My Annotated Yeast Gene
SSD1 (YDR293C)
The SSD1, Suppressor of SIT4 Deletion gene of Saccharomyces cerevisiae encodes
a 160 kDa cytoplasmic protein that can suppress mutations in a number of other
genes (Chen et al., 1998). The gene is located on Chromosome IV, and is encoded
on the crick strand. The encoded protein, found in the cytoplasm, has the ability
to bind to RNA as well as single stranded DNA, and belongs to the ribonuclease
II (RNB) family, involved in RNA modification (Usono et al., 1997). The protein
is implicated in the control of the cell cycle G1 phase (Sutton et al., 1991).
SSD1 protein participates in cell wall organization and biogenesis (SGD, 2003).
Several alleles of SSD1 exist in different yeast strains. SSD1 can suppress
the lethality due to deletion of SIT4, and partially defects due to BCY1 disruption
(Swiss-Prot, 2003). SSD1 gene is also involved in the tolerance to high concentration
of Ca2+ (Tsuchiya et al., 1996). Yeast is not usually pathogenic to healthy
individuals but increasingly has been isolated from immunocompromised patients.
Knocking out SSD1 causes increased virulence of yeast by changes in the composition
and cell wall architecture of the yeast cell surface. The hypervirulent ssd1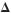 /ssd1
/ssd1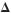 strain has been shown to cause lethal infections and accelerated death in mice,
and elicits greater proinflamatory cytokine induction in macrophages. Loss of
SSD1 showed many phenotypic differences in the cell wall from wild type yeast.
The ssd1
strain has been shown to cause lethal infections and accelerated death in mice,
and elicits greater proinflamatory cytokine induction in macrophages. Loss of
SSD1 showed many phenotypic differences in the cell wall from wild type yeast.
The ssd1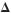 /ssd1
/ssd1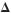 strain showed dramatically increased response to osmotin (plant anti-fungal
compound), changes in the composition of cell wall polysaccharides, and greater
sensitivity to cell wall damaging agent, calcoflour white (Wheeler et al., 2003).
strain showed dramatically increased response to osmotin (plant anti-fungal
compound), changes in the composition of cell wall polysaccharides, and greater
sensitivity to cell wall damaging agent, calcoflour white (Wheeler et al., 2003).
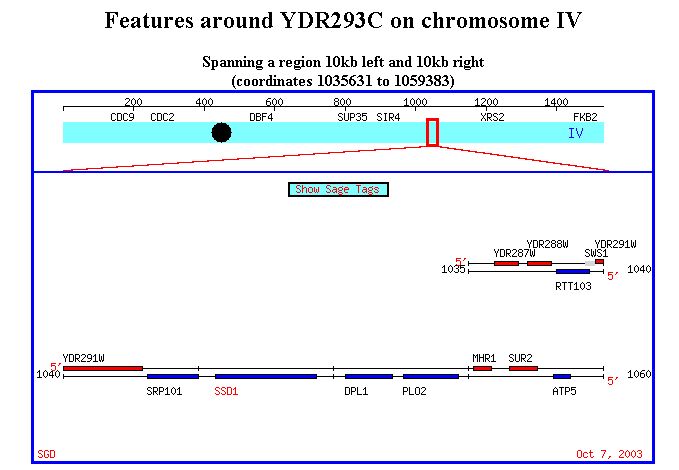
Figure 1. Physical map of SSD1 location, spanning from 1035631 to 1059383 on
chromosome IV.
