
The Proteomics of FUS1 and YCL048W
"This web page was produced as an assignment for an undergraduate course at Davidson College."
The final step in my investigation of the yeast genes FUS1 and YCL048W was to examine how they fit into the proteomics of this organism. The following is a compilation of the data located on the web at various institutions of learning and industry. The data located on the web comes from databases of the yeast genome as well as from experiments in yeast proteomics. While information was not always available, it is important to realize that this discovery is just as crucial as having a plethora of data.
FUS1
The following is the protein sequence encoded by the FUS1 annotated gene:
MVATIMQTTTTVLTTVAAMSTTLASNYISSQASSSTSVTTVTTIATSIRSTPSNLLFSNV
AAQPKSSSASTIGLSIGLPIGIFCFGLLILLCYFYLKRNSVSISNPPMSATIPREEEYCR
RTNWFSRLFWQSKCEDQNSYSNRDIEKYNDTQWTSGDNMSSKIQYKISKPIIPQHILTPK
KTVKNPYAWSGKNISLDPKVNEMEEEKVVDAFLYTKPPNIVHIESSMPSYNDLPSQKTVS
SKKTALKTSEKWSYESPLSRWFLRGSTYFKDYGLSKTSLKTPTGAPQLKQMKMLSRISKG
YFNESDIMPDERSPILEYNNTPLDANDSVNNLGNTTPDSQITSYRNNNIDLITARPHSVI
YGTTAQQTLETNFNDHHDCNKSTEKHELIIPTPSKPLKKRKKRRQSKMYQHLQHLSRSKP
LPLTPNSKYNGEASVQLGKTYTVIQDYEPRLTDEIRISLGEKVKILATHTDGWCLVEKCN
TQKGSIHVSVDDKRYLNEDRGIVPGDCLQEYD
(http://genome-www2.stanford.edu/cgi-bin/SGD/seqTools)

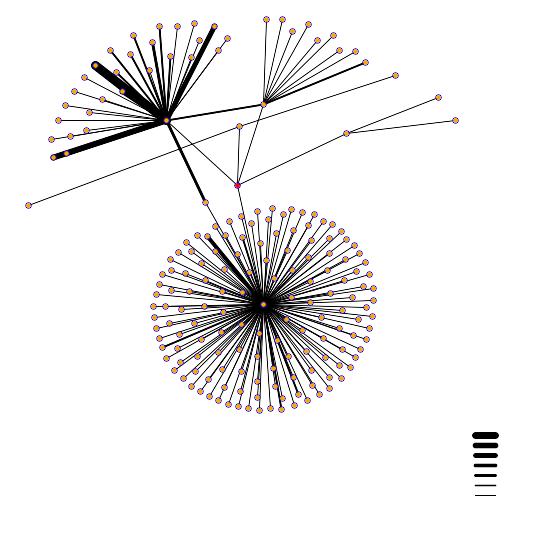
Figure 1: The above figure is a visual illustration of the proteins with which FUS1 interacts. FUS1 is represented by the red dot in the center of the figure and the other proteins are (clockwise from top left) ACT1, IQG1, SML1, and NUP116. The different amount of lines radiating from the centers indicates the varying number of proteins that interact with each. The key showing the different widths of lines indicates the certainty with which the connections are made. (http://dip.doe-mbi.ucla.edu/)
I then investigated the roles of each of the four proteins given in the above figure.
ACT1 (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=ACT1)
This protein has many roles in the yeast such as its identity as a structural
protein of the cytoskeleton, cell wall organization, bud growth, establishment
of cell polarity, and spindle orientation. In other words this is a major protein
involved in bud growth and mating, of which FUS1 is also a part. ACT1 seems
to be more of a pivotal protein in these processes as indicated by the fact
that a null mutant in this protein is not viable and by the first figure showing
that ACT1 is linked to many other proteins as compared to FUS1.
