Cell, Vol. 67, 469-481, November 1, 1991, Copyright © 1991 by Cell Press
* Department of Molecular Biology and Biochemistry
+ and the Developmental Biology Center
University of California
Irvine, California 92717
# Department of Molecular and Cell Biology
University of California
Berkeley, California 94720
Summary
Mutations in the Drosophila tolloid (tld) gene lead to a partial transformation of dorsal ectoderm into ventral ectoderm. The null phenotype of tld is similar to, but less severe than decapentaplegic (dpp), a TGF-ß family member required for the formation of all dorsal structures. We have cloned the tld locus by P element tagging. At the blastoderm stage, tld RNA is expressed dorsally, similar to that described for dpp. Analysis of a tld cDNA reveals three sequence motifs: an N terminal region of similarity to a metalloprotease, two EGF-like repeats, and five copies of a repeat found in human complement proteins C1r and C1s. tld sequence is 41% identical to human bone morphogenetic protein 1 (BMP-1); the closest members to dpp within the TGF-ß superfamily are BMP-2 and BMP-4, two other bone morphogenetic proteins. These findings suggest that these genes are members of a signal generating pathway that has been conserved between insects and mammals.
Introduction
Within the dorsal half of the Drosophila embryo, at least six zygotically acting genes, decapentaplegic (dpp), tolloid (tld), screw, shrew, zerknüllt, and twisted gastrulation directly influence pattern formation (Nüsslein-Volhard et al., 1984; Wieschaus et al.,1984, Jürgens et al.,1984; Wakimoto et al.,1984; Zusman and Wieschaus,1985; Irish and Gelbart, 1987). Among these genes dpp appears to play a pivotal role, since it is the only gene required for the production of all dorsal structures. Null mutations of dpp cause a complete transformation of dorsal epidermis into ventral ectoderm (Irish and Gelbart 1987). In contrast, null mutations of tld delete the amnioserosa and a portion of the dorsal ectoderm and cause a corresponding expansion of the neurogenic ectoderm. The similarity of phenotypes suggests that both gene products could be acting in close proximity in the dorsal-ventral (D-V) patterning pathway. Additional support for the close relationship between tld and dpp has been suggested by the fact that mutations in either gene lead to similar premature loss in zerknüllt steady-state mRNA levels in early embryos (Rushlow and Levine, 1990).
DNA sequence analysis of the dpp gene has shown that its protein product is a member of the TGF-ß superfamily of growth factors (Padgett et al.,1987). This family of secreted polypeptides represents a large collection of factors with diverse activities. Depending on the cell type involved, different family members can stimulate or inhibit cell growth or differentiation, as well as induce alterations in the composition of the extracellular matrix (for reviews, see Barnard et al., 1990; Massagué, 1990). In addition, many TGF-ß family members also appear to be involved in control of cell fate. Müllerian inhibiting substance was originally shown to induce regression of female genitalia primordium in mammalian male embryos (Blanchard and Josso, 1974) and may also directly affect testicular morphogenesis (Behringer et al.,1990), while injection of activin-related polypeptides induces a secondary developmental axis in Xenopus and chicken embryos (Thomson et al., 1990; Mitrani et al., 1990).
Within the TGF-ß superfamily, dpp is most closely related to human bone morphogenetic protein 2 (BMP-2), exhibiting 75% identity with its C-terminal growth factor domain (Wozney et al., 1988; Gelbart,1989). The BMPs were isolated as a copurifying mixture of proteins that was capable of inducing ectopic bone formation in rats (Wozney et al., 1988; Wang et al., 1988; Ozkaynak, 1990; Celeste et al., 1991). Of the seven BMPs that have been characterized, six show sequence similarity to the TGF-ß superfamily. The only BMP that is not a TGF-ß family member, BMP-1, has multiple structural motifs, including a region of similarity to a metalloprotease from the crayfish Astacus fluviatilis and a region of similarity to a domain present in the human complement proteins C1r and C1s (Wozney et al.,1988 and this report). Although the copurification of BMP-1 with the other BMPs suggested that the BMP-1 protein could physically interact with one or more of the TGF-ß BMPs, the role of BMP-1 in bone morphogenesis remains unclear.
In this report, we provide additional evidence for a biochemical link between D-V patterning in Drosophila and bone induction in mammals. Our analysis of the tld sequence indicates that it encodes a protein of 120 kd MW that is highly homologous to the BMP-1 protein. These results suggest a surprising conservation of molecular mechanisms between two diverse processes and organisms.
Results
tld Mutations Cause a Partial Ventralization of the Embryonic
Pattern
The tld gene was first identified in the third chromosomal screen
for zygotic pattern mutants (Jürgens et al.,1984). Compared with wild-type
embryos (Figure 1 A), tld mutant embryos lack a characteristic subset
of dorsally derived cuticular structures, indicating that the gene is required
for normal dorsal development (Figure 1 B). Embryos that lack tld
are missing some, but not all, structures derived from the dorsal 40% of
the blastoderm fate map. For example, null mutants of tldlack all
of the dorsally derived cuticular specializations of the head such as the
internal structures of the pharangeal skeleton, the antenno-maxillary sense
organs, and the cirri. A majority of tld null embryos also lack the
prominent dorsolaterally derived structure of the tail, the filzkörper.
Not all dorsal epidermal structures are deleted in tld embryos: cells
in the dorsal regions of the thorasic and abdominal segments in tld
null embryos still secrete the fine dorsal hairs characteristic of the wild-type
dorsal epidermis. Concomitant with the loss of some dorsal structures in
tld embryos, there is an expansion in the size of the neurogenic
ectoderm, as measured by an increase in the left-right extent of each denticle
band. Embryos lacking dpp activity have a more severe phenotype than
tld embryos. In dppnull embryos, all dorsally and dorsolaterally
derived structures are absent, and denticle bands circumscribe the entire
cuticle (Irish and Gelbart, 1987) (Figure 1 C). Since both tld and
dpp mutants display abnormalities in the pattern of gastrulation
similar to maternal effect mutants that ventralize the embryonic fate map
(Jürgens et al.,1984; Irish and Gelbart,1987), mutations in both genes
most likely cause shifts in the fate map of the blastoderm embryo (Figure
1D).

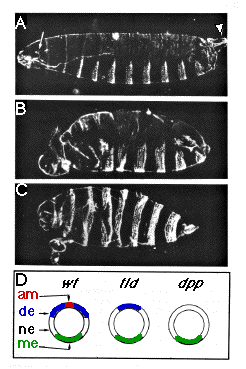
Figure 1. tld Mutations Result in a Partial Ventralization of
the Blastoderm Fate Map
(A) A lateral view of a wild-type embryo. In this figure, as in all others,
anterior is to the left and dorsal is up. In the head, the arrow notes the
internal pharangeal skeleton, which is derived mainly from cells located
at dorsal positions of the blastoderm fate map (Jürgens et al., 1986).
In the tail, the arrowhead notes the position of the paired respiratory
structures, the filzkörper, which are derived from a dorsolateral position
of the blastoderm fate map (Jürgens, 1987).
(B) An embryo lacking tld activity of genotype tld66 621tld6'.
In this embryo, all structures of the pharangeal skeleton and filzkörper
are deleted. In addition, the ventrally derived denticle belts are expanded
laterally.
(C) An embryo lacking dpp activity of genotype dpp48/Df(2L)
DTD2. In this embryo all dorsal structures have been deleted, and the denticle
belts circumscribe the cuticle.
(D) Schematic drawings of the blastoderm fate maps of wild-type, tld,
and dpp embryos. am, amnioserosa; de, dorsal ectoderm; ne, neurogenic
ectoderm; me, mesoderm.