SSD1 Nucleotide Sequence:
YDR293C Chr 4 reverse complement
ATGTCTAAAAATAGCAACGTTAACAACAATAGATCCCAAGAGCCAAATAACATGTTTGTG
CAAACCACAGGAGGTGGTAAAAACGCCCCAAAGCAGATTCATGTTGCACACAGACGTTCC
CAAAGTGAGTTGACAAATTTGATGATTGAACAATTCACTTTGCAGAAGCAGTTGGAGCAA
GTTCAAGCACAGCAGCAACAGTTGATGGCTCAGCAACAGCAATTGGCACAACAGACAGGA
CAATACCTGTCAGGAAATTCTGGCTCTAACAATCATTTCACGCCTCAACCGCCTCACCCT
CATTACAACTCAAACGGTAATTCACCTGGTATGAGTGCAGGTGGCAGCAGAAGTAGAACT
CACTCCAGGAACAACTCCGGATATTATCATAATTCATATGATAACAATAACAATAGCAAT
AATCCTGGGTCTAACTCACACAGAAAGACGAGTTCACAATCCAGCATATATGGCCATTCC
AGAAGACATTCTTTAGGTCTAAATGAAGCGAAAAAGGCTGCTGCGGAAGAACAAGCTAAA
AGAATATCTGGGGGTGAAGCAGGCGTAACTGTGAAGATAGATTCTGTTCAAGCTGATAGT
GGCTCAAATTCTACTACAGAACAATCTGATTTTAAATTTCCACCACCACCAAATGCTCAT
CAGGGCCATCGTCGCGCAACTTCAAACCTATCACCTCCCTCTTTCAAATTTCCTCCAAAC
TCTCACGGGGATAATGACGATGAATTCATAGCAACCTCTTCAACGCACCGCCGTTCAAAG
ACAAGAAACAATGAATATTCTCCAGGCATTAATTCCAACTGGAGAAACCAATCACAGCAA
CCTCAACAGCAGCTTTCTCCATTCCGCCACAGAGGATCTAATTCAAGGGATTACAATTCC
TTCAATACCTTAGAACCTCCTGCGATATTTCAGCAGGGACACAAACATCGTGCCTCTAAT
TCATCAGTTCATAGTTTCAGTTCACAAGGTAATAATAACGGAGGTGGACGTAAGTCCCTA
TTTGCACCCTACCTTCCCCAAGCCAACATTCCAGAGCTAATCCAAGAAGGGAGACTAGTA
GCTGGTATATTAAGAGTTAATAAAAAGAATAGATCGGATGCCTGGGTCTCTACAGATGGC
GCTCTTGATGCGGATATTTACATTTGCGGCTCCAAAGATCGTAATAGAGCACTTGAAGGT
GATTTAGTCGCGGTAGAACTATTAGTTGTGGACGATGTTTGGGAGTCCAAGAAAGAAAAG
GAAGAAAAGAAGAGGAGAAAGGATGCCTCTATGCAACACGATCTAATTCCTTTGAACAGT
AGTGACGATTACCACAACGATGCATCTGTTACTGCTGCAACAAGCAACAATTTTCTATCT
TCTCCCTCCTCGTCTGATTCGCTAAGCAAGGATGATTTATCCGTCAGAAGAAAGAGGTCA
TCTACTATCAATAATGATAGTGATTCCTTATCATCTCCTACCAAATCAGGAGTAAGGAGA
AGAAGTTCATTGAAACAACGTCCAACTCAAAAGAAAAATGACGATGTTGAAGTTGAAGGT
CAGTCATTGTTATTAGTTGAAGAAGAAGAAATCAACGATAAATATAAGCCACTTTACGCA
GGCCATGTCGTTGCTGTTTTGGACCGTATCCCTGGTCAGTTATTTAGCGGTACATTAGGT
TTGTTGAGACCATCCCAACAAGCTAATAGCGACAATAACAAACCACCACAAAGCCCAAAA
ATTGCTTGGTTCAAGCCTACTGATAAGAAGGTGCCATTAATTGCAATTCCTACAGAATTA
GCTCCAAAGGACTTTGTTGAAAACGCTGATAAATACTCCGAAAAGTTATTCGTTGCCTCT
ATTAAACGTTGGCCAATCACATCTTTGCATCCATTTGGTATTTTAGTTTCCGAACTTGGA
GATATTCACGATCCTGATACTGAAATTGATTCCATTTTAAGGGATAACAATTTTCTTTCG
AATGAATATTTGGATCAAAAAAATCCGCAAAAAGAAAAACCAAGTTTTCAGCCGCTACCA
TTAACGGCTGAAAGTCTAGAATATAGGAGGAATTTTACGGACACTAATGAGTACAATATC
TTTGCAATTTCCGAGCTTGGATGGGTGTCTGAATTTGCCTTACATGTCAGGAATAACGGA
AATGGTACCCTAGAGCTGGGTTGTCATGTTGTTGATGTGACCAGCCATATTGAAGAAGGC
TCCTCTGTTGATAGGCGTGCGAGAAAGAGGTCCTCTGCGGTGTTCATGCCACAAAAACTT
GTCAATTTATTACCACAATCGTTCAACGACGAACTGTCGTTGGCCCCTGGCAAGGAATCA
GCCACGCTGTCGGTTGTTTACACTCTAGACTCATCTACTTTAAGGATTAAATCTACTTGG
