each lane.
The 2.5 kb SalI-XhoI genomic fragment was used as a probe to screen cDNA libraries (Brown and Kafatos, 1988; Frigerio et al., 1986). The nearly full-length (2.3 kb) cDNA clone N5, as well as 3 kb of genomic DNA encompassing this cDNA, was sequenced (Figure 2). Comparison of cDNA and genomic sequences indicates that the nos primary transcript contains two introns, which are excised to produce a single 2.4 kb mRNA. The single large open reading frame encodes a protein of 400 amino acids with a predicted molecular weight of 43 kd (Figure 2). The nos polypeptide is somewhat basic (predicted pI = 9.1) and contains several stretches of polyglutamine (encoded by OPA sequences or M repeats) and polyasparagine, which are commonly found in Drosophila proteins (Wharton et al., 1985). No significant homology was found between nos and any protein listed in the latest available version of the PIR database (Altschul et al., 1990).

Figure 2. Sequence of the Predicted nos Polypeptide
This is a translation of the single large open reading frame encoded by
the nos gene. The transcriptional start site is designated as residue
1. The position of the transcriptional start was verified by primer extension
(data not shown) and lies 20 nucleotides upstream of the 5' end of the cDNA
N5. A potential TATA box sequence, ATTATT (underlined), occurs 25 nucleotides
upstream of the transcriptional start. The first AUG (underlined) begins
a short ORF, which is followed by several stop codons (the first in-frame
stop is underlined). The second AUG, at residue 263, begins the single large
open reading frame. This translational start site shows a two out of four
match with the Drosophila translation initiation consensus as determined
by Cavener (Cavener, 1987). The nos polypeptide sequence is shown
as a translation of the large open reading frame, which is interrupted by
two introns, spanning nucleotides 734-1281 and 1752-1824. The AATAAA polyadenylation
signal is underlined, and the polyadenylation site is marked by an arrow.
Distribution of nos mRNA
A central prediction of the model for establishment of the posterior pattern
is that nos should encode the posterior rescuing activity and that
the nos gene product should be localized to the posterior pole of
early embryos (Lehmann and Nüsslein-Volhard, 1991). Using a digoxigenin-labeled
nos cDNA as a probe for in situ hybridization, we find that the nos
transcript is concentrated at the posterior pole of freshly laid eggs
and cleavage stage embryos (stage 2; Figure 3A). At the pole bud stage (stage
3), the transcript segregates into the nascent pole cells (Figure 3B). The
transcript remains concentrated in the pole cells during blastoderm stages
(stages 4 and 5; Figure 3C). During gastrulation and germband extension,
when the pole cells are carried dorsally and internalized into the embryo,
staining for nos RNA is visible in the pole cells, although reduced
in intensity relative to earlier stages (stages 7 and 8, Figure 3D). While
the transcript is still visible in the pole cells at the fully extended
germband stage (stage 10, Figure 3E), localized staining is no longer detected
once the pole cells have left the pocket formed by the posterior midgut
invagination (stage 11, Figure 3F). In summary, the developmental profile
of nos RNA detected by in situ hybridization to embryos and by RNA
blot analysis suggests that maternally provided nos transcript is
present during oogenesis and early stages of embryogenesis and that transcript
localization is confined to the pole plasm and pole cells.
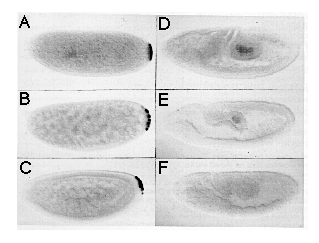
Figure 3. Whole-Mount In Situ Hybridization
The cDNA N5 (see Experimental Procedures) was used to prime synthesis of
a uniformly labeled digoxigenin-UTP probe, which was hybridized to wild-type
embryos.
(A) Early cleavage stage embryo (stage 2, ~0.5-1.3 hr). The probe recognizes
a transcript that is localized to the posterior pole.
(B) Embryo at pole bud stage (stage 3, ~1.3-1.5 hr). The transcript is concentrated
in the pole cells.
(C) Embryo at syncytial blastoderm stage (stage 4, ~1.5-2.5 hr). The transcript
is restricted to the pole cells and lacking from the blastoderm, which will
give rise to the somatic tissues.
(D) Embryo during early germband extension (stage 8, ~3.75-4.5 hr). Pole
cells are carried inside the embryo and are clustered inside a pocket of
cells that will give rise to the posterior midgut. Reduced levels of nos
RNA are detected in the pole cells.
(E) Embryo with fully extended germband (stage 9, ~4.5-5.15 hr). By this
stage most pole cells have migrated through the midgut epithelium. Only
a few pole cells are found within the midgut pocket, and only those show
residual staining.
(F) Embryo just prior to germband retraction (stage 10, ~5.15-6.25 hr).
No specific staining for nos RNA can be detected. In all cases anterior
is left, and dorsal is up. The final staining reaction was carried out longer
in embryos shown in (D)-(F) than in those shown in (A)-(C). The developmental
stages are as described in Campos-Ortega and Hartenstein (1985), and the
duration of stages at room temperature (22šC) is according to Wieschaus
and Nüsslein-Volhard (1986).