Identification and Cloning of a P Element Insert within the tld
Gene
In the course of screening several hundred P element transformant lines
containing Ubx-LacZ fusions (Simon et al.,1990), we obtained one line 68-62,
which was homozygous lethal, with homozygotes exhibiting partially ventralized
embryonic cuticles. The insertion mapped on chromosome 3 and by in situ
hybridization was localized to polytene chromosome subdivision 96A, the
known cytogenetic position of tld (Jürgens et al., 1984). In
crosses to other tld mutations, 68-62 behaved as a strong tld
allele (Figure 1 B). We concluded that this stock likely contained a P insertion
mutation within the tld gene, which we named tld68-62.
From a genomic library derived from tld 68-62 DNA, phage
clones of the genomic region surrounding the P insertion were recovered
and used to obtain comparable clones from a wild-type genomic library. Mapping
and hybridization experiments showed that, in addition to the P insertion,
the tld68-62 chromosome had also suffered a 3.5 kb deletion
of genomic DNA to the left of the insertion site (Figure 2).
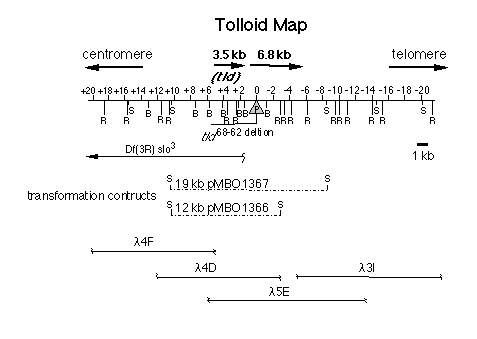
Figure 2. Structure of the tld Region
A restriction map of the 35 kb of DNA surrounding the site of the tld68-62
P element insertion is illustrated with the positions of the restriction
sites for the following enzymes noted: R. EcoRI; S. SalI; and B. BamHI.
The point of the P insertion is indicated by the stippled triangle. The
lines below the restriction map indicate the extents of the tld68-62
and slo3 (Atkinson et al., 1991) deletions. The heavy black arrows
above the line represent the map positions of two cDNAs located in the vicinity
of the P insertion. The locations of introns within these transcription
units have not yet been determined. The SalI fragments that were used for
P element rescue experiments are indicated by dotted lines below the map.
The DNA contained within four phage that were used to generate the restriction
map are shown at the bottom of the figure.
Identification of Transcripts within the Vicinity of the P Insert
To identify potential tld transcripts, a 2.8 kb BamHI genomic fragment
(Figure 2, coordinates +1.5 through -1), which included the P element insertion
site, was used as a hybridization probe on developmental Northern blots.
Two transcripts of approximately 6.8 and 3.5 kb were observed (Figure 3).
Full-length cDNA representatives of each transcript were obtained, and their
positions along the genomic map were determined by restriction enzyme analysis
and hybridization experiments (Figure 2). Based on these data, it was not
possible to determine unambiguously which transcript represented tld.
since both were disrupted by the P element insertion and its associated
deletion. To help resolve this issue, Southern blots were prepared with
DNA from 22 independent mutant tld alleles and screened by hybridization
with various genomic fragments covering the region from +12 through -14
kb. One deletion that failed to complement tld. slo3
(Atkinson et al., 1991), broke within the 0.8 kb BamHI fragment and removed
all DNA to the left of +2.1. This finding suggested that the 3.5 kb transcript
could be the tld product.
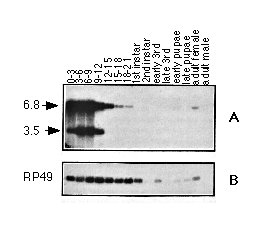
Figure 3. Developmental Profile of Two Transcripts from the tld
Region
Poly(A)+ RNA was isolated from whole organisms of the indicated stages.
Approximately 5 µg of RNA was loaded per lane and separated on a 1%
formaldehyde gel. The fractionated RNA species were then blotted to a nylon
membrane and hybridized with (A) a mixture of cDNA 6a (6.8 kb) and 4c (3.5
kb) or (B) with a probe for the RP49 ribosomal protein gene. Even on prolonged
exposure, there is no detectable tld expression in larvae pupal,
or adult stages (data not shown).
Transformation Rescue of tld Mutants
To determine whether the 3.5 kb transcript was the tld mRNA, we cloned
two different restriction fragments (Figure 2) into the P element transformation
vector Carnegie 20 (Karess and Rubin, 1984). Plasmid pMBO1367 contained
an 18 kb SalI fragment that included both the 6.8 and 3.5 kb transcripts,
as well as an additional 10.0 kb of flanking DNA. A second construct, pMBO1366,
contained a 14 kb SalI fragment that included only the 5' half of the 6.8
kb transcript, but otherwise left the 3.5 kb transcript and 5' flanking
sequences intact. Eleven independent transformant lines for each construct
were obtained, and 3 of each were tested for their ability to rescue tld
mutants. Both fragments were able to rescue tld mutations, and thus
we concluded that the 3.5 kb transcript is sufficient for tld function.
We also examined the phenotype due to loss of the 6.8 kb transcript by crossing
lines that contained the shorter construct back to the original tld68-62
mutation. Lack of the 6.8 kb transcript gave rise to a maternal effect mutation
with a phenotype quite distinct from that of tld (see Experimental
Procedures).
Pattern of tld Expression in Early Embryos
The spatial distribution of tld mRNA was detected by in situ hybridization
of digoxigenin-labeled tld probes to whole-mount embryos (Tautz and
Pfeifle,1989). We were particularly interested in how tld expression
compared with dpp in light of sequence data (see below) that suggested
the possibility of a physical interaction between the two proteins. tld
RNA was first detected at nuclear cycle 10-11 (Figure 4A). In the central
region of the embryo, only dorsally located nuclei expressed tld
while at the poles both dorsally and ventrally located nuclei are labeled.
This pattern intensified and expression peaked after nuclear division cycle
13 during the early stages of cellularization. At this point tld
staining encompassed the dorsal-most ~50% of the embryo (Figures 4B and
4C), and its expression pattern paralleled that seen for dpp (compare
Figure 4C with 4D; see also St. Johnson and Gelbart,1987; Ray et al., 1991).
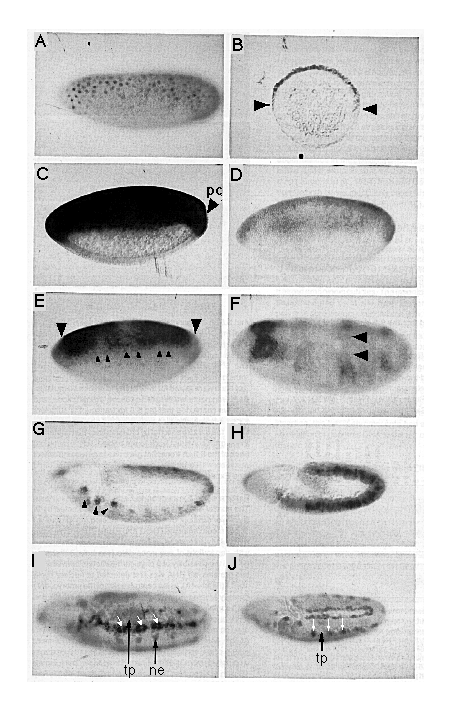
Figure 4. A Comparison of the Spatial Distributions of tld and
dpp RNA during Early Embryogenesis
Embryos were hybridized to digoxigenin-labeled probes as described. All
embryos are positioned with anterior on the left and dorsal on top unless
otherwise indicated. (A) Lateral view of nuclear cycle 10/11 embryos hybridized
with a tld probe. (B) Cross section at mid egg length of syncytial
blastoderm embryo hybridized with a tld probe. The arrowheads indicate
the D-V extent of the staining. (C) A lateral view of nuclear cycle 13 embryo
hybridized with a tld probe. Note absence of pole cell (pc) staining.
