
This page was produced as an assignment for an undergraduate course at Davidson College.
Yeast Gene Expression: MRS2 and YOR331C Revisited
Introduction:
The goal of this web page is to utilize public databases in order to find the gene expression for two types of genes using DNA Microarrays. The species that the two genes will be analyzed from is Saccharomyces cerevisiae. One annotated gene, MRS2, and one non-annotated gene, YOR331C, will be reevaluated using DNA microarrays found in public databases. To learn more about the genes MRS2 and YOR331C link to the previous web assignment (Yeast Genes).
To understand how DNA Microarrays work, link to this page. DNA microarrays measure the expression of every gene in the genome! With this data, researchers can subject the genome to many different experimental conditions and learn how the genome responds. All of the data is already waiting to be discovered, all researchers have to do is put the pieces together to learn how the genome works.
For the expression of genes on this web page, the colors green and red will be used to demonstrate repression and induction respectively. The brighter a color is, the more highly induced or repressed a gene is.
MRS2 Expression
As my previous web assignment informed, MRS2 is a magnesium ion transporter protein in the cell membrane of mitochondria in yeast cells. More simply, this gene encodes for an integral membrane protein of an organelle that produces ATP for yeast cells. As a mitochondria-related gene, it seems clear that MRS2 should be a highly regulated gene.
To begin my search for MRS2 expression, I searched the Expression Connection web site for MRS2. My results were not what I expected. Since MRS2 encodes a mitochondrial ion transport gene, it seemed clear to me that this gene would be strictly regulated since energy production is a critical cellular function, especially under stressed conditions. One induced expression that I found for MRS2 was the histone depletion experiment by JJ Wyrick et al (histone depletion).
All figures from Expression Connection. Permission pending.
Expression Scale

Figure 1: This figure shows the expression scale used to determine gene repression and induction (SGD 2002).
Effect of Histone Depletion on MRS2
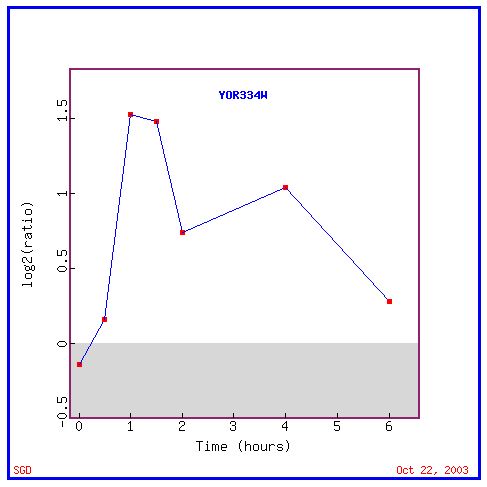
Figure2: This graph shows the induction of MRS2 over 6 hours while the yeast cell was histone depleted.
The graph gives a clear example of how DNA microarray data can be used individually to show the expression of a gene under a certain experimental condition. The rest of the figures will use only color coded lines to demonstrate the same data in a much simpler way.
The next step to understanding various genes and their expression in DNA Microarrays lead me to analyze various datasets with different experimental conditions. Under different experimental conditions, MRS2 should be clustered around similar genes with similar functions. Using this "guilt by association" method of finding similar genes, I should be able to find various datasets that have similar genes. Of course, the possibility that some genes share a common promoter with MRS2 is also one possible reason for expressions to be similar.
Expressions in Histone-Depleted Genes
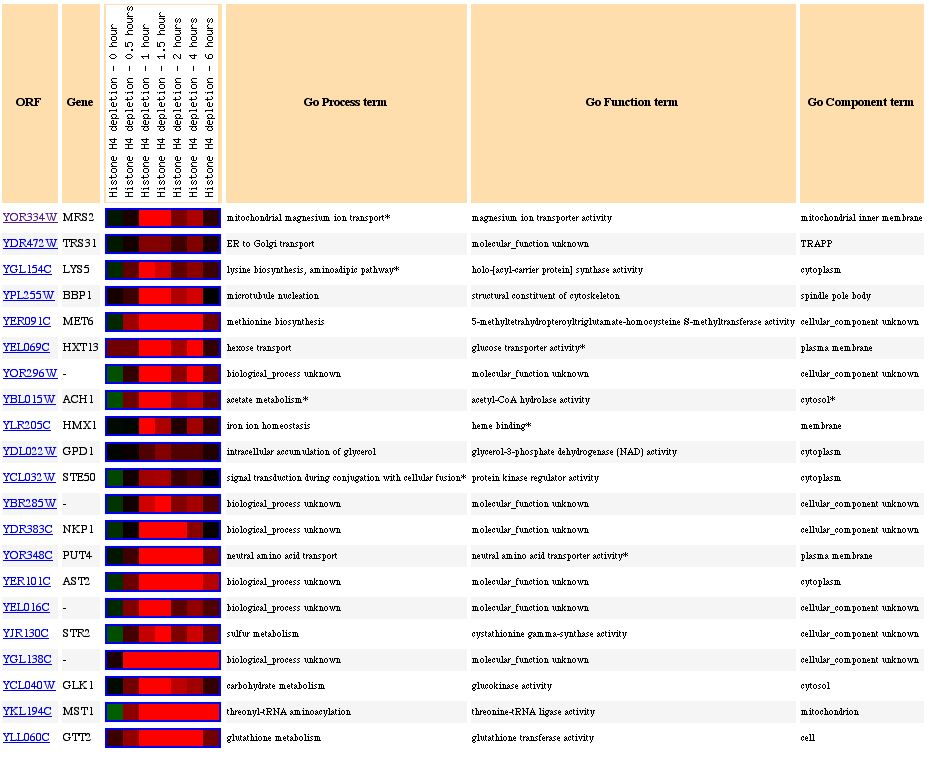 Figure
3: This figure shows genes that have similar expression profiles for histone-depletion.
Figure
3: This figure shows genes that have similar expression profiles for histone-depletion.
For Figure 3, notice the diversity in the types of genes that are presented by genes that are clustered by similar expressions. Clearly, not all of the genes in this figure have similar functions to MRS2, however, some similarity does exist for genes such as TRS31, HXT3, HMX1, and PUT4. All of these genes perform some type of transport between membranes, just like MRS2, which encodes for a magnesium ion transporter.
Expressions in Genes Undergoing Diauxic
Shift
Figure 4: This figure shows genes that have similarity in expression to MRS2, namely in their repression under the stress of diauxic shift.
Diauxic shift is the shift from aerobic respiration to anaerobic fermentation in yeast cells (JL DeRisi et al, 1997). Because ATP is produced aerobically by respiration in the mitochondria and the cell is shifting towards anerobic fermentation, the repression of MRS2 is not surprising.
Once again similar transport protein genes are clustered close to MSR2 such as SAM3 and BET3. Also, many genes have their cellular component in mitochondria for this clustering, such as MRPL50, ATP12, MRPL3 and SSQ1. In this circumstance, the idea of a circuit might be one possible explanation for the frequency that genes occur in mitochondria. The idea of a circuit involves the presumption that certain genes might be repressed at the same time by a certain mechanism. In this case, the diauxic shift might cause repression of certain mitochondria-linked genes that are regulated by a circuit.
Expression in Genes Exposed to Various
Alpha Concentrations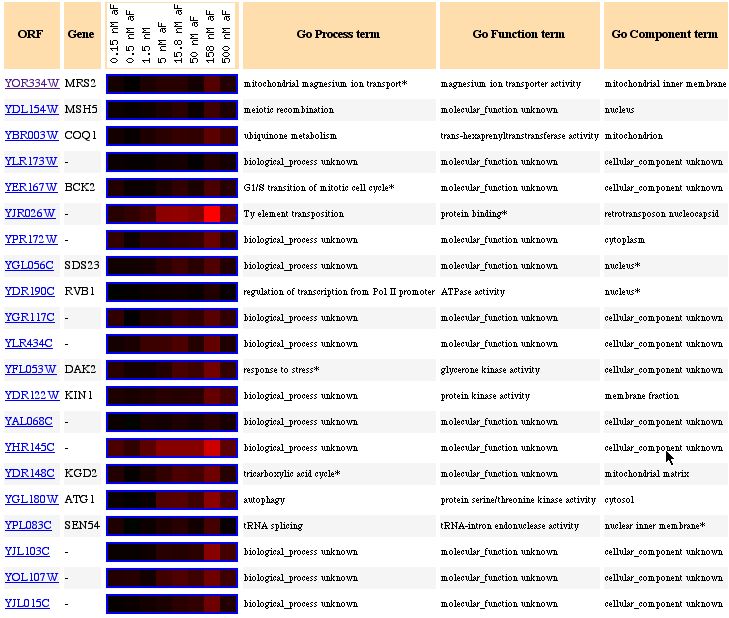
Figure 5: This figure shows an increasing in induction when yeast genes are exposed to various alpha concentrations.
In this figure, the data seems to give an interesting conclusion. Only one gene's cellular component is in mitochondria and no genes are present that have any type of transport activity. This clustering of genes with similar expressions shows that the guilt by association idea might not always work. Just because similar expression exists between two genes doesn't necessarily mean that the genes share the similar functions. When trying to find a trend in related function or similar cellular locations for genes, it seems very probable that with 6,000 genes
__________________________________________________________________________________________________________________________
YOR331C
From the data and results I posted in the last web assignment, YOR331C seemed to have a very close cellular role to MRS2(Favorite Yeast Gene). In my analysis of YOR331C using DNA microarrays, I will specifically look for trends in the expressions of genes clustered around YOR331C under various experiemental conditions. After I have presented the datasets, some type of cellular role for YOR331C should be evident. Then, I should be able to comment upon the hypothesis that MRS2 and YOR331C have similar cellular roles.
All figures taken from Expression Connection. Permission pending.
Expression for Genes undergoing DNA-Damage
 Figure
6: This figure shows that YOR331C shares a very similar expression profile with
VMA4, a vacuolar membrane protein involved in hydrogen transportation.
Figure
6: This figure shows that YOR331C shares a very similar expression profile with
VMA4, a vacuolar membrane protein involved in hydrogen transportation.
To view a continuation of figure 6 link to expression connection.
After DNA damaging elements are introduced into the cell, the cell cycle is arrested and the cell concentrates on DNA repair. During this type of cellular function, YOR331C is slightly induced and then later repressed. VMA4 shows a similar expression profile. I hypothesize that VMA4 at first carries out its normal function, transporting hydrogen until even this cellular function is arrested and DNA repair is fully focused on.
Expression with Genes Exposed to Alpha Factor over time
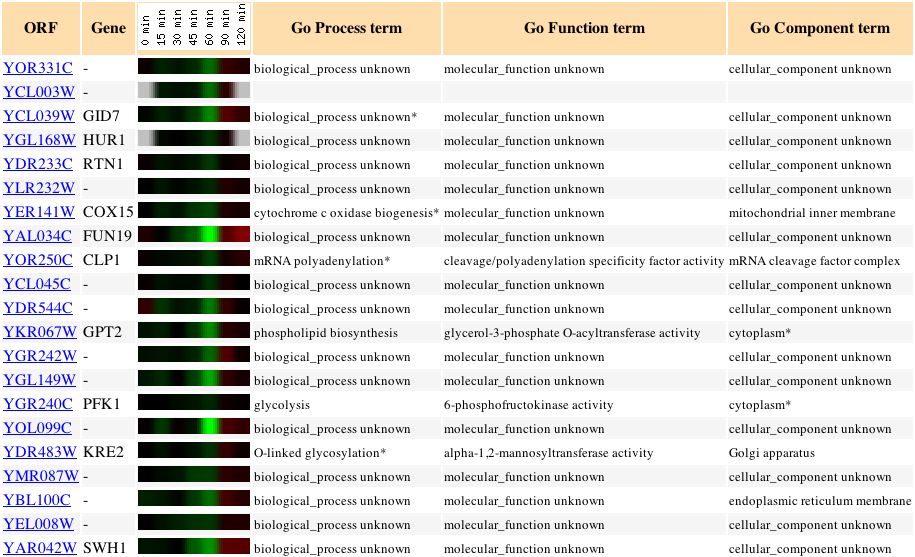
Figure 7: This figure shows genes clustered around YOR331C when yeast cells are exposed to alpha factor over a period of 2 hours.
Alpha factor is a signaling pheromone to other yeast cells that a particular yeast cell is interested in reproducing sexually. This expression shows how YOR331C gives a very weak expression during the time when the cell is exposed to alpha factor. One important gene to notice in this figure is COX15, a gene whose cellular role is in the mitochondria. If this gene has any functional relationship or similarity in cellular location, then YOR331C might be similar to MRS2.
Expression of Genes with Different Zinc Concentrations

Figure 8: This figure shows gene expression when yeast cells are exposed to zinc, a vital ion for yeast cell homeostasis.
This figure shows one important gene, VMA6. Because VMA4 had one similar expression profile, it seems likely that other VMA proteins would share similar expression profiles for certain conditions. The idea that VMA proteins might also share a similarity in function with YOR331C is another possibility. NFU1 is another gene that is clustered with YOR331C that might show a similarity in function and would support the more basic hypothesis that YOR331C is some type of transporter. For instance, YOR331C is similar to MRS2 because it could functionally be some type of transporter protein like NFU1 or the VMA proteins.
Expression of Genes Undergoing Sporulation
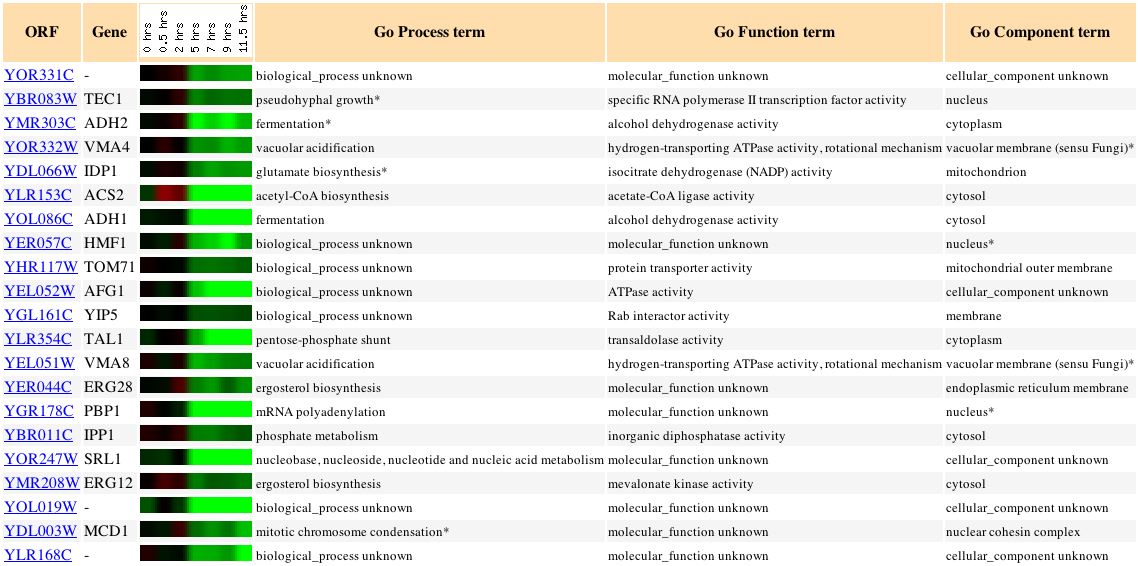
Figure 9: This figure shows the expression of genes in yeast cells that are undergoing sporulation.
This figure reinforces the idea that VMA genes such as VMA4 and VMA8 are linked in expression to YOR331C or perhaps linked in function.
Further Reinforcement of VMA Gene Expression Clustering with YOR331C
Reinforcement for similar expression between VMA genes and YOR331C is evident under other experimental conditions. For the environmental changes dataset, only VMA4 and YOR331C are clustered together (Environmental Changes). This data bolsters the idea that VMA4 and YOR331C might be linked functionally or through some type of expression mechanism that controls both VMA genes and YOR331C. Also, viewing the experiment in which expression is regulated by the PHO pathway shows VMA4 and VMA6 clustered with YOR331C.

Figure 10: This figure shows that VMA4 and VMA 6 are clustered with YOR331C when expression is regulated by the PHO pathway.
Not only do the two VMA genes cluster with YOR331C in this dataset, but these two VMA proteins are the first two that cluster with YOR331C meaning that they have the closest correlation to YOR331C.
__________________________________________________________________________________________________________________________________________________
Conclusion
Because there is an overwhelming data that support the similarity in expression between YOR331C and VMA genes, I believe that either YOR331C has a similar cellular role to VMA genes or shares a promoter with VMA genes. Since VMA genes are a type of ion transporter, my hypothesis that YOR331C's cellular role is similar to MRS2's cellular role is not completely unreasonable. If YOR331C is a hydrogen ion transporter or a transporter protein of any kind, then it does bear some similarity to MRS2.
When further investigation is made into the chromosomal location of VMA, the SGD database gives a physical map of chromosome XV and VMA is a neighbor to both YOR331C and MRS2 (Physical Map). For this reason, the idea that VMA might share a promoter with YOR331C seems like the most plausible explanation for their similarity in expression.
References
Chu S, DeRisi J, Eisen M, Mulholland J, Botstein D, Brown PO, Herskowitz I (1998) The transcriptional program of sporulation in budding yeast. Science 282(5389):699-705
DeRisi JL, Iyer VR, Brown PO (1997) Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278(5338):680-
Dolinski, K., Balakrishnan, R., Christie, K. R., Costanzo, M.
C., Dwight, S. S., Engel, S. R., Fisk, D. G., Hirschman, J. E., Hong, E. L.,
Issel-Tarver, L., Sethuraman, A., Theesfeld, C. L., Binkley, G., Lane, C., Schroeder,
M., Dong, S., Weng, S., Andrada, R., Botstein, D., and Cherry, J. M. "Saccharomyces
Genome Database"
http://www.yeastgenome.org/ 24 Oct 2003.
Expression Connection 2003. <ttp://genome-www4.stanford.edu/cgi-bin/SGD/expression/expressionConnection.pl>
Ferea TL, Botstein D, Brown PO, Rosenzweig RF (1999) Systematic changes in gene expression patterns following adaptive evolution in yeast. Proc Natl Acad Sci U S A 96(17):9721-6
Gasch AP, Huang M, Metzner S, Botstein D, Elledge SJ, Brown PO (2001) Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Mol Biol Cell 12(10):2987-3003
Lyons TJ, Gasch AP, Gaither LA, Botstein D, Brown PO, Eide DJ (2000) Genome-wide characterization of the Zap1p zinc-responsive regulon in yeast. Proc Natl Acad Sci U S A 97(14):7957-62
Ogawa N, DeRisi J, Brown PO (2000) New components of a system for phosphate accumulation and polyphosphate metabolism in Saccharomyces cerevisiae revealed by genomic expression analysis. Mol Biol Cell 11(12):4309-21
Roberts CJ, Nelson B, Marton MJ, Stoughton R, Meyer MR, Bennett HA, He YD, Dai H, Walker WL, Hughes TR, Tyers M, Boone C, Friend SH (2000) Signaling and circuitry of multiple MAPK pathways revealed by a matrix of global gene expression profiles. Science 287(5454):873-80
Wyrick JJ, Holstege FC, Jennings EG, Causton HC, Shore D, Grunstein M, Lander ES, Young RA (1999) Chromosomal landscape of nucleosome-dependent gene expression and silencing in yeast. Nature 402(6760):418-21
Email Ben with Questions or Comments.
This page was produced as an assignment for an undergraduate course at Davidson College.