Synthetic nos RNA Is Biologically Active and Depletes Maternal
hb in Embryos
The localization of nos RNA at the posterior pole of early embryos
and the presence of nos RNA in the pole cells suggests that nos
RNA is responsible for the rescuing activity assayed in transplantation
experiments (Lehmann and Nüsslein-Volhard, 1986). If nos RNA
is the rescuing activity found in the pole plasm, synthetic nos transcript
should provide rescuing activity in an embryo injection assay. Sense strand
RNA was synthesized in vitro from the nos cDNA, N5. This RNA was
injected into the prospective abdominal region of cleavage stage (stage
2) embryos derived from nosL7 homozygous
mutant females. Uninjected nos mutant embryos lack all abdominal
segments, as seen by the lack of ventral abdominal dentate belts (Figure
4A). By contrast, sibling embryos injected with nos RNA show rescue
of the abdominal segmentation defect (Figures 4B and 4C). Completely rescued
embryos are indistinguishable from wild type, and they hatch and develop
into normal, fertile adult flies.
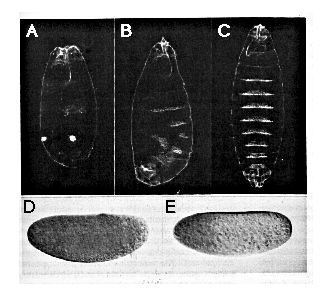
Figure 4. Rescue of the nos Mutant Phenotype by
In Vitro Synthesized nos Transcript
(A-C) Dark-field photographs of cuticle preparations of first instar larvae
derived from females homozygous for nos L7. (A) Uninjected
embryo, anterior up, dorsal aspect. Note lack of abdominal segments. Head
and thorasic segments are normal. (B) Partially rescued embryo, anterior
up, ventral aspect. This embryo shows bilaterally asymmetric rescue and
has developed six partial abdominal segments. (C) Fully rescued embryo,
anterior up, ventral aspect. This pattern is indistinguishable from wild
type.
(D-E) Embryos injected with nos RNA and stained for hb protein
using an anti-hb antibody. Embryos shown are at late cleavage stage (stage
2), anterior left, dorsal up. (D) shows embryo injected with RNA synthesized
from a template containing a frameshift mutation. (E) shows embryo injected
with RNA from a template containing wild-type nos RNA sequences.
The control embryo (D) shows uniform levels of hb protein, while
the embryo injected with functional nos RNA (E) shows reduced levels
of hb protein in the posterior but shows normal levels in the anterior
third. In some of the control embryos, we detected slightly reduced levels
of hb protein at the posterior pole, which may indicate that the
nos L7 mutation is not a complete lack of function mutation.
The role of nos in abdominal segmentation is mediated by the maternally provided product of the hb gene (Hülskamp et al., 1989; Irish et al., 1989; Struhl, 1989). In wild-type embryos, nos is required to exclude hb protein from the posterior half of the embryo (Tautz, 1988). The resulting gradient of maternal hb allows proper expression of abdominal gap genes such as knirps (kni) and giant (gt ) (Hülskamp et al., 1990; Eldon and Pirrotta, 1991; Kraut and Levine, 1991). To see whether the phenotypic rescue observed correlated with a change in the expression pattern of the maternal hb protein, we stained injected embryos with an antibody directed against hb (Figures 4D and 4E). RNA synthesized from the N5 nos cDNA template was injected into embryos derived from homozygous nosL7 females under the same conditions as described for phenotypic rescue (see above). After injection, the embryos were incubated for 1 hr and assayed for hb protein. These embryos were compared with embryos treated identically but injected with a nos RNA containing a frameshift mutation at amino acid 51 (Table 1). Uninjected or frameshift-injected (Figure 4D) embryos show a uniform pattern of hb protein staining at pole bud stage. By contrast, embryos injected with wild-type nos transcript lack detectable hb protein in the posterior region (Figure 4E). In most of these embryos, hb protein can be seen at the anterior, farthest from the site of injection (25%-50% egg length, where 0% is the posterior pole). Of 22 embryos injected with wild-type nos RNA and stained for hb protein, 5% showed homogeneous staining, 90% showed only anterior staining, and 5% gave no detectable staining. For comparison, of 19 embryos injected with nos frameshift RNA, 84% showed homogeneous staining, and 16% showed anterior staining. These results demonstrate that nos RNA depletes hb protein from the vicinity of the site of injection and strongly suggest that the phenotypic rescue of embryos by nos RNA injection is mediated by this interaction.
nos RNA Is Equivalent to the Posterior Rescuing Activity
The degree of rescue by nos RNA depends on the concentration of the
injected RNA. Rescue was tested over a 4-fold range in concentration (Table
1). At the lowest concentration tested, most embryos show only partial rescue
(Figure 4B). At the highest concentration tested, all embryos are completely
rescued and form all eight abdominal segments (Figure 4C). RNA transcribed
from a template containing a frameshift mutation fails to rescue the abdominal
defect when assayed by injection (Table 1). These results indicate that
the N5 cDNA contains a functional nos open reading frame and that
translation of nos RNA after egg deposition is sufficient to fully
restore the mutant defect.
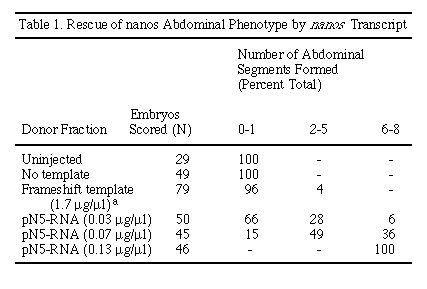
a Of the 79 cuticles scored, 3 developed more than one segment. Of these, 2 had two segments, and 1 cuticle had three segments. Note that the concentration of frameshift transcript injected was about 10-fold higher than the concentration of wild-type transcript used. We interpret the low degree of rescue observed as a reflection of some low level of readthrough translation or a residual activity of the 50 amino acid amino-terminal polypeptide.