GTAGGCGAATCTACAATTTCCCCCTCAAACATCTTGTCTTTAGAACAATTAGACGAAAAA
TTATCTACTGGAAGTCCCACTAGCTACCTCTCTACTGTACAGGAAATTGCTAGATCATTT
TATGCTAGAAGAATAAATGATCCAGAAGCTACATTACTTCCCACCCTGTCCTTATTGGAA
AGCTTGGATGACGAAAAAGTTAAGGTTGACTTGAACATCCTGGATAGAACTTTAGGCTTT
GTTGTAATTAATGAGATTAAAAGAAAGGTCAACTCCACTGTTGCAGAGAAAATTTACACC
AAACTTGGTGATCTAGCTCTTTTGAGAAGGCAGATGCAACCCATTGCAACCAAGATGGCG
TCATTTAGAAAGAAAATTCAAAATTTTGGTTACAATTTTGATACCAATACGGCGGATGAA
TTAATCAAAGGGGTGCTAAAAATTAAAGATGACGATGTTAGAGTCGGAATTGAAATTTTA
CTGTTTAAAACCATGCCAAGAGCTAGATACTTTATTGCTGGCAAAGTAGACCCGGACCAA
TATGGGCATTATGCCTTGAACCTACCTATCTACACACATTTCACAGCGCCAATGAGAAGA
TACGCTGATCATGTCGTTCATAGGCAATTAAAGGCCGTTATCCACGATACTCCATACACC
GAAGATATGGAAGCTTTGAAGATTACCTCCGAATATTGTAATTTTAAAAAGGACTGTGCT
TATCAAGCACAGGAACAAGCAATTCATCTATTGTTGTGTAAAACAATCAACGACATGGGA
AATACTACAGGACAATTATTAACAATGGCTACTGTCTTACAAGTTTACGAGTCCTCCTTT
GATGTATTTATTCCAGAATTTGGTATTGAAAAGAGAGTTCATGGAGATCAACTACCTTTG
ATCAAAGCTGAGTTTGATGGTACCAATCGTGTCTTGGAATTGCATTGGCAGCCCGGCGTA
GATAGTGCAACTTTTATACCAGCAGATGAAAAAAATCCAAAATCCTATAGAAATTCCATT
AAGAACAAATTCAGATCCACAGCCGCTGAGATTGCGAATATTGAACTAGATAAAGAAGCG
GAATCTGAACCATTGATCAGCGATCCATTGAGTAAGGAACTCAGCGATTTGCATCTAACA
GTACCAAATTTAAGGCTACCATCTGCAAGCGACAACAAGCAAAATGCTTTAGAAAAATTC
ATTTCTACTACTGAAACCAGAATTGAAAATGATAACTATATACAAGAAATACATGAATTG
CAAAAGATTCCTATTCTATTGAGAGCTGAGGTGGGGATGGCTTTGCCATGTTTAACCGTC
CGTGCATTAAATCCATTCATGAAGAGGGTATAA
SSD1 Protein Sequence:
1 MSKNSNVNNN RSQEPNNMFV QTTGGGKNAP KQIHVAHRRS QSELTNLMIE
51 QFTLQKQLEQ VQAQQQQLMA QQQQLAQQTG QYLSGNSGSN NHFTPQPPHP
101 HYNSNGNSPG MSAGGSRSRT HSRNNSGYYH NSYDNNNNSN NPGSNSHRKT
151 SSQSSIYGHS RRHSLGLNEA KKAAAEEQAK RISGGEAGVT VKIDSVQADS
201 GSNSTTEQSD FKFPPPPNAH QGHRRATSNL SPPSFKFPPN SHGDNDDEFI
251 ATSSTHRRSK TRNNEYSPGI NSNWRNQSQQ PQQQLSPFRH RGSNSRDYNS
301 FNTLEPPAIF QQGHKHRASN SSVHSFSSQG NNNGGGRKSL FAPYLPQANI
351 PELIQEGRLV AGILRVNKKN RSDAWVSTDG ALDADIYICG SKDRNRALEG
401 DLVAVELLVV DDVWESKKEK EEKKRRKDAS MQHDLIPLNS SDDYHNDASV
451 TAATSNNFLS SPSSSDSLSK DDLSVRRKRS STINNDSDSL SSPTKSGVRR
501 RSSLKQRPTQ KKNDDVEVEG QSLLLVEEEE INDKYKPLYA GHVVAVLDRI
551 PGQLFSGTLG LLRPSQQANS DNNKPPQSPK IAWFKPTDKK VPLIAIPTEL
601 APKDFVENAD KYSEKLFVAS IKRWPITSLH PFGILVSELG DIHDPDTEID
651 SILRDNNFLS NEYLDQKNPQ KEKPSFQPLP LTAESLEYRR NFTDTNEYNI
701 FAISELGWVS EFALHVRNNG NGTLELGCHV VDVTSHIEEG SSVDRRARKR
751 SSAVFMPQKL VNLLPQSFND ELSLAPGKES ATLSVVYTLD SSTLRIKSTW
801 VGESTISPSN ILSLEQLDEK LSTGSPTSYL STVQEIARSF YARRINDPEA
851 TLLPTLSLLE SLDDEKVKVD LNILDRTLGF VVINEIKRKV NSTVAEKIYT
901 KLGDLALLRR QMQPIATKMA SFRKKIQNFG YNFDTNTADE LIKGVLKIKD
951 DDVRVGIEIL LFKTMPRARY FIAGKVDPDQ YGHYALNLPI YTHFTAPMRR
1001 YADHVVHRQL KAVIHDTPYT EDMEALKITS EYCNFKKDCA YQAQEQAIHL
1051 LLCKTINDMG NTTGQLLTMA TVLQVYESSF DVFIPEFGIE KRVHGDQLPL
1101 IKAEFDGTNR VLELHWQPGV DSATFIPADE KNPKSYRNSI KNKFRSTAAE
1151 IANIELDKEA ESEPLISDPL SKELSDLHLT VPNLRLPSAS DNKQNALEKF
1201 ISTTETRIEN DNYIQEIHEL QKIPILLRAE VGMALPCLTV RALNPFMKRV
PDB search yielded no results.
Sources
Chen CY, Rosamond J. 1998. Candida
albicans SSD1 can suppress multiple mutations in Saccharomyces cerevisiae. Microbiology.
1998 Nov;144 ( Pt 11):2941-50.
<http://mic.sgmjournals.org/cgi/reprint/144/11/2941.pdf>
Kaeberlein M, Guarente L. 2002.
Saccharomyces cerevisiae MPT5 and SSD1 function in parallel pathways to promote
cell wall integrity. Genetics. 2002 Jan;160(1):83-95.
<http://www.genetics.org/cgi/content/full/160/1/83>
MIPS Comprehensive Yeast Genome
Database. 2003.
<http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YDR293C>
SGD database. 2003. Stanford University.
<http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=YDR293C>
Sutton A, Immanuel D, Arndt KT.
1991. The SIT4 protein phosphatase functions in late G1 for progression into
S phase. Mol Cell Biol. 1991 Apr;11(4):2133-48. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=1848673&dopt=Abstract>
Swiss-Prot database. 2003.
<http://us.expasy.org/cgi-bin/niceprot.pl?P24276>
Uesono Y, Toh-e A, Kikuchi Y. 1997.
Ssd1p of Saccharomyces cerevisiae associates with RNA. J Biol Chem. 1997 Jun
27;272(26):161039.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=9195905&dopt=Abstract>
Wheeler R. T., Kupiec M, Magnelli
P, Abeijon C, Fink G. R. A Saccharomyces cerevisiae mutant with increased virulence.
2003. Proc Natl Acad Sci U S A. 2003 Mar 4;100(5):2766-70. Epub 2003 Feb 14.
<www.pnas.org/cgi/reprint/100/5/2766.pdf>
Tsuchiya E, Matsuzaki G, Kurano
K, Fukuchi T, Tsukao A, Miyakawa T. 1996. The Saccharomyces cerevisiae SSD1
gene is involved in the tolerance to high concentration of Ca2+ with the participation
of HST1/NRC1/BFR1. Gene. 1996 Oct 17;176(1-2):35-8. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=8918228&dopt=Abstract>
My Non-annotated Yeast Gene
YDR288W
SGD listed no known molecular function, biological process and cellular component
for YDR288W. SGD says that YDR288W is an uncharacterized ORF, which means it
is “likely to be real due to the existence of orthologs in one or more other
species, but which are not supported with specific experimental data demonstrating
that a gene product is produced in S. cerevisiae” (SGD, 2003).
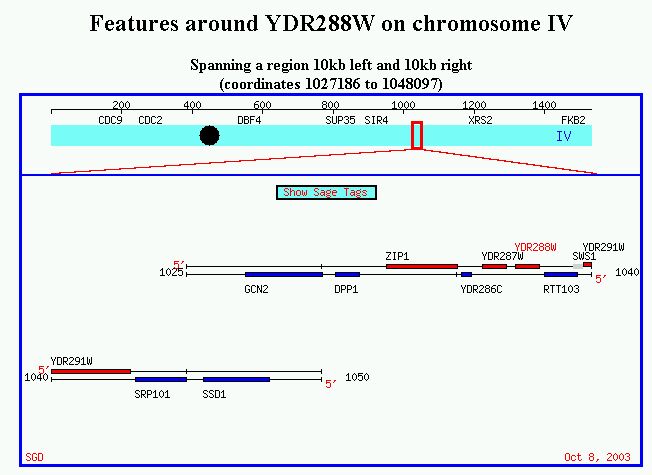
Figure 2. Location of YDR288W on chromosome IV, spanning from 1027186 to 1048097.
My annotated gene, SSD1, is visible on this figure as well.
NCBI protein database had the protein sequence information listed, and I used
this sequence information for a Kyte-Doolittle analysis.
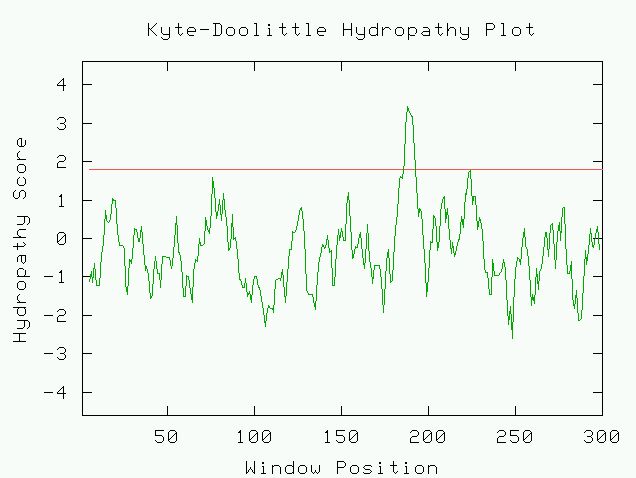
Figure 3. Kyte-Doolittle hydropathy plot of YDR288W. A high peak at amino acid
position about 190 shows that this protein is probably an integral membrane
protein.
NCBI protein database also returned a result for Ydr288wp, a protein required
for cell viability (NCBI, 2003). Blastn did not return any significant results;
however, Blastp returned information about Ydr288wp and listed YDR288W as a
probable membrane protein. See figures 4 and 5 below.
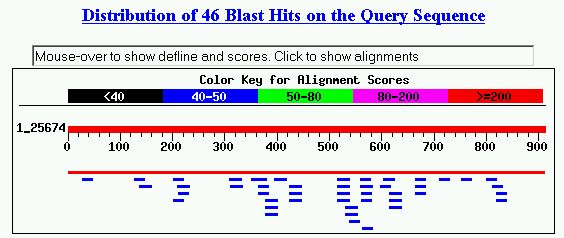


Figure 4. Blastn results with YDR288W nucleotide sequence. The first hit that
yielded E value of zero was Saccharomyces cerevisiae chromosome IV cosmid 9819,
which included many different genes, and YDR288W was one of them. The rest of
the hits are not very impressive, because the similarity regions are very fragmented,
and the E values are not significantly low. Also, most of the hits were not
characterized genes. This Blastn search did not help me predict the cellular
role of YDR288W.
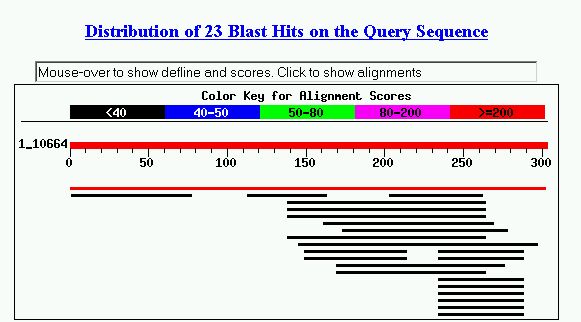

Figure 5. Blastp results with YDR288W protein sequence. The first hit listed
is Ydr288wp, which appeared in NCBI protein search earlier. The second hit is
for an unknown bacterial protein that is not characterized yet. Also, I see
many hits with Melano antigen, MAGE protein family in humans. The other hits
yielded fairly low bit scores and high E values, so they did not help me predict
the cellular role of YDR288W.
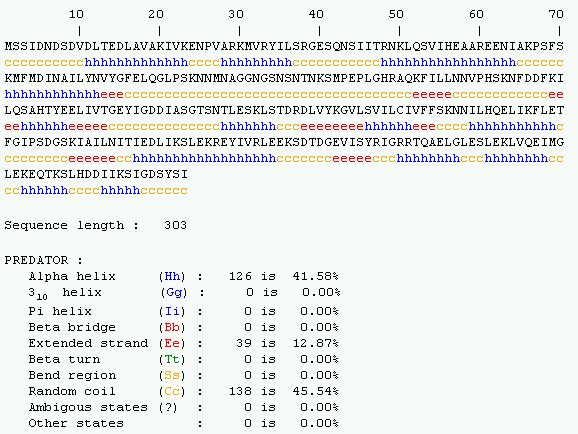
Figure 6. Predicting the secondary structure of YDR288W protein using PREDATOR.
This sequence is mostly composed of alpha helix (41.58%) and random coiling
(45.54%).
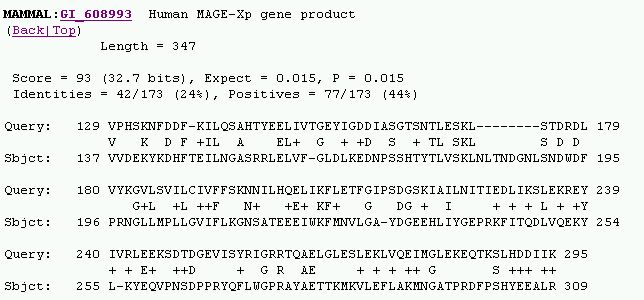
Figure 7. Mammalian Homology to Yeast analysis from SGD. The closest hit was
for Human MAGE-Xp gene product with 24% identity and 44% similarity. It is interesting
to note that homology analysis from SGD parallels the MAGE protein findings
from Blastp results.
Following the “Entrez Neighbors” link from SGD site on YDR288W, I found more
links about the human MAGE family hits, so I performed a Blast2 query, using
MAGE B-1 and YDR288W protein sequences.
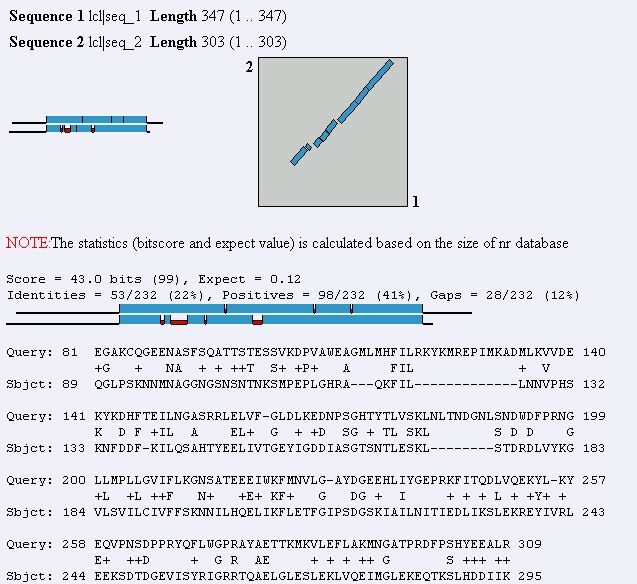
Figure 8. Blast2 result using MAGE B-1 and YDR288W protein sequences. The two
sequences show a fair similarity to each other.
So, I investigated further into MAGE proteins, and found out that MAGEB family
is expressed in a significant fraction of tumors of various histological types
(Lurquin, 1997).
The MAGE (melanoma antigen-encoding
gene) family are expressed in a wide variety of tumors but not in normal cells,
with the exception of the male germ cells, placenta, and, possibly, cells of
the developing embryo (NCBI, 2003 Melanoma antigen, family B, 1 [Homo sapiens]).
I found the fact that YDR288W had some similarity to MAGE proteins notable,
because MAGE proteins have been identified in tumors and male germ cells, which
may indicate that YDR288W may be associated with cell growth or cell growth
regulation.
Conserved Domain search returned no hits.
YDR288W Nucleotide Sequence
>YDR288W Chr4
ATGAGTTCTATAGATAATGACAGCGATGTGGATTTAACAGAAGATTTAGCCGTGGCCAAG
ATCGTTAAAGAGAATCCTGTAGCTAGAAAAATGGTTAGGTATATTTTATCGAGAGGAGAA
TCTCAAAATTCGATCATAACAAGAAACAAGTTACAATCTGTTATACATGAAGCTGCTCGC
GAGGAAAACATAGCAAAACCGTCTTTTAGTAAAATGTTTATGGACATAAATGCTATACTG
TATAACGTATATGGGTTTGAATTACAAGGATTGCCGTCAAAAAATAATATGAACGCTGGC
GGTAATGGAAGCAATAGCAATACAAATAAATCAATGCCTGAGCCACTGGGACATAGAGCC
CAGAAATTTATCTTGCTAAACAATGTCCCGCACTCAAAAAATTTTGACGATTTCAAGATT
TTACAAAGCGCTCACACTTATGAAGAGCTAATAGTTACCGGAGAGTATATTGGTGACGAT
ATAGCTTCAGGAACTTCCAATACCTTAGAAAGCAAGTTGAGTACAGATCGAGATCTTGTT
TACAAGGGTGTTTTGAGCGTCATTTTATGCATTGTTTTCTTCTCTAAGAATAACATATTA
CATCAGGAATTGATCAAATTTCTAGAGACATTTGGCATTCCAAGCGATGGTTCAAAGATT
GCTATACTGAACATTACTATTGAAGATTTGATAAAAAGTTTAGAAAAGCGCGAATATATA
GTCAGATTGGAGGAGAAATCCGATACCGACGGGGAAGTGATATCATACAGGATAGGTAGA
AGAACTCAGGCTGAGCTTGGACTAGAGTCTCTTGAAAAATTGGTGCAAGAAATCATGGGC
CTTGAAAAGGAGCAGACTAAAAGTTTGCACGATGATATAATAAAAAGCATTGGCGATTCA
TATTCTATATAG
Protein sequence of YDR288W
1 mssidndsdv dltedlavak
ivkenpvark mvryilsrge sqnsiitrnk lqsviheaar
61 eeniakpsfs kmfmdinail
ynvygfelqg lpsknnmnag gngsnsntnk smpeplghra
121 qkfillnnvp hsknfddfki
lqsahtyeel ivtgeyigdd iasgtsntle sklstdrdlv
181 ykgvlsvilc ivffsknnil
hqelikflet fgipsdgski ailnitiedl ikslekreyi
241 vrleeksdtd gevisyrigr
rtqaelgles leklvqeimg lekeqtkslh ddiiksigds
301 ysi
Bottom line
I have found that YDR288W protein
is probably a membrane protein, and is required for cell viability. Also, YDR288W
protein showed similarity to the human MAGE (melanoma antigen-encoding gene)
family, which may suggest that YDR288W is associated with cell growth or cell
growth regulation. Given that this protein is required for cell viability, I
assume this protein would play a critical role in cell cycle or growth regulation.
Since this is a membrane bound protein, it could be a receptor in a cell growth
signal pathway, that might be post-transcriptionally modified, such as being
phosphorylated. Another hypothesis I can propose is that YDR288W protein might
be a membrane bound transport, that may play a critical role in cell growth.
Sources
Lurquin C, De Smet C, Brasseur F,
Muscatelli F, Martelange V, De Plaen E, Brasseur R, Monaco AP, Boon T. 1997.
Two members of the human MAGEB gene family located in Xp21.3 are expressed in
tumors of various histological origins. Genomics. 1997 Dec 15;46(3):397-408.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=PubMed&cmd=Retrieve&list_uids=9441743&dopt=Citation>
NCBI Protein database. 2003. YDR288W.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=protein&list_uids=6320494&dopt=GenPept&term=YDR288W&qty=1>
NCBI Protein database. 2003. Melanoma
antigen, family B, 1 [Homo sapiens].
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=Protein&list_uids=15489351&dopt=GenPept>
NCBI Genbank database. 2003. MAGEB1.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=nucleotide&dopt=GenBank&list_uids=29171710>
Predator. 2003.
<http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_preda.html>
SGD database. 2003. Stanford University.
<http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=YDR288W>