(D) A nuclear cycle 13 embryo hybridized with a dpp probe. (E) A
lateral view of a tld hybridized embryo undergoing cellularization.
Note that cells in the vicinity of each pole as well as cells between the
three sets of arrowheads show reduced levels of staining. (F) A dorsolateral
view of a tld hybridized embryo at the completion of cellularization.
Note that cells in the dorsal-most regions (between arrowheads) no longer
show any labeling. (G) A stage 8 embryo hybridized with a tld probe.
The arrowheads highlight the strongest staining in the gnathal segments.
The apparent dark staining on the top is an artifact of the photograph.
(H) A stage 8 embryo hybridized with a dpp probe. (I) A ventral-lateral
view of a late stage 11 embryo hybridized with a tld probe. The dark
arrows highlight a patch of neurogenic ectoderm cells (ne) that labeled
and, for reference, a tracheal pit (tp). The white arrows denote three sets
of the most strongly labeled cells, which appear to be located near the
boundary of the neurogenic ectoderm and the dorsal ectoderm. (J) A stage
11 embryo hybridized with a dpp probe. The dark arrow points out
a tracheal pit, and the white arrows point out cells that appear to be equivalent
to those marked with white arrows in (I).
During cellularization, the pattern of tld expression was found to undergo rapid modulation. The first alteration was a reduction in the staining of cells located within 10% egg length from each pole. This was quickly followed by additional reductions in staining along the anterior-posterior axis, such that four distinct bands of expression were observed by the time cellularization was complete (Figures 4E and 4F). During this same time period, modulation along the dorsal-ventral axis also occurred, such that cells located in the dorsal-most 10%-20% of the embryo's circumference became less intensely stained (Figure 4F). During the rapid phase of germband elongation, the two strongest bands of staining persisted for some time, but faded by the time the germ band was 66% extended at the completion of stage 8 (data not shown).
A new tld expression pattern emerged during late stage 8 to early stage 9. Initially this pattern was composed of three lateral patches of cells located within the emerging gnathal segments, as well as additional patches of weakly labeled cells in more posterior segments (Figure 4G). The pattern continued to evolve with time and achieved its full complexity during stage 11. The most intense staining at this stage was seen in a series of epidermal cells shaped as repeating sets of parentheses. These cells are located ventral to each tracheal pit. Weaker staining was seen in cells that encircled each pit and in a series of segmentally reiterated patches of cells within the neurogenic ectoderm (Figure 4I). This pattern disappeared before the germband fully retracted during stage 12.
Unlike the significant overlap seen between the tld and dpp patterns at the syncytial blastoderm stage, dpp expression during stages 8 through 11 is substantially different than the tld pattern. In particular, during stages 8 through early 10, dpp is expressed in a much broader region, which encompasses the entire dorsal epidermis (Figure 4H; St. Johnston and Gelbart, 1987; Ray et al., 1991). Subsequently, during late stage 10 and early 11, the pattern of dpp expression resolves into two parallel groups of cells. One group comprises a continuous stripe of cells located between the dorsal epidermis and the amnioserosa, while the second group is composed of a segmentally repeated pattern of cells positioned near the boundary between the dorsal epidermis and the neurogenic ectoderm (Figure 4J; St. Johnston and Gelbart 1987; Ray et al.,1991). The location of this second group of cells relative to the developing tracheal pits suggests that they are nearly coincident with those tld -expressing cells that form the parentheses (compare Figure 4I with 4J). However, other aspects of the two patterns are quite distinct. For example, tld was not expressed in the thin strip of cells located between the dorsal epidermis and amnioserosa. dpp transcription then continues throughout the remainder of embryogenesis in the absence of any additional tld expression (St. Johnston and Gelbart, 1987; Ray et al., 1991).
DNA Sequence and Conceptual Translation of the tld Gene
The complete nucleotide sequence of one tld cDNA isolate, 4c, was
determined and is shown in Figure 5. Allowing for polyadenylation, this
3385 bp sequence is in close agreement with the transcript size of 3.5 kb
observed on Northern blots, suggesting that cDNA 4c is full-length or nearly
so. The cDNA sequence contains one large open reading frame (ORF) of 3171
bp, which is flanked by 102 bp of 5'-untranslated sequences and 112 bp of
3'-untranslated sequences. At the 5' end, five closely spaced methionine
codons are found in-frame with the large ORF. Of these, the fifth one shows
the best fit to the ANN(C/A)A(A/C)A/ C)ATGN consensus sequence for translation
start sites in Drosophila (Cavener, 1987). Assuming that this AUG
is used as the start codon, translation of the large ORF would produce a
1057 amino acid protein with an approximate Mr of 116,000 kd.
Hydropathy analysis by the method of Kyte and Doolittle (1982) revealed
one hydrophobic region at the N-terminus of the protein. This region conforms
to the rules that define signal sequences, and a probable cleavage site
for signal peptidase can be positioned after Gly 26 (von Heijne, 1985).
We also note the locations of eight potential sites for N-glycosylation
as well as two RGD sequences. In proteins of the extracellular matrix, RGD
sequences have been found to mediate interactions with integrins (Ruoslahti
and Piershcbacer, 1987). The cDNA is probably complete at the 3' end, since
a potential polyadenylation signal, AATAAA, falls 23 nucleotides upstream
of the terminal poly(A) track and 70 nucleotides downstream of the termination
of translation.
The Tld Protein Appears to Be Composed of Several Structural Domains
with a High Degree of Overall Similarity to Human BMP-1
The amino acid sequence of the deduced Tld protein was compared with the
sequences in the NBRF protein data base (release 19) and the translated
GenBank data base (release 62). As shown in Figure 6, a striking similarity
was found between Tld and human BMP-1. BMP-1 was originally isolated as
a component of a highly purified mixture of proteins which, upon subcutaneous
injection into rats, was able to induce ectopic bone formation (Wozney et
al., 1988). Tld and BMP-1 show 41% amino acid sequence identity over most
of their lengths. Two major differences between the two proteins are that
Tld contains an additional 298 amino acids on its C-terminal end relative
to BMP-1 and that the N-terminal 120 amino acids show little similarity.
The two proteins each contain three distinct sequence elements (Figures
5 and 6). Within the N-terminal region, Wozney et al. (1988) noted that
BMP-1 showed a high degree of similarity to a zinc metalloprotease from
the crayfish Astacus fluviatilis (Tianti, 1987). Tld also
shows this homology (boxed in Figure 5), and a comparison of the Astacus
sequence with the Tld and BMP-1 sequences is shown in Figure 6B. We find
30% identity among all three sequences over the entire length of the Astacus
protease. The Tld and BMP-1 sequences show an additional 15% identity in
this region. The Astacus protease has been shown to bind one zinc
ion per protease molecule, and chelating experiments have shown that this
metal ion is required for catalytic activity (Stocker et al.,1988). In thermolysin
the sequence HELXH has been shown by X-ray crystallographic studies to form
part of a metal ion binding site (Matthews et al.,1974). This same sequence
is found in Astacus protease, BMP-1, and Tld (Figure 6B, hatched
area) and may play a role in coordinating zinc in these proteins.

Figure 5. Nucleotide and Predicted Amino Acid Sequence of the tld
cDNA
The nucleotide sequence of the 3375 bp tld cDNA is shown together
with the conceptual translation product. The five possible AUG start codons
are boxed. The fifth one (shaded) shows the best fit to the consensus Drosophila
translation start site. Potential glycosylation sites are underlined with
black bars. Two RGD sequences are illustrated with hatched bars. The probable
polyadenylation signal is shown with a shaded box. The open boxed amino
acid sequence denotes a potential signal peptide as predicted by the method
of von Heijne (1985). The sequences in the blue box are homologous to a
crayfish metalloprotease (Titani et al., 1987). The sequences within the
green boxes show similarity to EGF-like repeats (Appella et al., 1988).
The Tld sequence also contains five copies of a repeat motif that has been
found in human complement proteins C1r and C1s (Journet and Tosi, 1986;
Mackinnon et al., 1987). Each repeat contains four conserved cysteine residues
that are boxed or circled in yellow alternate repeats.


Figure 6. Structural Organization of Tld, BMP-1, and C1 r/s and Comparison
of Similar Sequence Motifs
(A) Schematic representations of the domain structures of the Tld, BMP-1,
and C1r/s proteins are shown. The open boxes indicate the protease domains.
The hatched boxes represent C1 r/s type repeats, and the black boxes represent
EGF-like sequences. In C1 r and C1 s, two repeats that separate the CEC
domain from the protease domain are indicated by the thin open boxes. (B)
A comparison of the putative Tld, BMP-1, and Astacus protease domains. Vertical
dashes between sequences indicate identical amino acids. A potential metal
ion binding sequence, HELXH, is denoted with a shaded box. In the crayfish
protease, the two cysteines noted by asterisks have been shown to be linked
by disulfide bonds, as have the two cysteines connected by the bracket (Titani
et al., 1987). (C) A comparison of the Tld and BMP-1 EGF-like sequences.
The consensus sequence for a type 3 EGF repeat is from Appela et al. (1988).
(D) Comparison of the C1 r/s-like repeats. The open boxes indicate conserved
cysteines. Gaps to maximize homologies are indicated by blank spaces, while
loop outs are shown by amino acids being placed either above or below the
sequence line. In addition, the Tld and BMP-1 repeat units are compared
in pairs according to their linear order in the sequence with identity between
each pair indicated by vertical dashes. The last four sequences are from
C1r and C1s, and they are paired according to their linear order of appearance
within the C1 r and C1 s proteins. A consensus sequence is shown at the
bottom of the figure. Indicated residues were present in at least 8 of the
12 sequences at that position. The presence of two residues indicates that
at least 8 out of 12 sequences showed one or the other of the two amino
acids. Note that the amino acids between cysteine residues 1 and 2 show
the highest degree of conservation, with two notable exceptions: the first
Tld repeat appears to have a fairly large insertion relative to the others,
and the first C1 r and C1s repeats do not contain cysteines 1 and 2. The
last portion of the repeat, distal to cysteine 4, shows the greatest heterogeneity
in both length and sequence.
The remaining C-terminal portions of Tld and BMP-1 are composed of two types of repeating sequence motifs. The first motif is a 36 amino acid sequence with a high degree of similarity to epidermal growth factor (EGF)-like sequences. Two EGF-like sequences are found in Tld beginning at residues 585 and 747 (Figure 5), whereas BMP-1 has only one (see Figure 6C for a comparison). These EGF-like sequences contain the subsequence, Asp X Asp/ Asn Cys Xn Cys Xn Cys X Asp*/Asn*-X4 Tyr/Phe X Cys X Cys X4 Glu, which is thought to signal posttranslational hydroxylation of the third Asp or Asn residue (*) of the consensus (Stenflo et al.,1987; Rees et al.,1988). In the case of Tld and BMP-1, this would lead to a ß-hydroxy asparagine residue. This consensus sequence and/or the ß-hydroxylate residue may play a role in forming a high affinity calcium binding site (Öhlin et al., 1988; Rees et al., 1988; Handford et al., 1990).
The second type of repeat found in Tld and BMP-1 is based on the conserved spacing of four cysteine residues and is, on average, 113 amino acids long. There are five such repeats in Tld: two tandem repeats precede the first EGF repeat, one falls between the EGF repeats, and then two more tandem copies follow the second EGF repeat. BMP-1 contains the first three repeats, which are similarly positioned about the EGF-like sequences, but because of the C-terminal truncation, BMP-1 does not contain the final two repeats.
A computer search of available data bases revealed that this type of repeat is also found in the two complement proteins, C1r and C1s (Journet and Tosi, 1986; Mackinnon et al., 1987). These proteins are serine proteases which, together with C1q, form the first component of the classical complement cascade. Like Tld and BMP-1, the EGF-like sequences of C1r and C1s are sandwiched between two of the 113 amino acid repeats (Figure 6A), suggesting that a functional unit may actually comprise one or more of these repeats flanking an EGF-like sequence. A comparison of 11 C1r/s-type repeats is given in Figure 6D along with a derived consensus sequence.
Discussion
Copurification of BMPs Suggests a Possible Physical Interaction between
tld and dpp
We have shown by DNA sequencing that tld gene product is very similar
in primary sequence and overall domain structure to human BMP-1. The fact
that the original mixture of BMPs copurified as a complex suggests that
a physical interaction takes place between BMP-1 and one or more of the
TGF-ß-like BMPs. If Tld provides the same biochemical function as
BMP-1, then by analogy to the BMPs, one might expect that Tld should also
physically interact with a TGF-ß molecule and that perhaps this molecule
is the dpp product. Evidence supporting this hypothesis comes from
the observation that embryos heterozygous for certain tld alleles
and a recessive, partial loss-of-function dpp allele die with a partially
ventralized phenotype (E. L. F. and K. Anderson, unpublished data; L. Raftery
and tld. Gelbart, personal communication). As this intergenic failure
of complementation is not observed in similar crosses using a deficiency
of the tld locus, it is possible that these unusual tld alleles
fail to complement the dpp mutation because they produce aberrant
products that form nonfunctional complexes with the dpp protein.
The tld Transcript Shows Two Distinct Rounds of
Expression in Early Embryos: The Precellularization Pattern of tld
Transcription Overlaps the dpp Pattern
The tld expression pattern prior to cellularization is consistent
with a possible interaction with dpp. For both genes, transcripts
begin to accumulate during nuclear division cycle 10 or 11, and with the
exception of nuclei located near the poles, expression is limited to the
dorsal surface (St. Johnston and Gelbart,1987 and this work). tld
expression reaches a maximum level just after nuclear division cycle 13,
and its expression domain encompasses the dorsal-most 50% of the embryo
surface. This is slightly broader than the 40% limit that has been reported
for dpp.
It is noteworthy that the early transcription pattern of zerknüllt,
a third zygotic gene required for patterning of dorsal tissue, is also confined
to the dorsal-most 40%50% of the embryo's circumference (Rushlow et al.,1987).
For all three genes, the lateral borders of their expression domains are
thought to be set by dorsal (dl ), since in dl mutant
backgrounds each gene is uniformly expressed around the entire circumference
of the embryo (Rushlow et al.,1990; Ray et al.,1991; K. Arora and M. B.
O., unpublished data). The dl morphogen has been found to be a DNA-binding
protein related to NF-![]() B and the rel oncogene (Ip
et al., 1991; Thisse et al., 1991; Stewart, 1987). There is a gradient of
nuclear localization of dl such that the highest concentration of
dorsal protein is observed in the ventral-most nuclei, and little or no
protein is present in dorsal nuclei (Rushlow et al., 1989; Roth et al.,
1989; and Steward, 1989). The similarities in the extent of expression for
all three genes along the D-V axis may indicate that the dl nuclear
localization gradient establishes one final response threshold at 40%-50%
egg circumference. Thus, the dl gradient may not provide positional
information to pattern the dorsal regions of the embryo, suggesting that
the spatial organization of this region may be specified by interaction
among zygotic genes.