The degree of rescue by nos RNA depends on the age of the mutant embryo and the position of injection. Injection of nos RNA into nos mutant embryos results in optimal rescue when embryos are injected before pole cell formation (Figure 5A). We conclude that nos function is required early in embryogenesis, at the time when the concentration gradient of maternal hb product is established (Tautz, 1988). Furthermore, optimal rescue is achieved if nos RNA is injected into the prospective abdominal region (25%-50% egg length) (Figure 5B), indicating that the injected RNA is translated at the site of injection and does not require localization to the posterior pole for translation. In summary, the activity of synthetic nos RNA is equivalent to the rescuing activity found in posterior pole plasm with respect to the spatial and temporal parameters of rescue (Lehmann and Nüsslein-Volhard, 1986, 1991).
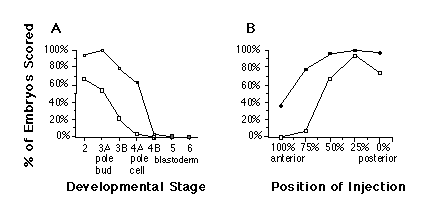
Figure 5. Developmental Stage and Position Dependence of
Rescue
"Strong rescue" (open squares) indicates more than five abdominal
segments, and "rescue" (solid squares) indicates two or more abdominal
segments, as judged by the presence of abdominal ventral denticle belts.
Fifteen to 80 cuticles were scored for each data point.
(A) Developmental stage dependence. Staging of recipient embryos as in Campos-Ortega
and Hartenstein, 1985 (stages 1 and 2, cleavage stage; stage 3, pole bud
formation; stage 4, syncytial blastoderm; stage 5, cellular blastoderm;
stage 6, beginning of gastrulation). Developmental events are indicated
along horizontal axis.
(B) Position dependence. Cleavage stage embryos were injected at the positions
shown along the horizontal axis.
Synthetic nos RNA Rescues Abdominal Phenotype of All Posterior
Group Mutants
Embryos derived from females defective for seven of the nine posterior group
genes lack the specialized posterior pole plasm, including the characteristic
polar granules, and subsequently fail to form pole cells, the germline precursors
(Lehmann and Nüsslein-Volhard, 1986; Schüpbach and Wieschaus,1
986; Boswell and Mahowald, 1985; Manseau and Schüpbach, 1989). Cytoplasmic
transplantation experiments have shown that these embryos lack posterior
rescuing activity (Lehmann and Nüsslein-Volhard, 1986, 1991). To demonstrate
the exclusive role of nos as a localized posterior determinant, we
tested the ability of nos RNA to rescue the abdominal phenotype of
all posterior group mutants. Injection of nos RNA completely rescues
the abdominal phenotype of the genes osk , stau , tud ,
vas, vls, capu, and spir (Table 2). The germ
cell defect, however, remains. These results indicate that nos acts
as a posterior determinant and suggest that these posterior group genes
are dispensable for the translation and/or activity of nos protein.
Since embryos from mothers mutant for these genes lack the specialized posterior
pole plasm, we propose that localization of nos mRNA to the posterior
pole depends on the presence of an intact pole plasm. The abdominal segmentation
defect of these mutants may therefore be a secondary effect of failure to
localize or stabilize pole plasm components, including the nos mRNA.
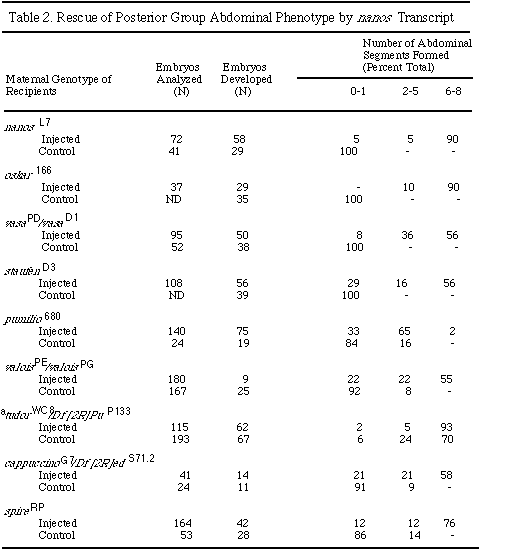
"Control" indicates phenotype of uninjected embryos.
ND, not determined.
a No tudor alleles with complete penetrance for the abdominal phenotype
have been identified. For the tudor embryos, the phenotypic scores
for control and injected populations were compared by the Wilcoxon-Mann-Whitney
ranks test and shown to be significantly different with P < 0.004.
Embryos derived from pum mutant mothers have intact pole plasm and form germ cells. Cytoplasmic transplantation experiments have shown that the posterior pole plasm of pum embryos contains normal levels of rescuing activity for all posterior group mutants (Lehmann and Nüsslein-Volhard, 1987, 1991). pum mutant embryos injected with nos RNA show rescue of the abdominal segmentation phenotype. However, rescue is limited to six abdominal segments or less (Table 2), and segmentation is restored only locally, near the site of injection (data not shown). We conclude that pum affects the ability of nos to act at a distance on abdomen formation and propose that pum affects the distribution of nos protein, rather than the localization of nos RNA. Additional experiments are required to determine whether pum function involves intracellular transport, stabilization, abundance, or activity of nos protein.