IQG1 (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=IQG1)
This protein occupies similar cellular roles as FUS1 in that it is a cytoskeleton
protein involved in actin filament organization and cytokenisis. It is also
a crucial protein as the null mutant is not viable.
SML1 (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=SML1)
This protein is thought to be involved in DNA repair, but the molecular function
is unknown.
NUP116 (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=NUP116)
The molecular function of this protein is one of a structural protein. NUP116
is involved in nuclear protein imports and exports.
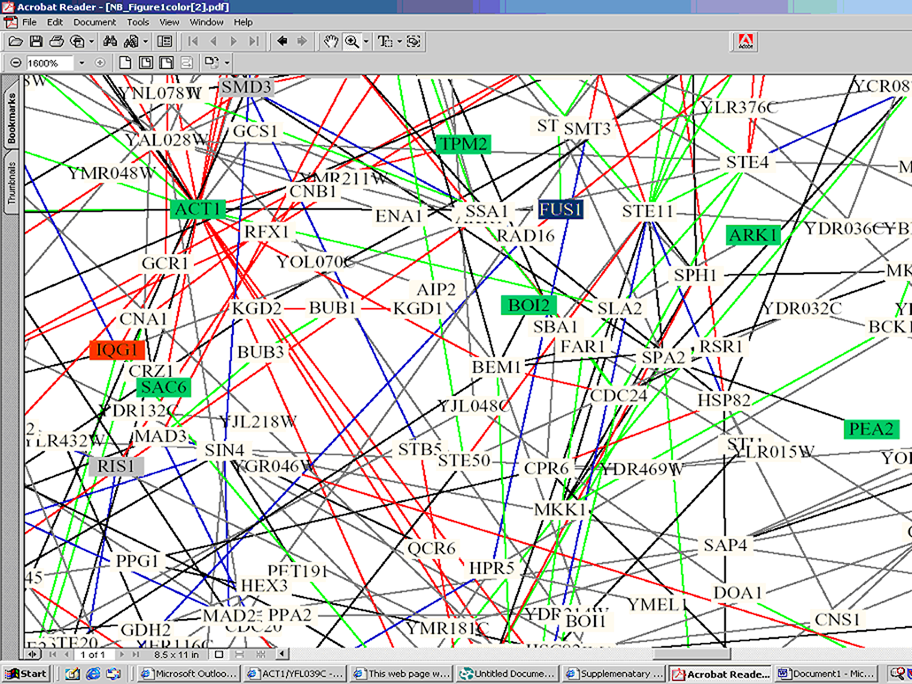
Figure 2: The above figure is a portion of an interaction map for yeast put together by Stan Fields and Peter Uetz at the University of Washington and CuraGen Corporation with the help of Benno Schwikowski from the Institute for Systems Biology. The overall figure shows 2,358 interactions among 1,548 proteins. The subset above is zoomed in such that FUS1 is blocked in blue toward the center and then to the left ACT1 and IQG1 can be found. This further illustrates that FUS1 is linked to several of the proteins presented in the dip doe figure. Below is the key for the color scheme of the boxes and lines.
This is the color key for figure 2.
· red = cellular role and subcellular localization of interacting proteins
are identical;
· blue = localizations are identical but functions differ;
· green = cellular roles are identical but localizations differ;
· black = cellular role and localization are different or unknown
Here is the color code for the boxes in this figure:
· blue = membrane fusion
· gray = chromatin structure
· green = cell structure
· yellow = lipid metabolism
· red = cytokinesis
In addition, I visited the web site with the combined efforts of Stan Fields and Peter Uetz, and I did not find any new information but rather affirmed that Fus1 does interact with ACT1 (http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastList).
At the Yale
Gerstein Lab website, there was little additional information available.
The website did show that they had no protein chip data available. But the Yale
data enforces the notion that this protein is membrane related.
According to the Stanford website link
to the PDB BLAST report, the protein product of YCL027W (FUS1) has no known
3D structure and does not show significant sequence similarity to any protein
of known 3D structure.
According to Protein Explorer there is no structure information (http://www.umass.edu/microbio/chime/explorer/#search).
There was no information at the Protein Data Bank (http://www.rcsb.org/pdb/status.html).
YCL048W
I next began my quest for information about the proteomics of my non-annotated gene. Unfortunately, my search was not terribly fruitful.
Below is the protein sequence encoded by the YCL048W gene:
MNRITRKSCLFAIIFASLFVTHALGAAIDPPRRPHNVKPFHNGNLELQRRANEPFFEIDV
KSLNTNSPISELCKKDLHVIESSHDLFHLQNQCEFILGSLKVTNYDSNILDLNSLRAIGG
DLIIQDSPELIRIQAGNLNKIEGLFQLQGLTSLVSVEIPTLKFCQSLEWKVVPILNYVSM
DSQNIEIIKDIVISDTSLANIENFNKVQEIDTFNINNNRFLETIHSNVKTIRGQFSVHAN
AKELELEMPHLREVENITIRDTSLVYLPQLTKVKSSLEFIENYFYELNLNNLQKIGGTLG
IINNVNLIKVNLENLTDIQGGLMIADNESLEDITFLPNLKQIGGAIFFEGSFKDIMFDSL
KLVKGSAFIKSSSNVLDCNKWTNPSNGRSIIRGGKFTCISGKKENTLNVKQDGTIIEKGY
KDLTQEGEDSKKRVISKYANSANPSMQLDPLLFGTCLVAMLLF
(http://genome-www2.stanford.edu/cgi-bin/SGD/getSeq?map=p3map&seq=YCL048W&flankl=0&flankr=0&rev=)
After searching the interaction map of selected yeast genes as mentioned above, I could not locate this gene in the sampling.
I next went to Stanford's Function Junction and was able to locate a hint of possible protein-protein interactions with my gene. Below is a summary of my findings.
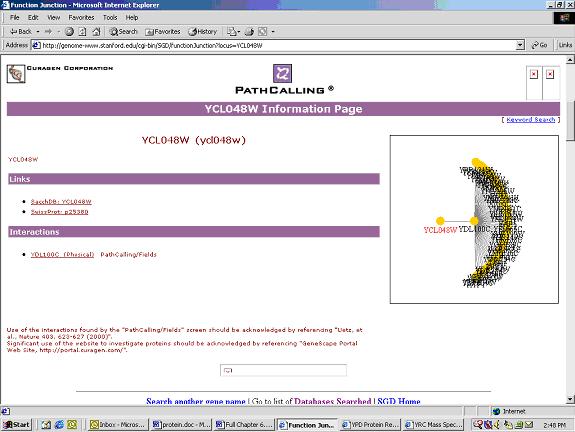
Figure 3: This is the interaction figure from the Yeast PathCalling Home Page sponsored by the University of Washington and CuraGen. This shows that YCL048W interacts with the protein YDL100C. This is the only protein shown to interact with YCL048W (http://genome-www.stanford.edu/cgi-bin/SGD/functionJunction?locus=YCL048W).
After investigating the role of YDL100C, I could find little information about this ORF which is also not annotated. YDL100C is a hypothetical ORF with no known biological or molecular function (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=YDL100C)
Also from the above Function Junction website, I found three proteins that show sequence similarity to YCL048W. These proteins were in the section comparing the worm and yeast genome. These three proteins are ECM33, YDR055W, and SPS2.
After investigation of the roles of each protein,
ECM33: This gene was shown to be involved in cell wall biogenesis (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=ecm33).
YDR055W: The gene product has been detected among the proteins secreted
by regenerating protoplasts, but this gene is mostly unknown (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=ydr055w).
SPS2: This gene is turned on late in meiosis. The molecular process
is unknown (http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=sps2).
Gerstein Lab (http://bioinfo.mbb.yale.edu/genome/smartlink.cgi?id=ycl048w) had no results.
I then visited the website sponsored by Stan Fields and found no useful information in his study of protein-protein interactions (http://depts.washington.edu/%7Eyeastrc/th_11.htm).
Experiment 1
According to the Function Junction database specifically the SGD worm-yeast
protein comparison, there are three genes with which YCL048W shows sequence
similarity. These genes include ECM33, YDR055W, and SPS2. ECM33 and SPS2 are
annotated genes and thus would provide some insight if YCL048W did have protein-protein
interaction with them. I believe that a yeast two-hybrid (Y2H) method would
be an excellent and straightforward way to investigate these possible interactions.
The Y2H method is designed specifically to find proteins that physically interact
with one another in the bait and prey set up.
In order to do this, the bait cells are composed of the gene of interest, YCL048W,
fused to the DNA binding domain (DBD). This construct is encoded by a plasmid
also found in the cell. This bait cell will remain constant. The bait is then
combined with different prey cells in order to deduce which proteins interact.
Prey-1 is composed of the gene ECM33 fused onto the activation domain (AD)
which is capable of activating RNA polymerase.
Prey-2 is composed of the gene YDR055W fused onto the same AD.
Prey-3 is composed of the gene SPS2 fused onto the same AD.
Prey-4 serves as a control and is composed of YDL100C fused onto the same AD.
This is a control because it is known that YCL048W interacts with this protein
and so there should be transcription.
Prey-5 also serves as a control and is composed of UBR1 fused onto the same
AD. This should serve as a control because UBR1 is an outlier protein that is
shown to interact with only one other protein in the yeast proteome and this
protein is not the one encoded by YCL048W. Hence, there should be no transcription.
When the bait and prey plasmids are combined into the same cell, if the prey ORF-encoded protein can bind to the bait protein then the RNA polymerase should be able to transcribe the reporter gene of HIS3. Cells will be plated on a medium without the protein histidine. Without the HIS3 protein, cells cannot grow in the absence of histidine. If a cell colony grows, then those proteins interact.
Experiment 2
In the last investigation using microarray data, I predicted that YCL048W was a hypothetical ORF involved in the signaling of conjugation. It is known that pheromones are a part of the mating process, and the presentation of pheromones elicit the beginning of conjugation. This experiment can predict when exactly transcription of YCL048W begins as indicated by the time lapse of pheromone presentation which should mimic the presentation during mating.
First, yeast will be plated such that the medium experiences increases in pheromone
concentration. The same yeast will be used throughout but aliquots will be taken
at the differing pheromone concentrations. Then, the YCL048W protein will be
extracted from the cell samples and plated in wells much like the ones used
in Mark Snyder's lab where individual wells contain proteins under different
conditions. Then the plate will be submerged in the protein produced by YDL100C
in order to interact with each well. This experiment relies on the fact that
YCL048W will interact with YDL100C when the gene is turned on in the cell. The
conditions under which it binds to YDL100C can be the conditions, or the concentration
of pheromone, in which it is activated. This provides information about when
this protein is activated and may shine light on the function in the cell.
References
Campbell, A. Malcolm. "Chapter Six: Proteomics." Genomics, Proteomics, and Bioinformatics. 2001.
Database of Interacting Proteins. UCLA. 1999. <http://dip.doe-mbi.ucla.edu/dip/represent_JAVA.cgi?PKEY=1041> Accessed on 11-08-01.Function Junction. Saccaromyces Genome Database. 2001. <http://genome-www.stanford.edu/cgi-bin/SGD/functionJunction>
Accessed on 11-08-01.
Protein Data Bank. 2001. <http://www.rcsb.org/pdb/index.html> Accessed on 11-08-01.
Proteome Database. "YPD." 2001. <http://www.proteome.com/databases/YPD/YPDsearch-quick.html>
Accessed on 11-08-01.
For questions or comments contact elsellars@davidson.edu