B and the rel oncogene (Ip
et al., 1991; Thisse et al., 1991; Stewart, 1987). There is a gradient of
nuclear localization of dl such that the highest concentration of
dorsal protein is observed in the ventral-most nuclei, and little or no
protein is present in dorsal nuclei (Rushlow et al., 1989; Roth et al.,
1989; and Steward, 1989). The similarities in the extent of expression for
all three genes along the D-V axis may indicate that the dl nuclear
localization gradient establishes one final response threshold at 40%-50%
egg circumference. Thus, the dl gradient may not provide positional
information to pattern the dorsal regions of the embryo, suggesting that
the spatial organization of this region may be specified by interaction
among zygotic genes.
The Postcellularization Pattern of tld Transcription Diverges
from dpp
During cellularization and subsequent stages of embryogenesis, the patterns
of tld and dpp expression show some overlap, but they are
not as closely aligned as at early stages. In particular, during stage 11,
tld expression extends both ventrally into the neurogenic ectoderm
and dorsally around the developing tracheal pits, where dpp expression
is not seen. The fact that the tld and dpp patterns only partially
overlap may indicate that tld has an additional, dpp-independent
function during this time period. Alternatively, since these proteins are
likely to be secreted, functional protein interactions may take place at
sites removed from their synthesis. What is difficult to assess, is how
far these proteins might diffuse from their source cells. Immunolocalization
studies on developing embryos have suggested that both dpp and the
Drosophila int-1 homolog wingless (another secreted
growth factor) accumulate to high levels within distances of only one or
two cell diameters from their source cells (Immergluck et al.,1990; Reuter
et al.,1990; Panganiban et al.,1990; van den Heuvel et al., 1989). Since
the distances between some groups of tld - and dpp-expressing
cells differ by more than two nuclei, there could be significant accumulation
of one protein in the absence of the other.
The dpp gene product is also required for the proper development and morphogenesis of the imaginal disks that produce the cuticular structures of the adult fly (Spencer et al.,1982). If, as we postulate, the Tld protein provides an integral component of the signal generating pathway for TGF-ß molecules, it is surprising that Tld expression is restricted to the first 8 hr of embryogenesis. This paradox has been partially resolved by the identification of a Tld homolog that appears to be expressed throughout development (J. Jamal, M. J. S., and M. B. O., unpublished data; and R. Padgett, personal communication). However, the involvement of this protein in later developmental events remains to be established.
Structural Features of the Tld and BMP-1 Proteins: Implications for
Function
Analysis of the deduced amino acid sequence suggests that Tld contains at
least three distinct sequence motifs, each of which is also found in human
BMP-1 and two of which are found in complement proteins C1r and C1s. Figure
6A shows a schematic comparison of the domain structures of tld,
BMP-1, C1r, and C1s as inferred from sequence homology data. Both Tld and
BMP-1 have potential metalloprotease domains located in the N-terminal regions,
whereas C1r and C1s have serine protease domains located in the C-terminal
portion of the protein. Of particular interest is the fact that all four
proteins share what we refer to as a CEC supermotif. This motif is composed
of two C1r/s repeats flanking an EGF-like sequence. There is substantial
evidence that the CEC supermotifs of C1r and C1s are important for calcium-dependent
tetrameric complex formation between two C1r-C1s dimers and the association
of this complex with C1q to form the mature C1 molecule (reviewed by Arlaud
et al.,1987). By analogy we might expect that the CEC regions of Tld and
BMP-1 could also form interaction domains which bind calcium and promote
the formation of multimeric complexes that may include a TGF-ß-like
molecule.
The only extensive region of nonhomology between Tld and BMP-1 is the N-terminal portion up to the start of the protease domain. In Tld, this region measures 126 amino acids, while in BMP-1 it is 120 residues. This sequence may serve some pathway- or species-specific function or, by analogy to several of the blood clotting proteases and matrix metalloproteases, these regions could serve to maintain the proteins as deactivated zymogens (Furie and Furie, 1988; Matrisian, 1990). We note the presence of a tetrabasic (Tld) or dibasic (BMP-1) sequence immediately prior to the start of the protease domain, which could be proteolytically processed to activate the zymogen.
What are the biochemical roles of Tld and BMP-1 ? The most obvious possibility is that they are the processing proteases that cleave the precursor forms of DPP and BMP-2 (and/ or BMP-3), respectively, to liberate the mature C-terminal TFG-ß-like peptides. Both EGF and nerve growth factor (NGF) are known to be complexed with processing proteases when secreted from certain tissues (Taylor et al., 1974; Server and Shooter, 1976). Such a role would explain the copurification of BMP-1 with BMP-2 and BMP-3 and would be consistent with the genetic interactions noted between tld and dpp.
Another possible function for Tld and BMP-1 would be to activate latent complexes of DPP and BMP-2/3, respectively. TGF-ß1, for example, is known to be secreted in a latent form that is comprised of two mature C-terminal peptides noncovalently associated with two N-terminal chains and a third protein known as TGF-ß-binding protein (TGF-ß1-BP) (Miyazono et al., 1988; Wakefield et al., 1988; Kanzaki et al.,1990). The in vivo mechanism for activating the latent complex is not known, but Lyons et al. (1988) and Sato and Rifkin (1989) have suggested that proteolytic processing might be involved. At present neither DPP nor any of the BMPs has been shown to form latent complexes. Other possibilities can also be envisioned, and the availability of a tld cDNA will facilitate the assignment of biochemical activities to the tld gene product and will allow us to test various models.
Drosophila D-V Patterning and Mammalian Bone Morphogenesis
Employ Similar Molecules
Sequence comparisons of BMP-2 and BMP-4 with known TGF-ß family members
first raised the possibility that these molecules are the vertebrate analogs
of dpp. The results described in this report not only strengthen
this view but suggest that additional aspects of an ancient signaling process
have been conserved since the split of arthropods and chordates. Such conservation
of underlying molecular mechanisms, in seemingly unrelated developmental
processes, presumably signifies the greater ease with which evolution can
usurp an existing pathway for a new purpose rather than evolve a new molecular
mechanism. Are other aspects of this signaling pathway conserved? At least
two other zygotically acting D-V genes, shrew and screw, appear
to function within the same pathway as tld and dpp (Jürgens
et al., 1984; K. Arora and C. Nüsslein-Volhard, personal communication;
E. L. F. and K. Anderson, unpublished data). Presumably many downstream
molecules, including the receptor for the TGF-ß-like peptides and
the intracellular signal transduction systems, could all be conserved with
only the ultimate target genes having evolved to give pathway and species-specific
diversities. This suggests the possibility that by studying D-V patterning
in the fly, one may in fact be able to gain additional insights into the
signaling pathway that operates to produce a distinctly vertebrate trait
such as bone morphogenesis (see also Kaplan et al., 1990). Like dpp,
however, the BMPs may also play a multitude of roles during early mammalian
development (Lyons et al.,1989; Jones et al.,1991), making them quite similar
to their Drosophila homologs in function.
Experimental Procedures
Drosophila Strains and Crosses
The tld68-62 mutation arose in a transformant line after
microinjection of P element plasmid pMBO141 (abx6.8b) (Simon et al., 1990).
The line was maintained over the TM2 balances A homozygous ry502 stock was
used as the recipient for all transformation experiments described in this
paper. The balancer stock T(2:3) ApXa/CyO; TM2, ry was used to assess the
chromosomal linkage of inserts. For transformation rescue experiments, transformant
lines 1366-65, carrying an insert of the 12 kb SalI fragment (Figure 4)
on chromosome 2 and 1367-43, carrying an insert of the 18 kb SalI fragment
(Figure 4) on chromosome 2 were crossed to the balancer stock listed above,
and male progeny of the genotype P [tld+]/CyO; +/TM2 were selected.
Single males were then crossed to females with the genotype +/CyO, S; tld5H/
TM3 or +/CyO,S; tld68-62/ TM3 and male progeny of the
genotype P [tld +/CyO,S; tld5H or tld68-62/TM2
were selected. These flies were then crossed to females of the genotype
tld'110E/TM3, and progeny were scored for the survival of tld
/tld offspring. In each case more than 90% of the expected tld
/tld progeny were found. All non-TM2, non-TM3 flies were found to
have inherited the P [tld +] chromosome (i.e., not CyO, S) indicating
linkage of the rescuing activity with the second chromosome insert.
To determine the phenotype that was associated with disruption of the 6.8 kb transcript, we crossed transformant lines containing the smaller 12 kb construct (pMBO1366) on the second chromosome to the original tld 68-62 allele. The resulting F1 flies were crossed inter se and scored for progeny bearing the transformant DNA on the second chromosome and homozygous for the tld68-62 mutant. Any defects associated with such flies should be attributable to lack of the 6.8 kb transcript, since the transformant chromosome will rescue the tldtranscription unit. Adult flies of the appropriate genotype were obtained, and except for a slight notching of the wings in about 50% of the progeny, these flies were of normal appearance. Adult females, however, were sterile and produced very few eggs. While not examined in detail, these embryos appeared to undergo abnormal cellularization and exhibited very little subsequent development. These defects cannot be attributable to other mutations on the tld 68-62 chromosome, since transformant lines containing the larger insert (pMBO1367) are able to completely rescue the tld 68-62 allele to full viability and fertility.
P Element Transformation
Germline transformation was performed essentially as described (Spradling
and Rubin, 1982). Host embryos (ry502) were injected with DNA solutions
containing 50 µg/ml each of the two helper plasmids p25.7 wc (Karess
and Rubin, 1984) and pUChsP![]() 2-3 (Mullins et al., 1989) and 600
µg/ml of either pMBO1366 or pMBO1367. Transformant flies were identified
by ry+ eye color.
2-3 (Mullins et al., 1989) and 600
µg/ml of either pMBO1366 or pMBO1367. Transformant flies were identified
by ry+ eye color.
Cloning of tld and Isolation of tld cDNAs
A genomic library was prepared from the tld 68-62/TM2
stock by ligating ~10-20 kb Sau3A partial fragments into the BamHI site
of ![]() EMBL3. P element-positive phage were detected after hybridizing
plaque lifts with the 0.5 kb BamHI-HindIII fragment of Carnegie 3 (Karess
and Rubin, 1984). Five positive clones were purified and mapped with several
restriction enzymes. A 1.9 kb BamHI genomic fragment (+2.2 to +4.1) which
flanked the P element insertion site was identified (Fig. 2) and used to
isolate wild-type genomic clones from a Canton S
EMBL3. P element-positive phage were detected after hybridizing
plaque lifts with the 0.5 kb BamHI-HindIII fragment of Carnegie 3 (Karess
and Rubin, 1984). Five positive clones were purified and mapped with several
restriction enzymes. A 1.9 kb BamHI genomic fragment (+2.2 to +4.1) which
flanked the P element insertion site was identified (Fig. 2) and used to
isolate wild-type genomic clones from a Canton S ![]() EMBL3
library kindly provided by R. Blackman. Standard protocol for library screening,
Southern hybridization, and
EMBL3
library kindly provided by R. Blackman. Standard protocol for library screening,
Southern hybridization, and ![]() phage purification were employed
(Maniatis et al., 1982).
phage purification were employed
(Maniatis et al., 1982).
The 2.7 kb (coordinates +1.5 to -1.3) and 1.9 kb BamHI (+2.2 to +4.1)
fragments from ![]() 5E (Figure 3), were used to screen a 0-8 hr embryonic
cDNA library kindly provided by Nick Brown (Brown and Katatos,1988). Approximately
80,000 colonies were screened and 22 positive clones identified. These fell
into two classes, which represented the adjacent 6.8 kb and the 3.5 kb transcripts.
Clone 6a is a full-length representative of the 6.8 class, while clone 4c
is a full-length representative of the 3.5 kb (tld) class.
5E (Figure 3), were used to screen a 0-8 hr embryonic
cDNA library kindly provided by Nick Brown (Brown and Katatos,1988). Approximately
80,000 colonies were screened and 22 positive clones identified. These fell
into two classes, which represented the adjacent 6.8 kb and the 3.5 kb transcripts.
Clone 6a is a full-length representative of the 6.8 class, while clone 4c
is a full-length representative of the 3.5 kb (tld) class.
Analysis of tld RNA Expression
Total mRNA was prepared from staged embryos, larvae, or adults by the hot
phenol method (Jowett, 1986). Poly(A)+ was purified by one cycle of binding
to oligo(dT)-cellulose (Collaborative Research). RNA was fractionated on
formaldehyde-agarose gels and transferred to nitrocellulose by capillary
blotting. Hybridization probes were prepared by random priming according
to standard methods.
For in situ hybridization, embryos were collected and fixed for 20 m in in a 2:1 mixture of PBS: heptane containg 4% formaldehyde. The embryos were devitellinized by washing in methanol and then stored until use at -20° C in 95% ethanol . DNA probes were labeled by digoxigenin-dATP (Genius kit, Boehringer Mannheim) according to the manufacturers protocol. Hybridization and detection were carried out by the procedure of Tautz and Pfeifle (1989) as modified by C. Oh and B. Edgar (personal communication). Embryonic stages are as described by Campos-Ortega and Hartensteing (1985). For cross sections, embryos were quick frozen on dry ice in OCT media (Miles Inc.) before sectioning.
DNA Sequencing and Computer Analysis
The tld cDNA 4c was subcloned as three separate EcoRI fragments into
plasmid pEMBL18 (Vieira and Messing, 1987). Both orientations of these subclones
were subjected to exonuclease digestion (Erase a Base Kit, Promega) in order
to generate a series of nested deletions. Deletions that occurred at approximately
every 200 bp were selected for sequencing. The dideoxy chain termination
method (Sanger et al., 1977) was used with Sequenase Version 2.0 (U. S.
Biochemical) enzyme. In most cases, double-stranded plasmid DNA, as prepared
by mini alkaline lysis (Maniatis et al.,1982), was sequenced. In several
GC-rich regions, however, single-stranded DNA templates were prepared by
infection with the filamentous phage M13K07 (Vieira and Messing,1987) .
Sequencing reactions were run on 6% polyacrylamide wedge gels. Both strands
were sequenced and, with a few exceptions, every base was read on average
three times. Sequencing data were entered and analyzed using the IBI Super
Sequencing System (MacVector Program). The deduced Tld protein sequence
was compared with the NBRF (version 19) data base using the program FASTA.
Alignments were done initially using the ALIGN program and subsequently
scanned by eye to maximize alignments
Acknowledgments
We wish to acknowledge Eileen Kimm for her assistance in analyzing the nested tld deletions and Martin Smith for the generous time and advice he gave us when using his IBI Sequencing System. We thank Yishi Jin for contribution of a developmental Northern blot, and Rob Ray and Francesca Pignoni for advice on in situ hybridization to wholemount embryos. We are grateful to Nigel Atkinson for supplying the slo3 stock. We thank Bill Gelbart for initially bringing BMP-1 to our attention, and we are grateful to Kathryn Anderson Larry Marsh, Mark Peifer, and Jeff Simon for critical evaluation of the manuscript. This work was supported by awards from the Academic Senate of UC Irvine, the Cancer Research Coordinating Committee of the University of California, and P. H. S. grant GM42546 to M. B. O. E. L. F. was supported by a Helen Hay Whitney fellowship, a NIH postdoctoral training grant, HD07299, and by P. H. S. grant GM35437 to Kathryn Anderson .
The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked "advertisement" in accordance with 18 USC Section 1734 solely to indicate this fact.
Received June 27, 1991; revised August 23, 1991.
References
Arlaud, G. J., Colomb, M. G., and Gagnon, J. (1987). A functional model of the human C1 complex. Immunol. Today 8, 106-110.
Appella, E., Weber, l. T., and Blasi, F. (1988). Structure and function of epidermal growth factor-like regions in proteins. FEBS Lett. 231, 1-4.
Atkinson, N. S., Robertson, G. A., and Ganetzky, B. (1991). A component of calcium-activated potassium channels is encoded by the Drosophila slo locus. Science 253, 551-555.
Barnard, J. A., Lyons, R. M., and Moses, H . L. (1990). The cell biology of transforming growth factor ,B. Biochim. Biophys. Acta 1032, 79-87.
Behringer, R. R., Cate, R. L., Froelick, G. J., Palmiter, R. D., and Brinster, R. L. (1990). Abnormal sexual development in transgenic mice chronically expressing Müllerian inhibitory substance. Nature 345, 167-170.
Blanchard, M., and Josso, N. (1 974). Source of anti-Müllerian hormone synthesized by the fetal testis; Müllerian inhibiting activity of fetal bovine Sertoli cells in culture. Pediatr. Res. 8, 968-971.
Brown, N., and Kafatos, F. C. (1988). Functional cDNA libraries from Drosophila embryos. J. Mol. Biol. 203, 425-437.
Cavener, D. R. (1987). Comparison of the consensus sequence flanking translation start sites in Drosophila and vertebrates. Nucl. Acids Res. 15, 1353-1361.
Campos-Ortega, J. A., and Hartenstein, B. (1985). The Embryonic Development of Drosophila melanogaster (Berlin: Springer-Verlag).
Celeste, A. J., Iannazzi, J. A., Taylor, R. C., Hewick, R. M., Rosen, V., Wang, E., and Wozney, J. M. (1990). Identification of transforming growth factor ,B family members present in bone-inductive protein purified from bovine bone. Proc. Natl. Acad. Sci. 87, 9843-9847.
Furie, B., and Furie, B. A. (1988). The molecular basis of blood coagulation. Cell 53, 505-518.
Gelbart, W. M. (1989). The decapentaplegic gene: a TGF-ß homolog controlling pattern formation in Drosophila. Development [Supp] 107, 65-74.
Handford, P. A., Baron, M., Mayhew, M., Willis, A., Beesley, T., Brownlee, G. G., and Campbell, l. D. (1990). The first EGF-like domain from human factor IX contains high-affinity calcium binding site. EMBO J. 9, 475-480.
Immergluck, K., Lawrence, P. A., and Bienz, M. (1990). Induction across germ layers in Drosophila mediated by a genetic cascade. Cell 62, 261-268.
Irish, V. F., and Gelbart, W. M. (1987). The decapentaplegic gene is required for dorsal-ventral patterning of the Drosophila embryo. Genes Dev. 1, 868-879.
Ip, Y. T., Kraut, R., Levine, M., and Rushlow, C. A. (1991). The dorsal morphogen is a sequence-specific DNA-binding protein that interacts with a long-range repression element in Drosophila. Cell 64, 439-446
Jones, M. C., Lyons, K. M., and Hogan, B. (1991). Involvement of bone morphogenetic protein-4 (BMP-4) and Vgr-1 in morphogenesis and neurogenesis in the mouse. Development 111, 531-542.
Journet, A., and Tosi, M. (1986). Cloning and sequencing of full-length cDNA encoding the precursor of human complement component C1 r. Biochem. J. 240, 783-787.
Jowett, T. (1986). Preparation of nucleic acids. In Drosophila: A Practical Approach, D. B. Roberts, ed. (Washington: IRL Press), pp. 279.
Jürgens, G. (1987). Segmental organization of the tail region in the embryo of Drosophila melanogaster. A blastoderm fate map of the cuticule structures of the larval tail region . Roux's Arch . Dev. Biol .196, 141-157.
Jürgens, G., Lehmann, R., Schardin, M., and Nüsslein-Volhard, C. (1986). Segmental organization of the head in the embryo of Drosophila melanogaster. A blastoderm fate map of the cuticle structures of the larval head. Roux's Arch. Dev. Biol. 195, 359-377.
Jürgens, G., Wieschaus, E., Nüsslein-Volhard, C., and Kluding, H. (1984). Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster. II. Zygotic loci on the third chromosome. Roux's Arch. Dev. Biol. 193, 283-295.
Karess, R. R., and Rubin, G. M. (1984). Analysis of P transposable element functions in Drosophila. Cell 38, 135-146.
Kaplan, F. S., Tabas, J. A., and Zasloff, M. A. (1990). Fibrodysplasia ossificans progressiva: a clue from the fly? Calcif. Tissue Int. 47,117125.
Kanzaki, T., Olofsson, A., Moren, A., Wernstedt, C., Heliman, U., Miyazono, K., Claesson-Welsh, L., and Heldin, C.-H. (1990). TGF-ß1 binding protein: a component of the large latent complex of TGF-ß1 with multiple repeat sequences. Cell 61, 1051-1061.
Kyte, J ., and Doolittle, R. F. (1982). A simple method for displaying the hydropathy character of a protein. J. Mol. Biol. 157, 105-132.
Lyons, R. M., Keski-Oja, J., and Moses, H. L. (1988). Proteolytic activation of latent transforming growth factory from fibroblast-conditioned medium. J. Cell Biol. 106, 1659-1665.
Lyons, K. M., Pelton, R. W., and Hogan, B. L. M. (1989). Patterns of expression of murine Vgr-1 and BMP-2a RNA suggest that transforming growth factor-ß-like genes coordinately regulate aspects of embryonic development. Genes Dev. 3, 1657-1668.
Mackinnon, C. M., Carter, P. E., Smyth, S. J., Dunbar, B., and Fothergill, J. E. (1987). Molecular cloning of cDNA for human complement component C1s. Eur. J. Biochem. 169, 547-553
Maniatis, T., Fritsch, E. F., and Sambrook, J. (1982). Molecular Cloning: A Laboratory Manual (Cold Spring Harbor, New York: Cold Spring Harbor Laboratory). \
Massagué, J. (1990). The transforming growth factory family. Annu. Rev. Cell Biol. 6, 597-641.
Matrisian, L. M. (1990). Metalloproteinases and their inhibitors in matrix remodeling. Trends Genet. 6, 121-125.
Matthews, B. tld., Wear, L. M ., and Kester, W. R. (1974). The conformation of Thermolysin. J. Biol. Chem. 249, 8030.
Mitrani, E., Ziv, T., Thomsen, G., Shimoni, Y., Melton, D. A., and Bril, A. (1990) Activin can induce the formation of axial structures and is expressed in the hypoblast of the chick. Cell 63, 495-501.
Miyazono, K., Heliman, U., Wernstedt, C., and Heldin, C.-H. (1988). Latent high molecular weight complex of transforming growth factor 1: purification from human platelets and structural characterization. J. Biol. Chem. 263, 6407-6415.
Mullins, M. C., Rio, D. C., and Rubin, G. M. (1989). Cis-acting DNA sequence requirements for Pavement transposition. Genes Dev. 3, 729-738.
Nüsslein-Volhard, C., Wieschaus, E., and Kluding, H. (1984). Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster 1. Zygotic loci on the second chromosome. Roux's Arch. Dev. Biol. 193, 267-282.
Öhlin, A.-K., Landes, G., Bourdon, P., Oppenheimer, C., Wydro, R., and Stenflo, J. (1988). Calcium binding to the epidermal growth factor homology region of bovine protein C. J. Biol. Chem. 263, 7411-7417.
Ozkaynak, E., Rueger, D. C., Drier, E. A., Corbett, C., Ridge, R. J., Sampath, T. K., and Oppermann, H. (1990). Op-1 cDNA encodes an osteogenic protein in the TGF-ß family EMBO J. 9, 2085-2093.
Padgett, R. W., St. Johnston, R. D., and Gelbart, W. M. (1987). A transcript from a Drosophila pattern gene predicts a protein homologous to the transforming growth factory family. Nature 325, 81-84.
Panganiban, G. F., Router, R., Scon, M. P., and Hoffmann, F. M. (1990b). A Drosophila growth factor homolog, decapentaplegic, regulates homeotic gene expression within and across germ layers during midgut morphogenesis. Development 110, 1041-1050.
Ray, R. P., Arora, K., Nüsslein-Volhard, C., and Gelbart, W. M. (1991). The control of cell fate along the dorsal-ventral axis of the Drosophila embryo. Development, in press.
Rees, D. J. G., Jones, l. M., Handford, P. A., Walter, S. J., Esnouf, M. P., Smith, K. J., and Brownlee, G. G. (1988). The role of ß-hydroxyaspartate and adjacent carboxylate residues in the first EGF domain of human factor IX. EMBO J. 7, 2053-2061.
Reuter, R., Panganiban, G. E. F., Hoffmann, F. M., and Scott, M. P. (1990). Homeotic genes regulate the spatial expression of putative growth factors in the visceral mesoderm of Drosophila embryos. Development 110, 1031-1040.
Roth, S., Stein, D., and Nüsslein-Volhard, C. (1989). A gradient of nuclear localization of the dorsal protein determines dorso-ventral pattern in the Drosophila embryo. Cell 59, 1189-1202.
Ruoslahti, E., and Pierschbacher, M. D. (1987). New perspectives in cell adhesion: RGD and integrins. Science 238, 491-497.
Rushlow, C., Doyle, H., Hoey, T., and Levine, M. (1987a). Molecular characterization of the zerknüllt region of the Antennapedia gene complex in Drosophila. Genes Dev. 1, 1268-1279.
Rushlow, C., Han, K., Manley, J. L., and Levine, M. (1989). The graded distribution of the dorsal morphogen is initiated by selective nuclear transport in Drosophila. Cell 59, 1165-1177.
Rushlow, C., and Levine, M. (1990) Role of the zerknüllt in dorsal-ventral pattern formation in Drosophila. Adv. Genet. 27, 277-307.
Sanger, F., Nicklen, S., and Couison, A. R. (1977). DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74,5463-5467.
Sato, Y., and Rifkin, D. B. (1989). Inhibition of endothelial cell movement by pericytes and smooth muscle cells: activation of a latent transforming growth factor-ß1-like molecule by plasmin during co-culture. J. Cell Biol. 107, 1199-1205.
Server, A. C., and Shooter, E. M. (1976). A comparison of the arginine esteropoptidases associated with the nerve and epidermal growth factor J . Biol . Chem . 251, 165-173.
Simon, J., Peifer, M ., Bender, W., and O'Connor, M. (1990). Regulatory elements of the bithorax complex that control expression along the anterior-posterior axis EMBO J. 9, 3945-3956
Spencer, F. A., Hoffman, F. M., and Gelbart, W. M. (1982). &laqno; Decapentaplegic: a gene complex affecting morphogenesis in Drosophila melanogaster. Cell 28, 451-461.
Spradling, A. C., and Rubin, G. M. (1982). Transposition of cloned P elements into Drosophila germane chromosomes. Science 218, 348353.
Stenflo,J., Lundwall,A.,and Dahlack, B. (1987). ß-Hydrosyasparagine in domains homologus to the epidermal growth factor precursor in vitamin K-dependent protein S. Proc. Natl. Acad. Sci. USA 84, 368-372.
Steward, R. (1987). dorsal, an embryonic polarity gene in Drosophila, is homologous to the vertebrate proto-oncogene, c-rel/. Science 238, 692-694.
Steward, R. (1989). Relocalization of the dorsal protein from the cytoplasm plasm to the nucleus correlates with its function. Cell 59,1179-1188.
St. Johnston, R. D., and Gelbart, W. M . (1987). Decapentaplegic transcripts are localized along the dorsal-ventral axis of the Drosophila embryo. EMBO J. 6, 2785-2791.
Stocker, W., Wolz, R. L., Zwilling, R., Strydom, D. J., and Auid, D. S. (1988). Astacus protease, a zinc metalloenzyme. Biochemistry 27, 5026-5032.
Tautz, D., and Pfeifle, C. (1989). A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translation control of the segmentation gene hunchback. Chromosoma 98, 81-85.
Taylor, J. M., Mitchell, W. M., and Cohen, S. (1974). Characterization of the binding protein for epidermal growth factor. J. Biol. Chem.249, 2188-2194.
Thisse, B., Stoetzel, C., Gorostiza-thisse, C., and Perrin-Schmitt, F. (1988). Sequence of the twist gene and nuclear localization of its protein in endomesodermal cells of early Drosophila embryos. EMBO J. 7, 2175-2183.
Thisse, C., Perrin-Schmit, F., Stoetzel, C., and Thisse, B. (1991). Sequence-specific transactivation of the Drosophila twist gene by the dorsal gene product. Cell 65, 1191 -1201.
Titani, K., Torte H.4., Hormel, S., Kumar, S., Walsh, K. A., Rodl, J., Neurath, H., and Zwilling, R. (1987). Amino acid sequence of a unique protease from the crayfish Astacus fluviatilis. Biochemistry 26, 222226.
Thomsen, G., Woolf, T., Whitman, M., Sokol, S., Vaughan, J., Vale, W. and Melton, D. A. (1990). Activins are expressed early in Xenopus embryogenesis and can induce axial mesoderm and anterior structures. Cell 63, 485-493.
van den Heuvel, M., Nusse, R., Johnston, P., and Lawrence, P. A. (1989). Distribution of the wingless gene product in Drosophila embryos: a protein involved in cell-cell communication. Cell 59,739-749.
Vieira, J., and Messing, J. (1987). Production of single-stranded plasmid DNA. Meth. Enzymol. 153, 3-11.
von Heijne, G. (1985). Signal sequences: the limits of variation. J. Mol. Biol. 184, 99-105.
Wakefield, L. M., Smith, D. M., Planders, K. C., and Sporn, M. B. (1988). Latent transforming growth factor-~ from human platelets: a high molecular weight complex containing precursor sequences. J. Biol. Chem. 263, 7646-7654.
Wakimoto, B. T., Turner, F. R., and Kaufman, T. C. (1984). Defects in embryogenesis in mutants associated with the Antennapedia gene complex of Drosophila melanogaster. Dev. Biol. 102, 147-172.
Wang, E. A., Rosen, V., Cordes, P., Hewick, R. M., Kriz, M. J., Luxenberg, D. P., Sibley, B. S., and Wozney, J. M. (1988). Purification and characterization of other distinct bone-inducing factors. Proc. Natl. Acad. Sci. USA 85, 9484-9488.
Wieschaus, E., Nüsslein-Volhard, C., and Jürgens, G. (1984). Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster . Zygotic loci on the X-chromosome and fourth chromosome. Roux's Arch. Dev. Biol. 193, 296-307.
Wozney, J. M., Rosen, V., Celeste, A. J., Mitsock, L. M., Whitters, M. J., Kriz, R. W., Hewick, R. M ., and Wang, E. A. (1988). Novel regulators of bone formation: molecular clones and activities. Science 242,1528-1534.
Zusman, S. B., Sweeton, D., and Wieschaus, E. F. (1988).shortgastrulation, a mutation causing delays in stage-specific cell shape changes during gastrulation in Drosophila melanogaster. Dev. Biol. 129, 417-427.
GenBank Accession Number
The accession number for the sequence reported in this article is M76976.
Return To Molecular Biology Main Page
© Copyright
2000 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu