My Favorite Yeast Protein
This site was created for an undergraduate Genomics course at Davidson College
SSL2!!!
The first place I went to find interaction information for SSL2 was the Pathcalling Database. From the figure below it is apparent that this site found only one protein interaction for SSL2.

DIP Data
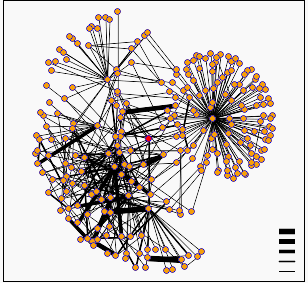
Figure 2: Diagram of protein interactions surrounding yeast gene SSL2. SSL2 is represented by the red dot in the center of the figure. This figure provided by the Database of Interacting Proteins (DIP)
When I searched the DIP database for proteins that interact with SSL2 I found the graph of interactions shown above. In this diagram the darker lines represent increased confidence in the predicted protein interactions. A list of the proteins interacting with SSL2 are available from DIP. The first thing I noticed upon examining this list is that the protein MSL1 that was predicted by the Pathcalling database to interact directly with SSL2 was not present. I then looked through the functions of the interacting proteins and for the most part they fit very well with the function of SSL2 as described in Web Assignment #1. Remember, RAD25 is another name for SSL2!!
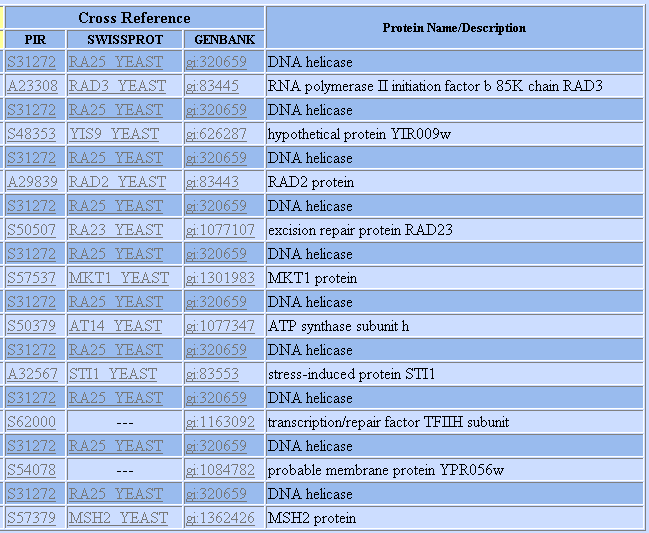
Figure 3: List of proteins shown to interact with SSL2 according to the DIP database. Graph provided by DIP.
We know from previous experiments that RAD2, RAD23, and RAD3are involved in nucleosome excision repair, and thus are not surprised to find out they interact with SSL2. In addition, one of the unannotated genes shown to interact with SSL2 (gi 1163092) takes part in the same TFIIH subunit that SSL2 does. We thus see that the interaction data is able to confirm much of what we already knew about SSL2's role in NER. Furthermore, an association with stressed induced protein STI1 perhaps makes sense since SSL2's role as a DNA repair helicase is upregulated when yeast face stress, as seen in the microarray experiments before.
The next search I conducted utilized the Interactions GRID Database. I linked to this page from the SGD page for SSL2. This interaction database described four protein interactions for SSL2. Of these four proteins, three of them (MKT1, AT14, and STI1) were identified by the DIP database. GRID showed interaction with MSL1 just as Pathcalling did.
I also looked at the "Benno Figure 1" available at The Genomics Place under Chapter 6 links. I zoomed in and located SSL2 and was able to observe a few of the interactions described by the previous database searches. Specifically I located Rad3 and MSL1 which are both visible in the following figure.
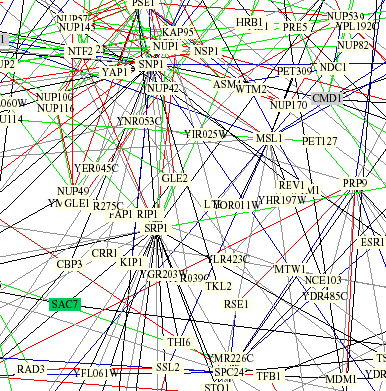
Figure 4: Excerpt of protein interaction grid from "Benno Figure 1". The red line connecting SSL2 and RAD3 represents that both of the interacting proteins play the same subcellular role and localizations. The blue line connecting MSL1 and SSL2 refers to the fact that both proteins are localized to the same place but serve different functions.
Using the key to interpret what the interaction lines means sheds some light on the interaction of SSL2 and MSL1. Before I was confused as to why these two proteins with very different function were showing up together, but it appears that they are showing up as interacting due to very similar localization. However, I found contradictory information at the MIPS site (discussed below) that referred to the interaction between SSL2 and MSL1 as a physical interaction.
Another site that proved very helpful was the MIPS Comprehensive Yeast Genome Database page. Unfortunatly I cannot link to the results page for SSL2. Please click the above link, then type SSL2 in the top right Gene/ORF search box. Click 'Go', then click the link that pops up. This provides useful data regarding the function of SSL2. Of new importance is the information on many SSL2 mutant strains. We learn from this data 1) without the C-terminus of SSL2, yeast are hyper sensitve to UV-light. 2) SSL2, or its functional complex, may be heat dependent. 3) An inability of SSL2 to perform DNA excision repair does not necesarily lead to lack of transcription.
Another helpful piece of SSL2 info I found at this site was an interactive diagram of the RNA Polymerase II Holoenzyme. The active diagram of the figure below can be found by clicking the link below "Scheme" on the MIPS page link above. Clicking on the TFIIH subunit sends you to a link of all of the subunits, including SSL2.
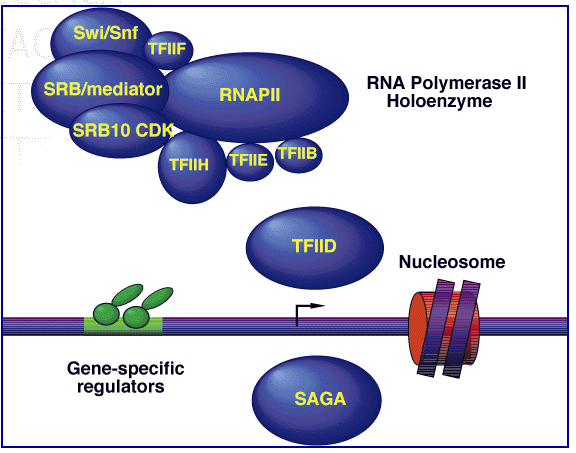
Figure 5: Model of RNA Polymerase II Transcription Initiation Machinery.The machinery depicted here encompasses over 85 polypeptides in ten (sub) complexes: core RNA polymerase II (RNAPII) consists of 12 subunits; TFIIH, 9 subunits; TFIIE, 2 subunits; TFIIF, 3 subunits; TFIIB, 1 subunit, TFIID, 14 subunits; core SRB/mediator, more than 16 subunits; Swi/Snf complex, 11 subunits; Srb10 kinase complex, 4 subunits; and SAGA, 13 subunits. This figure provided by Comprehensive Yeast Genome Database.
Unfortunatly the PROWL database is down.
YIL137C!!!
The first database I searched was Pathcalling. This database provided no information on protein interactions for YIL137C.
The next site I tried was the Interactions GRID Database. This link shows two proteins shown to interact with YIL137C. The first of these is YTM1, a cytoskeletal binding protein. This protein takes part in the building of the cytoskeleton. This does not fit any of the predictions or data previously collected for YIL137C. The second protein is JSN1 (aka PUF1) an mRNA binding protein that takes place in mRNA catabolism. Because YIL137C is predicted to be an aminopeptidase, specifically a zinc protease, it appears that this interaction with another protein that takes place in catabolism may shed light on the function of YIL137C. However, while it may be possible that despite YIL137C being involved in protein catabolism and PUF1 being involved in mRNA catabolism they are functionally related, I have never studied anything regarding the relationship between breaking down proteins and breaking down mRNA's. Because the relationship between PUF1 and YIL137C was discovred using the yeast two-hybrid test it appears that these two proteins physically interact. While my previous web pages have described domains and possible functionalities of YIL137C, it is still worth checking out the MIPS page on this ORF. As before, type in YIL137C in the upper right and click 'Go', the click the link that will pop up. Furthermore, taking a second look at the microarray data I presented on Web Page #3 shows a number of co-expression patterns with catabolic proteins... something I overlooked the first time through.
I next searched the Y2H database and found no information on this ORF.
Utilizing the DIP database provided interaction information relating to the same two genes I found using the GRID database. Below I include the map of interactions, but just for show really.
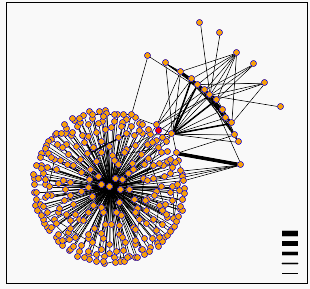
Figure 6: Digram of protein interactions around hypothetical yeast ORF YIL137C. YIL137C is represented by the red dot at the center of the diagram (though I'll mention that being slightly colorblind, and human, I find this representation next to useless). Diagram provided by DIP.
Using the DIP database I found the same two protein interactions predicted by the GRID database. These are YTM1 and JSN1/PUF1. Check out the interaction page here. We learn that the JSN1 interaction was predicted by yeast two-hybrid tests and YTM1's interaction was found by immunoprecipitation.
I attempted to find interaction data via the "Benno Figure #1" pdf file, but YIL137C is not included in this particular diagram.
I also searched the ExPasy site to try and find some 2D gel information on this ORF, but the database did not include my ORF.
In addition I checked out the Triples Database and was unable to find information on my protein.
PROWL database is unavailabe.
Overall I have learned very little new information about YIL137C by studying the interaction databases. The two proteins that it was shown to interact with are unrelated and certainly not clearly related to the predicted function of YIL137C. However, since I didn't search the MIPS database for YIL137C until this assignment, and from this site I found most of my predictions regarding YIL137C confirmed, I do have increased confidence in my predictions regarding the probably role of YIL137C as a zinc protease. Below I include a few experiments that would improve our knowledge of this ORF.
Experiments!!!
Over the last couple of assignments I have made a number of predictions about the structure and function of yeast hypothetical ORF YIL137C. One issue I have not addressed, or found evidence about, is the cellular localization of YIL137C. When I first examined this protein I utilized a Kyte-Doolittle Hydropathy Plot. This plot had two peaks that suggested it may be a transmembrane protein. However, the MIPS database describes YIL137C's similarity to Ape2, a zinc protease, and Ape2 is an extracellular secreted protein that takes part in protein degredation. I would thus perform protein localization experiments utilizing the detergent Triton X-114. This detergent allows a seperation of proteins into two layers, one layer includes transmembrane proteins, and the other contains membrane bound proteins. I would then run the proteins on a gel and look for the correct size band, then test by sequencing until I found my ORF. I would then have determine whether or not it is transmembrane or not. Once I determined whether or not YIL137C is membrane bound, I would use flourescently tagged antibodies to this protein and apply them to yeast cells. Upon viewing the cells I could determine where this protein is located within the cell and thus gain insight into its possible function. A final experiment I would like to perform a functional assay using YIL137C in order to test whether or not it functions as a zinc protease. This experiment would require the purification of a large amount of this protein. In addition I would have to learn more about the action of a zinc protease and I could do this by studying Ape2 or other such zinc proteases. The difficulty in performing this experiment would be lack of controls. While I could look at Ape2's results, it would be poor experimental design to compare YIL137C's functional assay results to that of another protein. However, if I put in another protein that I knew played no role in proetin catabolism (actin for example), I could use this for a baseline against which I could compare YIL137C's functionality. If the results showed increased catabolic activity in YIL137C over actin, then I would gain confidence in calling it a probably zinc protease; however, this would not yet prove anything definitively.
One important issue I would like to clear up regarding SSL2 is why the databases show it to interact with some (RAD3, RAD23, RAD2) proteins involved in the RNA Polymerase II Holoenzyme, but not all of the proteins involved in the TFIIH transcription factor subunit. These proteins include the following:

I would thus like to utilize the yeast two-hybrid experiment to determine whether or not SSL2 physically interacts with the other 8 proteins in this subunit. This would involve using SSL2 fused to the DNA binding domain as the bait. The prey proteins would be fused to the activation domain. For this test each of the of other 8 proteins listed above (one at a time) would be the prey proteins. If SSL2 interacts with any of these other proteins a reporter gene will be transcribed and this will be viewable to the reseearcher. Performing this test is important because while it is known that all of these proteins work together in order to perform their role in Nucleotide Excision Repair, it is not understood how they come together to perform this task. (See Web Assignment #2). It also be important to perform yeast two-hybrid experiments looking at all interactions between these 9 proteins. Perhaps understanding this entire units interactions would lead to a model of how they group together at the site of damaged DNA. Finally this could be expanded to look at the entire RNA Polymerase II Holoenzyme. In addition to looking at protein interactions I would like to study the microarray data for each of the gene in this holoenzyme. Just as we saw on Review 2 that protein cascades can be elucidated from microarray data, I would like to see if this is possible in the case of this holoenzyme. Perhaps we would see which subunits are transcribed first and from this order, combined with interaction data, we may gain insight into the routes by which many protein complexes come together to perform a unified task.
Contact Kevin James with any comments or questions.