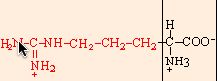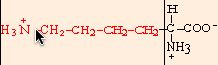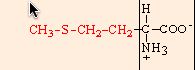This web page was produced as an assignment for an undergraduate course at Davidson College
Annotated Yeast Gene CYT1(YOR065W) and Non-Annotated Yeast GeneYOR066W
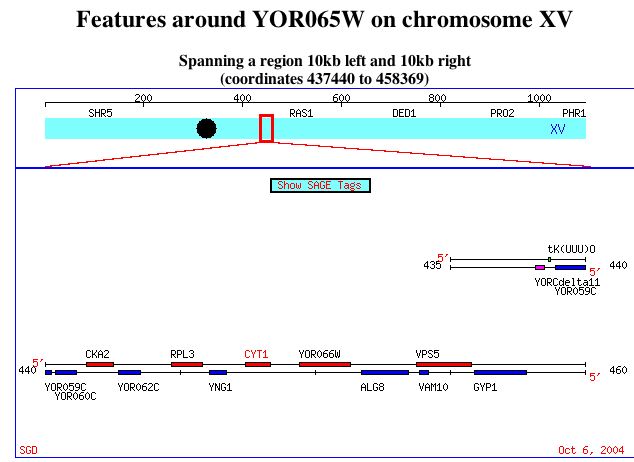
image from: http://db.yeastgenome.org/cgi-bin/ORFMAP/ORFmap?locus=YOR065W
The figure shows the relative locations of annotated gene CYT1 (also called YOR065W) and non-annotated gene YOR066W, on Saccharomyces cerevisiae chromosome 15.
Annotated CYT1
Molecular Function:
CYT 1 codes for an electron transporter, which transfers electrons within CoQH2-cytochrome c reductase complex. To test the necessity of CYT1 an experiment, using dominant-negative mutations of the CYT1 gene, was conducted. A single arginine at amino acid 166 substituted by glycine, or substituted by glycine and another amino acid, caused a nearly nonfunctional cytochrome c1.
arginine |
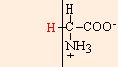
glycine |
lycine
|
methionine |
leucine |
chemical structures from: http://www.web-books.com/MoBio/Free/Ch2A2.htm
R166K, R166M, and R166L resulted in normal, or nearly normal, function. Arg-166 is conserved in all cytochromes c1, but Cytp does not appear to interact directly with any of the physiological partners of the cytochrome bc1 complex. It appears that the large size of the side chain (shown above, in red) at position 166 is critical for a functional cytochrome c1, but that R166 is not imperative for cytochrome c1 interaction with the cytochrome bc 1 complex (Ahmad, 2001).
Biological Process:
Cytp 1 (cytochrome c) is one of ten protein subunits that forms a monomer, with which the cytochrome bc1 complex (QCR) dimerizes (Ahmad, 2001). Cytochrome bc1, an integral membrane protein of the respiratory chain, catalyzes the electron transfer from ubiquinol to cytochrome c (Lang, 2001).
 |
Figure. Saccharomyces cerevisiae binding of phospholipids (L1-L5) and detergent (UM) shown in yellow, seen in the transmembrane region of QCR, is viewed from the matrix side on the membrane plane. The core of the dimeric complex is made up of eight helices of COB (red) per monomer. Single helices of RIP1 (green), CYT1 (yellow), QCR8 (purple) and QCR9 (gray) are attached on each side at the periphery. The heme b cofactors (light gray) as well as UQ6 and Stig (dark gray) are shown. The helices are depicted as ribbon drawings, and further molecules are depicted as ball-and-stick representation. Standard colors are used for all non-carbon atoms. |
image and caption from: http://embojournal.npgjournals.com/cgi/content/full/20/23/6591
Cellular Function:
CYT1 is a yeast gene that codes for cytochrome c1 in Saccharomyces cerevisiae. It is involved in the mitochondrial electron transport chain and, structurally, in the mitochondrial inner membrane.
To see whether this protein is conserved, I used the amino acid sequence in a blastp search: http://www.ncbi.nlm.nih.gov:80/BLAST/Blast.cgi There were many hits with high bit scores and low E values, indicating that the amino acid sequence is conserved. The most closely aligned hits were those of yeast proteins, but mammalian proteins were also among the higher bit scores. Cytochrome c1 is seen in Homo sapiens with a bit score of 298 and an E-value of 1e - 79. Similarity between a yeast and human protein, indicates that cytochrome c1 is highly conserved.
Because sequence similarity is more likely to be found in amino acid sequences, than it is in nucleotide sequences, resulting from "third base wobble," I used the CYT1 nucleotide sequence in a a blastn search: http://www.ncbi.nlm.nih.gov:80/BLAST/Blast.cgi As suspected, there were fewer hits with high bit scores and low E values for the nucleotide comparison. Conservation between the yeast and mammalian cytochrome c nucleotide sequence, bit score 48 and E value .037, was obtained from a human clone of the Saccharomyces cerevisiae gene on chromosome 15.
Non-annotated YOR066W
YOR066W is located near CYT1 on chromosome 15 of Saccharomyces cerevisiae. The nucleotide and protein sequences of this Open Reading Frame (ORF) are indicated below.
Nucleotide Sequence:
atggataaaa gcatgatcaa aaaaagaggc aggcctccca tcaccaaaga ttatcccaat
61 cccctacaaa gccccatggc gcattcttca atgcaggttc agaagcaggg gcctcacagt
121 tttgccaaac ctttgatgaa agtggggcaa tctagtccat ccccgaataa aagaagactc
181 agcatcgatc atcatcataa tttggcggca actacgagaa agggcagata caggggcgtt
241 cttctttcta ctcccaccaa gaaatcgagc actaatggct ctacgccgat ctctaccccg
301 tcttcgaatg acagctataa taacacggtg ttttctgaaa caaggaaaac ctttcttcag
361 agttcgccgc ctatcatgac gtcttctcca gctttccaaa agaaaaatga ctacatgttt
421 ccttcacagg aacaattcaa actttcttta accattaccg aaagtgggaa ggccgttatt
481 gcaggatctt tgcctttctc tccatcgtca aaatcgtctc atttaatgaa caataataat
541 aaaaagatca tgcaaaatga aaaaattcac aaaggaagca agaaaaatgc acccaagttt
601 gagaagagac gtattctttc tctcttaaaa cagatgaaga atgagaaata ttgtgatact
661 gatacgctcc ccgaagcacc tccagcgaaa ccaagcagat ctgacatcat cgacacagaa
721 ttgccgacta tcatagaaac ttcagcttct ccaataggca gtgctagaaa caataatata
781 ctgctttcgc aaccaccaca atcaccccct tcaagcgcgc aacttaagcc cccatccact
841 ccaaagagtt ctctacaatt caggatggga tttactccca atgttgcact aaattccgta
901 tcattaagtg atactatatc caaatctaca aatgctgtgg gtgccagcaa taataacaat
961 cagaacggta attccattag caatatcgcg gatgctaata ctttacttac cttgactaat
1021 agcccgggag ttttcttatc acctcgaaac aaaatgttgc ccaagtcgac tactgcatct
1081 aacgaacaac agcaggaatt cgtgtttaag ttttcaagtg gtgacccgtt attgcttact
1141 gacgacgcag atgggaattg gccagaaatg ctgttcaatg tatctaatac gcccagacgt
1201 caaaaatgtt ttaacacacc tccttcatgg ataaactttg gttcacctgg attattcagc
1261 ccaccaagaa gctctaatgt catggtaaat ggcaccactg tagccaccgc atcggatagc
1321 ggcaacgtcc atagacaatt acaagctcaa ttggaggcgc aagtgcaagt gcaatcgcaa
1381 tccaattctc ctactcaaag gcaacaacaa cagcgtcaat ttcaaatacc cccacctcac
1441 ataaacatga actcttcccc gccacaaatc aatatcgctt ccccacctca tcaatctatg
1501 tctcgggtgt catctattta cttcaacaaa gaaaaaacca caacaggggt agcaaatatg
1561 ctaggtaaca ccaaaagcga aaacttacag ccgccggcga atttatttac cgctgcccat
1621 ggaccttcaa ctcctagaaa tcaagaattt caattaccca ctttgattga atgcactcca
1681 ttaatccagc aaaccatgaa tggctcatta ggaaccaaat acataccagg tacttcaatt
1741 tcgaacagtg caactcctaa tttacacggg ttccctgttg gaaccgggaa agctccctct
1801 agctttgacg attcactaaa acaaaacccg tacagcaaca aacaggatga tgctagaact
1861 gccttgaaaa gattaatcga tgatcagtga
Amino Acid Sequence:
1 mdksmikkrg rppitkdypn plqspmahss mqvqkqgphs fakplmkvgq sspspnkrrl
61 sidhhhnlaa ttrkgryrgv llstptkkss tngstpistp ssndsynntv fsetrktflq
121 ssppimtssp afqkkndymf psqeqfklsl titesgkavi agslpfspss ksshlmnnnn
181 kkimqnekih kgskknapkf ekrrilsllk qmknekycdt dtlpeappak psrsdiidte
241 lptiietsas pigsarnnni llsqppqspp ssaqlkppst pksslqfrmg ftpnvalnsv
301 slsdtiskst navgasnnnn qngnsisnia dantlltltn spgvflsprn kmlpksttas
361 neqqqefvfk fssgdplllt ddadgnwpem lfnvsntprr qkcfntppsw infgspglfs
421 pprssnvmvn gttvatasds gnvhrqlqaq leaqvqvqsq snsptqrqqq qrqfqippph
481 inmnssppqi niaspphqsm srvssiyfnk ektttgvanm lgntksenlq ppanlftaah
541 gpstprnqef qlptliectp liqqtmngsl gtkyipgtsi snsatpnlhg fpvgtgkaps
601 sfddslkqnp ysnkqddart alkrliddq
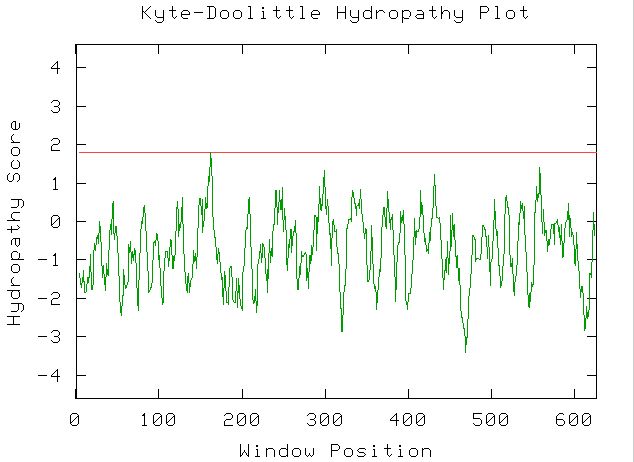
Figure. window for the Kyte-Doolittle hydropathy plot is set at window size 9. No portions of the hydropathy plot for yeast gene YOR066W appear above cut off 1.8. This plot indicates that YOR066W is probably not a transmembrane protein, nor is it likely that it contains regions of globular proteins.
Blast searches did not detect conservation of YOR066W sequences. http://www.ncbi.nlm.nih.gov:80/BLAST/Blast.cgi.
Although this non-annotated gene has "unknown molecular function, biological process, and cellular component," the nucleotide sequence indicates that this gene potentially codes for Cdc28p substrate.
Cdc28, in Saccharomyces cerevisiae is a catalytic subunit of the main cell cycle cyclin-dependent kinase (CDK). This protein associates with G1 cyclins and G2/M cyclins, directing CDK to specific substrates during cellular processes. The amino acid sequence for Cdc28 is as follows:
msgelanykr lekvgegtyg vvykaldlrp gqgqrvvalk kirlesedeg vpstaireis
61 llkelkddni vrlydivhsd ahklylvfef ldldlkryme gipkdqplga divkkfmmql
121 ckgiaychsh rilhrdlkpq nllinkdgnl klgdfglara fgvplrayth eivtlwyrap
181 evllggkqys tgvdtwsigc ifaemcnrkp ifsgdseidq ifkifrvlgt pneaiwpdiv
241 ylpdfkpsfp qwrrkdlsqv vpsldprgid lldkllaydp inrisarraa ihpyfqes
The blastp query for Saccharomyces cerevisiae Cdc28 shows a great degree of conservation. http://www.ncbi.nlm.nih.gov:80/BLAST/Blast.cgi
The blastn nucleotide sequence for Saccharomyces cereviisiae (below) contained some sequence alignments. http://www.ncbi.nlm.nih.gov:80/BLAST/Blast.cgi
atgagcggtg aattagcaaa ttacaaaaga cttgagaaag tcggtgaagg tacatacggt
61 gttgtttata aagcgttaga cttaagacct ggccaaggtc aaagagtagt cgcattgaag
121 aaaataagac tagagagtga agacgagggt gttcccagta cagccatcag agaaatctca
181 ttattgaagg aattaaaaga cgataatatt gtcagattat acgatattgt tcactctgat
241 gcacacaagc tatatcttgt ttttgagttc ctcgatttgg acctgaaaag atatatggag
301 ggtattccaa aggaccaacc gttaggagct gatattgtta agaagtttat gatgcaactt
361 tgtaagggta ttgcatactg ccactcacac cgtattctgc atcgtgattt aaaaccgcag
421 aacttattga ttaacaaaga tgggaatcta aaactaggtg attttggctt agcgcgtgct
481 tttggtgttc cgttgagagc ttacacacat gaaattgtta ctctatggta tagagctccg
541 gaggtattac tgggtggaaa acaatatagt acaggtgtcg atacatggtc catcggctgt
601 atatttgccg aaatgtgtaa caggaaacca atcttcagtg gcgatagtga gatcgatcag
661 attttcaaga tattcagagt attgggaacg ccgaatgaag ctatatggcc agatattgtc
721 tacttgcctg atttcaagcc aagctttcct caatggcgca gaaaagacct atcacaagtg
781 gtaccaagtc tagatccacg cggtattgat ttgttggaca aactcctcgc gtatgaccct
841 attaaccgga ttagcgccag aagagcagcc atccacccct acttccaaga atcataa
Conclusions
The data indicate that non-annotated YOR066W might code for a substrate of the catalytic subunit Cdc28. The gene YOR066W is located on chromosome 15 of the Saccharomyces cerevisiae genome, near CYT1. CTY1 is expressed on the inner mitochondrial membrane, allowing protein interaction in the mitochondrial electron transport chain. If YOR066W is involved in the same cellular processes as CYT1, then it follows that YOR066W would also be located in, or near the mitochondria.
Since data show that YOR066W may be associated with the CDC28 gene in Saccharomyces cerevisiae. We must consider the function of CDC28. CDC28 is the master regulator of cell division in the budding yeast (Mendenhall et al, 1998). CDC28 encodes cyclin-dependent protein kinase (CDK), which plays a role in DNA replication, spindle formation and chromosome separation (Mendenhall et al, 1998). If YOR066W is a CDC28 substrate, and we suspect that it contains no transmembrane proteins, than YOR066W is likely expressed, in the cytoplasm and/or the nucleus, near CDC28.
The large degree of conservation in CDC28 and the fact that domain conservation was essentially lacking in YOR066W might be explained by substrate, rather than ligand, specificity.
CDC28 is involved in several imperative biological functions. A protein that is involved in processes such as DNA replication, spindle formation and chromosome separation is probably highly conserved throughout species. At this point, the biological process and molecular function of YOR066W are unknown. A receptor, that does not require a transmembrane domain, of a ligand protein, such as CDC28, is a reasonable possibility for this protein.
References
Ahmad, Z., and Sherman, F. 2001. Role of Arg-166 in yeast cytochrome C1. J Biol Chem 276(21):18450-6.
Alberman, D.B., et al. 1997. The nucleotide sequence of Saccharomyces cerevisiae chromosome XV. Nature 387(6632 Suppl):98-102.
Lang, C., et al. 2001. Specific roles of protein–phospholipid interactions in the yeast cytochrome bc1 complex structure. The EMBO Journal. 20(23): 6591-6600.
Mendenhall, M., Hodgel, A. 1998. Regulation of Cdc28 Cyclin-Dependent Protein Kinase Activity during the Cell Cycle of the Yeast Saccharomyces cerevisiae. Microbiol Mol Biol Rev. 62 (4): 1191–1243. <http://www.pubmedcentral.gov/articlerender.fcgi?tool=pmcentrez&artid=98944> Accessed October 6, 2004
Saccharomyces Genome Database <http://www.geneontology.org> Accessed October 5, 2004.
Yeast Deletion Project and Proteomics of Mitochondria Database<http://www-deletion.stanford.edu/YDPM/YDPM_index.html> Accessed October 6, 2004.
Kyte Doolittle Hydropathy Plot <http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/activities/kd/slrkd.pl> Acccessed October 6, 2004.
Emily Wilson's Genomics Home Page
Email Questions, Comments or Suggestions : emwilson@davidson.edu