Presence of nos at the Anterior Suppresses bcd
Previous studies have shown that transplantation of posterior pole plasm
to an ectopic anterior position suppresses the formation of head and thorasic
structures and directs the formation of a second abdomen at the anterior
end of the embryo (Nüsslein-Volhard et al., 1987; Frohnhöfer et
al., 1986). If this respecification is due to nos, injection of nos
RNA into the anterior of embryos should suppress anterior development
and cause development of ectopic posterior structures. We injected synthetic
nos RNA into the prospective head region of nos mutant embryos
and followed their development (Figure 6).
The cephalic furrow, which normally forms during gastrulation (stages 6-7) at 65% egg length, serves as a marker for the anterior region of the embryonic fate map. In many cases, injected embryos show an anterior shift in the position of the cephalic furrow, and cuticle preparations of first instar larvae reveal a reduction of head skeletal structures. In more extreme instances, injected embryos lack a cephalic furrow altogether, and an ectopic posterior midgut invagination is observed at the anterior end (Figures 6A and 6E). The result of this extreme fate map shift is seen more clearly in cuticle preparations. In the case shown, head structures are completely absent, and structures normally found at the posterior, including a telson and several abdominal denticle belts, are duplicated in mirror image at the anterior ("bicaudal" phenotype; Figure 6F). Of 57 embryos injected anteriorly and scored for cuticle phenotype, 32% had normal head structures, 30% showed reduction of head structures, 33% lacked head structures altogether, and 5% gave a bicaudal phenotype that included reversal of abdominal segments. These results demonstrate that anteriorly introduced nos RNA is sufficient to suppress formation of head and thorasic structures and is able to induce the formation of posterior structures.
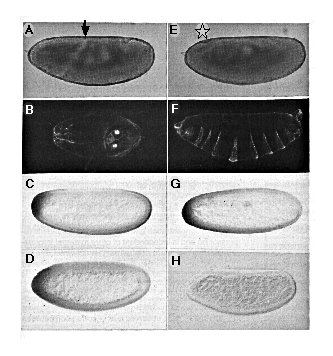
Figure 6. Ectopically Introduced nos Alters Embryonic
Pattern
(A-D) Control embryos derived from nos L7 mutant
females ([A] and [B], uninjected; [C] and [D], injected with in vitro synthesized
nos RNA containing a frameshift mutation) show normal head development
([A] and [B]) and wild-type pattern of bcd RNA (C) and bcd protein
(D) distribution .
(E-H) Embryos derived from nosL7 mutant females
injected with RNA synthesized from a wild-type nos cDNA template
develop posterior structures at the expense of head structures ([E] and
[F]) and show normal levels and distribution of bcd RNA (G), while
levels of bcd protein are reduced (H).
Top panel ([A] and [E]): living embryos during gastrulation (stage 7); the
position of the cephalic furrow is indicated in (A) by an arrow. Star in
(E) indicates the position of an ectopic posterior midgut invagination at
the anterior. Second panel ([B] and [F]): dark-field photographs of cuticle
preparations from first instar larvae developed from embryos similar to
those shown above. Embryo in (F) shows a bicaudal phenotype such that the
telson and posterior abdomen is duplicated in mirror image at the expense
of head and thorasic structures. Third panel ([C] and [G]): syncytial blastoderm
(stage 4) embryos stained for bcd RNA. Bottom panel ([D] and [H]):
syncytial blastoderm (stage 4) embryos stained for bcd protein. In all cases
anterior is to the left, dorsal up. All embryos were injected at 75% egg
length.
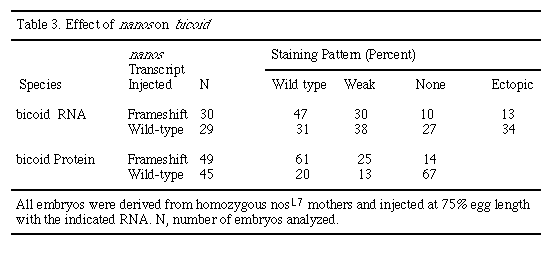
The suppression of head structures observed in nos-injected embryos suggests that nos affects genes specifying head and thorasic structures. These anterior defects resemble those of bcd and bicaudal mutants (Frohnhöfer and Nüsslein-Volhard, 1986; Nüsslein-Volhard, 1977). In no case were phenotypes characteristic of zygotic hb mutants, such as thorasic segment deletions, produced. To further distinguish between a direct effect of nos on bcd and an effect of nos on the zygotic hb product, which is under the control of bcd (Driever and Nüsslein-Volhard, 1989; Struhl et al., 1989), we stained embryos with antibodies against bcd protein (Driever and Nüsslein-Volhard, 1988a). At the syncytial blastoderm stage (stages 3-4), levels of bcd protein are reduced in embryos injected with nos RNA, whereas normal levels of bcd protein are found in embryos injected with a frameshifted nos RNA (Figures 6D and 6H; Table 3). In addition, suppression of anterior structures is only observed after nos RNA injection in regions where bcd RNA is normally localized (data not shown). In situ hybridization analysis of injected embryos indicates that the stability and distribution of bcd mRNA is not significantly affected by the ectopic introduction of nos (Figures 6C and 6G; Table 3). These results taken together suggest that nos can affect the synthesis and/or stability of bcd protein.
Discussion
Previous genetic studies led to identification of nine genes that are required for normal abdomen formation in Drosophila (Boswell and Mahowald, 1985; Schüpbach and Wieschaus, 1986; Lehmann and Nüsslein-Volhard, 1986, 1987, 1991; Manseau and Schüpbach, 1989). These studies revealed a factor concentrated in the pole plasm of wild-type embryos that rescues the abdominal defect of posterior group mutants (Lehmann and Nüsslein-Volhard, 1991). It was suggested that nos is involved in the synthesis of this factor (Lehmann and Nüsslein-Volhard, 1991). In this article, we have demonstrated that the nos gene product is the rescuing factor. nos mRNA is localized at the posterior pole and serves as a source of the nos protein, which acts as a signal in the prospective abdomen. Synthetic nos RNA is biologically active and rescues the abdominal phenotype of nos and all other posterior group mutant embryos. This result shows that the other posterior group genes are not required for nos function but rather play a role in localization or distribution of nos gene products. Finally, transplantation of nos RNA to the anterior suppresses head and thorasic development and promotes the formation of posterior structures in reversed orientation at the anterior end. These results indicate that a localized source of nos product is sufficient to specify abdominal segmentation.
How does the nos product determine posterior pattern? The anterior, posterior, and terminal systems of maternal genes regulate the spatial domains of gap gene transcription in the embryo (Nüsslein-Volhard et al., 1987; Ingham, 1988). For the posterior system, nos activity is required for the correct embryonic transcription of at least three gap genes: Krüppel (Kr ), kni , and gt (Gaul and Jäckle, 1987; Nauber et al., 1988; Eldon and Pirrotta, 1991; Kraut and Levine, 1991). However, nos does not regulate embryonic transcription directly. nos negatively regulates the abundance of the maternally provided hb gene products such that a concentration gradient of hb RNA and hb protein is established, with highest concentrations in the anterior (Tautz, 1988). Maternally provided hb itself is believed to act as a transcriptional repressor, such that low levels of hb in the abdomen allow proper expression of the gap genes Kr , kni , and gt (Hülskamp et al., 1989, 1990; Irish et al., 1989; Struhl, 1989).
Changes in the levels of nos along the anterior-posterior axis shift the embryonic fate map (Lehmann, 1988) and can alter the polarity of the segmental pattern (Lehmann and Nüsslein-Volhard, 1991), presumably by affecting the maternal hb concentration gradient (Tautz, 1988; Wharton and Struhl, 1989). However, recent experiments indicate that, under some conditions, the maternal hb gradient is dispensable for the formation of a normal segmental pattern (Hülskamp et al., 1989; Irish et al., 1989; Struhl, 1989). Embryos that lack nos and maternal hb products, thereby completely lacking maternally provided posterior information, develop with a normal segmentation pattern (Hülskamp et al., 1989; Irish et al., 1989; Struhl, 1989), which probably reflects normal expression of gap genes. Segmentation in such embryos must operate by a second, "nos-independent" pathway. Gap gene expression in the abdomen of these embryos is set by neighboring gap genes, whose spatial domains of expression depend in turn on the anterior and terminal maternal systems (Hülskamp et al., 1990). Thus, in the abdomen, two redundant systems can establish the spatial expression of gap genes, which control segmentation and polarity. The nos system acts via a concentration gradient of the transcriptional repressor hb and is independent of other maternal information (Nüsslein-Volhard et al., 1987). By contrast, the zygotic system acts via gap gene interactions established by positional cues provided by the anterior and terminal maternal genes (Lehmann and Frohnhöfer, 1989; Hülskamp et al., 1990).
This feature of the abdominal pattern forming system may account for the observed lack of correlation between the local concentration of nos and the abdominal structures formed. For example, injection of nos RNA into nos mutant embryos at various positions results in the formation of abdominal segments of normal polarity (Figure 4; Lehmann and Frohnhöfer, 1989). According to the model, nos removes the repressor hb , such that kni and gt can be expressed in the abdominal region. The relative position of kni and gt expression, however, is determined by interaction with neighboring gap genes (Lehmann and Frohnhöfer, 1989). This model for posterior pattern formation can also explain how injection of nos RNA into the anterior of a nos mutant embryo can result in the formation of two abdomens in mirror image (Figure 6): first, nos suppresses bcd function in the anterior (see below); then, ectopically introduced nos eliminates maternal hb throughout the embryo. This allows expression of abdomen-specific gap genes in both the anterior and posterior halves of the embryo. The polarity of the resulting abdomens is most likely set by the terminal genes (Lehmann and Frohnhöfer, 1989), which are active at either end (Pignoni et al., 1990).
Transplantation of nos RNA to the anterior not only specifies the formation of abdominal segments but also suppresses formation of head and thorasic structures. This suggests that nos can negatively regulate bcd , the anterior determinant. Indeed, antibody staining of injected embryos shows that bcd protein levels are decreased after injection of nos RNA in the anterior (Figure 6). In addition, in situ hybridization analysis of injected embryos shows that the stability and localization of bcd RNA is not significantly affected (Figure 6). This suggests that nos acts on bcd at the level of translation or protein stability.
We favor the idea that nos regulates the translation of hb and bcd , since the presence of nos has a pronounced effect on levels of the hb and bcd proteins, whereas the effect of nos on hb and bcd RNA levels is more subtle and appears to occur later (Driever and Nüsslein-Volhard, 1988b; Tautz, 1988; Wharton and Struhl, 1989; Table 3). One possibility is that nos protein binds to hb and bcd RNA, decreases their rate of translation, and ultimately these transcripts are degraded. Evidence from experiments involving replacement of the hb 3' region with the lacZ gene (Hülskamp et al. , 1989) and examination of the bcd RNA in genetically bicaudal embryos (Wharton and Struhl, 1989) suggests that targets of nos action reside in the 3' regions of the bcd and hb RNAs. Analysis of the nos protein sequence, however, has not revealed homology to sequences known to be involved in RNA binding (Bandziulis et al., 1989). Additional experiments are necessary to determine whether nos protein directly binds bcd and hb RNA and to identify sequences within the nos protein required for such an interaction.
Experimental Procedures
Cloning of nos
The nos genomic region is defined by the chromosomal deficiencies
Df[3R]DI-HD28 and Df[3R]DI-KX12, which complement all nos mutants,
and Df[3R]DI-A143, which uncovers the nos gene (Vässin and Campos-Ortega,
1987; Alton et al., 1988; Lehmann and Nüsslein-Volhard, 1991). The
region defined by the Df[3R]DI-A143 and Df[3R]DI-KX12 interval contains,
in addition to nos, three lethal complementation groups: I(3)91Fb
(= I(DIX43)c3), I(3)91Fe (= I(DIX43)c4), and I(3)91Fc (Vässin and Campos-Ortega,
1987; Alton et al., 1988). Representative alleles of each of these complementation
groups complement nos mutants. Two lambda phage genomic libraries
(in Charon 4 and EMBL 4 vectors) were used for the chromosomal walk, which
was carried out according to standard procedures (Sambrook et al., 1989).
The direction and progress of the walk was monitored by in situ hybridization
of biotinylated phage DNA to salivary gland chromosomes of larvae heterozygous
for the deficiencies Df[3R]DI-A143, Df[3R]DI-KX12, and Df[3R]DI-HD28 (Vässin
and Campos-Ortega, 1987; Alton et al., 1988). The nos region was
further mapped by blot hybridization of DNA prepared from flies heterozygous
for the deficiencies Df[3R]DI-A143 and Df[3R]DI-HD28 with representative
clones from the walk (F. Pelegri and R. L., unpublished data). To identify
the genomic region encoding nos, radiolabeled cDNA from specific
stages was hybridized to a blot of genomic DNA from the walk. By this analysis,
the 2.5 kb fragment described hybridizes to the most strongly expressed
maternal transcript in this region (data not shown).
Northern Blot Analysis
Poly(A)+ RNA prepared from the indicated stages (1.5 µg
per lane) was fractionated on a 1.2% agarose-formaldehyde gel. After transfer
to nylon membranes, the filter was ultraviolet cross-linked and probed with
the appropriate 32P-labeled DNA fragment. The 2.5 kb SalI-XhoI
DNA fragment indicated in Figure 1A was used as the nos probe. Blots
were exposed to X-ray film with an intensifying screen for 48, 24, or 6
hr, for nos, bcd , or actin, respectively. Preparation of
poly(A)+ RNA as well as hybridization and washing of blots was
done by standard methods (Sambrook et al., 1989).
cDNA Cloning and DNA Sequencing
The 2.5 kb SalI-XhoI genomic DNA fragment described in Figure 1A was labeled
with [32P]dCTP by random primer oligolabeling and used to screen
both lambda and plasmid cDNA libraries. Three independently isolated cDNA
clones were subcloned into Bluescript vectors (Stratagene) and nested deletions
were made using exonuclease III and mung bean nuclease according to manufacturers'
protocols (Stratagene). These deletions were sequenced using Sequenase (United
States Biochemical). The genomic DNA was sequenced using sequence-specific
oligonucleotide primers, or, for some regions, by subcloning small fragments.
In all cases, sequence was confirmed on the opposite strand.
In Situ Hybridization
In situ hybridization was performed as described previously by Tautz and
Pfeifle (1989).
RNA Injections
RNA for injection assays was synthesized in vitro from the full-length cDNA
clone N5, which includes a 43 nucleotide ß-globin 5' untranslated
leader sequence fused to the complete nos open reading frame (Brown
and Kafatos, 1988). Methods for RNA synthesis were as described by Krieg
and Melton (1984). The in vitro transcription products were ethanol precipitated
and resuspended in diethyl pyrocarbonate-treated water. Concentrations were
determined by optical density measurement at 260 and 280 nm and are corrected
for the presence of residual template DNA. Since the level of active transcription
products obtained varies, the product of a single transcription reaction
was used for any given experiment. "No template" embryos were
injected with the product of an in vitro transcription reaction from which
template DNA was omitted. The frameshift template was constructed by filling
in a PstI site at nucleotide 406 of the insert, followed by addition of
a BamHI linker and religation, creating a frameshift mutation after amino
acid 50 of the open reading frame.
Injections and cuticle preparations were carried out as previously described (Lehmann and Nüsslein-Volhard, 1986, 1991). All embryos for injection were derived from nosL7 mutant females and injected at early cleavage stage (stage 2) at 25%-50% egg length unless otherwise indicated. Final concentration of donor RNA was 2 µg/µl in diethyl pyrocarbonate-treated water. In general, 50%-80% of injected embryos developed a cuticle and could be scored for segmentation phenotype. For the injections into other posterior group mutants (Table 2), all allele combinations used were the strongest available for the abdominal segmentation phenotype (Boswell and Mahowald, 1985; Schüpbach and Wieschaus, 1986; Lehmann and Nüsslein-Volhard, 1986, 1987, 1991; Manseau and Schüpbach, 1989). In experiments in which rescued embryos were allowed to develop to adulthood, both heterozygous and homozygous progeny were obtained. For the experiment in Figure 6, embryos were injected at 75% egg length, observed through gastrulation by phase microscopy, and photographed; they were then allowed to develop through the end of embryogenesis and mounted for scoring of cuticle phenotype. Staging of embryos was as described by Campos-Ortega and Hartenstein, 1985.
hb and bcd Antibody Staining of Injected Embryos
Embryos derived from nosL7 mutant females were injected
with the appropriate RNA at 25%-50% egg length for the hb stainings
and at 75% egg length for the bcd stainings. Injected embryos were
aged at 18šC for 1 to 1.5 hr after injection, then fixed at pole bud
and syncytial blastoderm stages (stages 3 and 4) and devitellinized by hand.
Antibody distribution was detected using a biotinylated anti-mouse secondary
antibody and a commercially available horseradish peroxidase detection kit
(Vector labs).
In Situ Hybridization of Injected Embryos
Injected embryos were prepared, fixed, and devitellinized as described above
for the antibody staining, then hybridized with a bcd cDNA probe
for whole-mount in situ hybridization as described by Tautz and Pfeifle
(1989).
Acknowledgments
We thank H. Vässin for the clone HP25.3, M. Noll, N. Brown, and J. M. Dura for libraries, W. Driever for bcd antibody, and D. Tautz for hb antibody. We thank F. Pelegri for deficiency breakpoint mapping, L. Dickinson for in situ hybridization analysis, and D. Curtis for providing the embryos shown in Figures 3D-3F. We also thank S. Burden, G. Fink, T. Orr-Weaver, and D. Rio and members of the laboratory for helpful comments on the manuscript. R. L. thanks M. Wilsox and other members of the LMB, MRC in Cambridge, England for advice and support. This work was supported by an NSF predoctoral fellowship (C. W.), the NIH, and the HHMI.
The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked "advertisement" in accordance with 18 USC Section 1734 solely to indicate this fact.
Received April 9, 1991; revised June 10, 1991.
References
Alton, A. K., Fechtel, K., Terry, A. L., Meikle, S. B., and Muskavitch, M. A. T. (1988). Cytogenetic definition and morphogenetic analysis of Delta , a gene affecting neurogenesis in Drosophila melanogaster . Genetics 118, 235-245.
Altschui, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403-410.
Bandziulis, R. J., Swanson, M. S., and Dreyfuss, G. (1989). RNAbinding proteins as developmental regulators. Genes Dev.3,431 -437.
Berleth, T., Burri, M., Thoma, G., Bopp, D., Richstein, S., Frigerio, G., Noll, M., and N0sslein-Volhard, C. (1988). The role of localization of bicoid RNA in organizing the anterior panern of the Drosophila embryo. EMBO J . 7, 1749-1756.
Boswell, R. E., and Mahowaid, A. P. (1985). tudor, a gene required for assembly of the germ plasm in Drosophila melanogaster . Cell 43,97104,
Brown, N. H., and Kafatos, F. C.,(1988). Functional cDNA libraries from Drosophila embryos. J. Mol. Biol. 203, 425-437.
Campos-Ortega, J. A., and Hartenstein, V. (1985). The Embryonic Development of Drosophila melanogaster (Heidelberg: SpringerVerlag).
Cavener, D. R. (t 987). Comparison of the consensus sequences flanking translational start sites in Drosophila and vertebrates. Nucl. Acids Res. 15, 1353-1361.
Driever, W., and Nüsslein-Volhard, C. (1988a). A gradient of bicoid protein in Drosophila embryos. Cell 54, 83-93.
Driever, W., and Nüsslein-Volhard, C. (1988b). The bicoid protein determines position in the Drosophila embryo in a concentrationdependent manner. Cell 54, 95-104.
Driever, W., and Nüsslein-Volhard, C. (1989). The bicoid protein is a positive regulator of hunchback transcription in the early Drosophila embryo. Nature 337, 138-143.
Eldon, E. D., and Pirrota, V. (t 99t ). Interactions of the Drosophila gap gene giant with maternal and zygotic pattern-forming genes. Development 111, 367-378.
Frigerio, D., Burri, M., Bopp, D., Baumgartner, S., and Noll, M. (1986). Structure of the segmentation gene paired and theDrosophila PRD gene set as part of a gene network. Cell 47, 735-746.
Frohnhöfer, H. G., and Nüsslein-Volhard, C. (1986). Organization of the anterior pattern in the Drosophila embryo by the maternal gene bicoid . Nature 324, t20-t25.
Frohnhöfer, H. G., and Nüsslein-Volhard, C. (1987). Maternal genes required for the anterior localization of bicoid activity in the embryo of Drosophila . Genes Dev. 1, 880-890.
Frohnhöfer, H. G., Lehmann, R., and Nüsslein-Volhard, C. (1986). Manipulating the anteroposterior panern of the Drosophila embryo. J. Embryol. Exp. Morphol. (Suppl.) 97, 169-179.
Fyrberg, E. A., Mahaffey, J. W., Bond, B. J., and Davidson, N. (1983). Transcripts of the six Drosophila actin genes accumulate in a stageand tissue-specific manner. Cell 33, 115-123.
Gaul, U., and Jäckle, H. (1 987). Pole region-dependent repression of the Drosophila gap gene Krüppel by maternal gene products. Cell 51, 549-555.
Hülskamp, M., Schroder, C., Pfeifle, C., J§ckle, H., and Tautz, D. (1989). Posterior segmentation of the Drosophila embryo in the absence of a maternal posterior organizer gene. Nature 338, 629-632.
Hülskamp, M., Pfeifle, C., andTautz, D. (1990). Amorphogeneticgradient of hunchback protein organizes the expression of the gap genes Krippel and knirps in the early Drosophila embryo. Nature 346, 577580.
Ingham, P. W. (1988). The molecular genetics of embryonic pattern formation in Drosophila . Nature 335, 25-34.
Irish, V., Lehmann, R., and Akam, M. (1989). The Drosophila posterior group gene nanos functions by repressing hunchback activity. Nature 338, 646-648.
Klingler, M., Erdeleyi, M.; Szabad, J ., and Nüsslein-Volhard, C. (1 988). Function of torso in determining the terminal anlagen of theDrosophila embryo. Nature 335, 275-277.
Kraut, R., and Levine, M. (1991). Spatial regulation of the gap gene giant during Drosophila development. Development 111, 601-609.
Krieg, P. A., and Melton, D. A. (1984). Functional messenger RNAs are produced by SP6 in vitro transcription of cloned cDNAs. Nucl. Acids Res. 12, 7057-7070.
Lehmann, R. (1988). Phenotypic comparison between maternal and zygotic genes controlling the segmental pattern of the Drosophila embryo. Development (Suppl.) 104, 17-27.
Lehmann, R., and Frohnhöfer, H. G. (1989). Segmental polarity and identity in the abdomen of Drosophila is controlled by relative position of gap gene expression. Development (Suppl.) 107, 21-29.
Lehmann, R., and Nüsslein-Volhard, C. (1986). Abdominal segmentation, pole cell formation, and embryonic polarity require the localized activity of oskar , a maternal gene in Drosophila . Cell 47, 141-152.
Lehmann, R., and Nüsslein-Volhard, C. (1987). Involvement of the pumilio gene in the transport of an abdominal signal in the Drosophila embryo. Nature 329, 167-170.
Lehmann, R., and Nüsslein-Volhard, C. (1991). The maternal gene nanos has a central role in posterior pattern formation of the Drosophila embryo. Development 112, 679-693.
Manseau, L. J., and Schüpbach, T. (1989). cappuccino and spire: two unique maternal-effect loci required for both the anteroposterior and dorsoventral panerns of the Drosophila embryo. Genes Dev. 3,14371452.
Nauber, U., Pankratz, M., Kienlin, A., Seifert, E., Klemm, U., and Jäckle, H. (1988). Abdominal segmentation of the Drosophila embryo requires a hormone receptor-like protein encoded by the gap gene knirps. Nature 336, 489-492.
Nüsslein-Volhard, C. (1977). Genetic analysis of pattern-formation in the embryo of Drosophila melanogaster . Roux's Arch. Dev. Biol. 183, 249-268.
Nüsslein-Volhard, C., Frohnhöfer, H. G., and Lehmann, R. (1987). Determination of anteroposterior polarity in Drosophila . Science 238, 1675-1681.
Pignoni, F., Baldarelli, R. M., Steingrimsson, E., Diaz, R. J., Patapoutian, A., Merriam, J. R., and Lengyel, J . A. (1990). The Drosophila gene tailless is expressed at the embryonic termini and is a member of the steroid receptor superfamily. Cell 62, 151-163.
Sambrook, J., Fritsch, E. F., and Maniatis, T. (1989). Molesular Cloning: A Laboratory Manual (Cold Spring Harbor, New York: Cold Spring Harbor Laboratory).
SchOpbach, T., and Wieschaus, E. F. (1986). Maternal-effect mutations altering the anterior-posterior pattern of the Drosophila embryo. Roux's Arch. Dev. Biol. 195, 302-317.
St Johnston, D., Driever, W., Berleth, T., Richstein, S., and Nüsslein-Volhard, C. (1989). Multiple steps in the localization of bicoid RNA to the anterior pole of the Drosophila oocyte. Development (Suppl.) 107, 13-19.
Struhl, G . (1989) . Differing strategies for organizing anterior and posterior body pattern in Drosophila embryos. Nature 338, 741-744.
Struhl, G., Struhl, K., and Macdonald, P. M. (1989). The gradient morphogen bicoid is a concentration-dependent transcriptional activator. Cell 57, 1259-1273.
Tautz, D. (1988). Regulation of the Drosophila segmentation gene hunchback by two maternal morphogenetic centers. Nature 332,281 284.
Tautz, D., and Pfeifle, C. (1989). A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback . Chromosoma 98, 81-85.
Vässin, H., and Campos-Ortega, J . A. (1 987). Genetic analysis of Delta , a neurogenic gene of Drosophila melanogaster . Genetics 116, 433445.
Vässin, H ., Bremer, K. A., Knust, E., and Campos-Ortega, J. A. (1 987). The neurogenic gene Delta of Drosophila melanogaster is expressed in neurogenic territories and encodes a putative transmembrane protein with EGF-like repeats. EMBO J. 6, 3431-3440.
Wharton, K. A., Johansen, K. M., Xu, T., and Artavanis-Tsakonas, S. (1985). Nucleotide sequence from the neurogenic locus Notch implies a gene product that shares homology with proteins containing EGF-like repeats. Cell 43, 567-581.
Wharton, R. P., and Struhl, G. (1989). Structure of the Drosophila BicaudalD protein and its role in localizing the posterior determinant nanos . Cell 59, 881-892.
Wieschaus, E. F., and Nüsslein-Volhard, C. (1986). Looking at embryos. In Drosophila : A Practical Approach, D. B. Roberts, ed. (Washington, DC: IRL Press), pp. 199-227.
GenBank Accession Number
The accession number for the sequence reported in this article is M72421.
Return To Molecular Biology Main Page
© Copyright
2000 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu