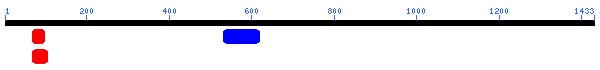CAT8 & YMR279C: My Favorite Yeast Genes |
If dogs hadn't gotten it first, Saccharomyces cervisiae would probably have been bequeathed the title of "Man's Best Friend." Although they don't fetch or bark or radiate a sense of everlasting companionship, the unicellular fungi can be counted on to metabolize sugar by fermentation. Scientists discovered that this process generates free ethanol and carbon dioxide around dawn of agricultural civilization, and humankind has been domesticating yeasts like S. cervisiae for the production of bread and beer ever since.1
Science has changed a lot over the last 3,500 years. Beer and bread have become staples of our diet, more often found in refrigerators than on the cutting edge, and yeast itself has become something of a staple for our science. S. cervisiae is one of the best-characterized organisms for studying genetics, and in 1996 became the first eukaryote to have its complete genome sequenced.2
 |
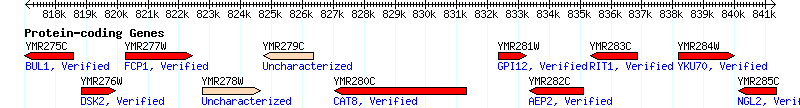
But sequence is only the first step to understanding. The S. cervisiae genome consists of 12,156,590 base-pairs divided amongst 16 chromosomes, encoding a predicted 6,591 ORFs.3 While functional information has been determined for a majority of these, roughly one-third of the ORFs remain totally unannotated. To explore what the differences in data are between annotated and unannotated genes, I present as much information as I could find, excluding microarray & protein interaction data sources or resorting to actual experimentation, on two genes that are located directly next to each other on chromosome 13.
Overview |

|
Gene |
CAT8 (YMR280C) |
|||||||||||||||||||||||||||||||||||||||||||||||||
Molecular Function |
-"Specific RNA polymerase II transcription factor activity" | Molecular Function unknown | ||||||||||||||||||||||||||||||||||||||||||||||||
Biological Process |
-"Positive regulation of gluconeogenesis" |
Biological Process unknown | ||||||||||||||||||||||||||||||||||||||||||||||||
Cellular Component |
-"Nucleus" | Cellular Component unknown | ||||||||||||||||||||||||||||||||||||||||||||||||
Protein Information |
|
|
||||||||||||||||||||||||||||||||||||||||||||||||
| Mutant Phenotype | "Null mutant is viable but unable to grow on non-fermentable carbon sources due to failure to derepress all major gluconeogenic enzymes; overexpression of Cat8p suppress inability of snf1 and snf4 mutants to grow on ethanol." | "viable" | ||||||||||||||||||||||||||||||||||||||||||||||||
Sequence Information |
Click here for sequence data | |||||||||||||||||||||||||||||||||||||||||||||||||
CAT8 |
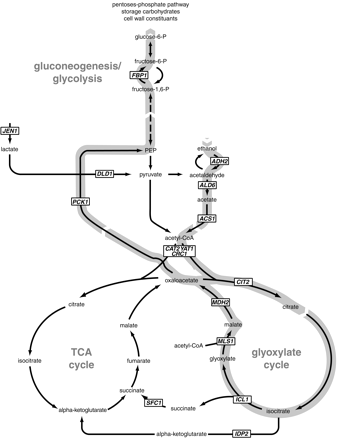 |
Yeast prefers to metabolize glucose by fermentation, but if environmental sugars run out, it can transform to a nonfermentative metabolism and use the ethanol (or other C2 or C3 energy substrate) it may have previously produced as an energy source.4 This change is called the diauxic shift, and as one might expect for a process that drastically alters metabolic input, changes the expression of a wide variety of genes. CAT8 encodes a transcriptional activator zinc-cluster protein designated Cat8p which derepresses at least 34 other genes that are highly induced during the initial stages of the diauxic shift.5 Almost all 34 of these genes have the CSRE (carbon source-responsive element) motif somewhere in their promoter region, which is the suspected site by which Cat8p regulates their expression.5
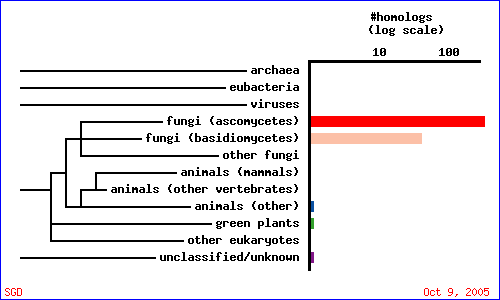
To guage how CAT8 is conserved among different species, a PSI-BLAST search was conducted with the gene's amino acid sequence against the UniRef database. Figure 5 and table 2 summarize the results, and indicate that CAT8 is conserved only in fungi.
| Acession number | Organism | Description | E-value | % aligned |
|---|---|---|---|---|
| Q6FJW6 | Candida glabrata | Candida glabrata strain CBS138 chromosome M complete sequence | 1.0e-261 | 99.8 |
| P39113 | Saccharomyces cerevisiae | Regulatory protein CAT8 | 1.0e-261 | 100 |
| O74229 | Kluyveromyces lactis | Cat8p | 1.0e-174 | 98.7 |
| Q75DZ4 | Eremothecium gossypii | ABL121Cp | 1.0e-160 | 99.9 |
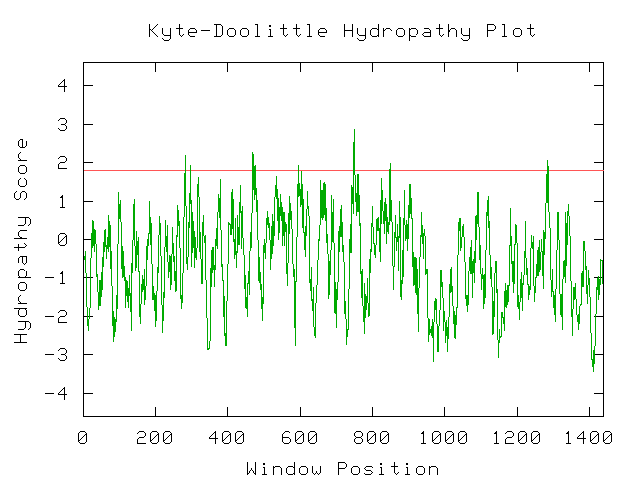
Unfortunately, there are no solutions to CAT8 in the protein database, and the closest homolog (ID66) only has 51 amino acids in common(4.1%) with an e-value of 1.7e-05. An NCBI6 search for conserved domains (fig 6.) only produced three hits, two of which were found in GAL4-like transcription regulators that had a zinc-finger, and one of which was simply called "Fungal specific transcription factor." A Kyte-Doolittle hydropathy plot (fig. 7) revealed several regions with hydropathicity scores greater than 1.8, but none were of realistic extent for a true transmembrane protein. Besides, CAT8 has been well characterized as a transcriptional controller, so perhaps the peaks have to do with certain regions that interact with DNA.
|
|
PREDATOR : Alpha helix (Hh) : 364 is 25.40% 310 helix (Gg) : 0 is 0.00% Pi helix (Ii) : 0 is 0.00% Beta bridge (Bb) : 0 is 0.00% Extended strand (Ee) : 115 is 8.03% Beta turn (Tt) : 0 is 0.00% Bend region (Ss) : 0 is 0.00% Random coil (Cc) : 954 is 66.57% Ambigous states (?) : 0 is 0.00% Other states : 0 is 0.00% |
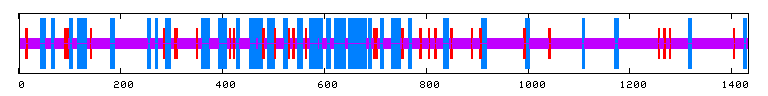 |
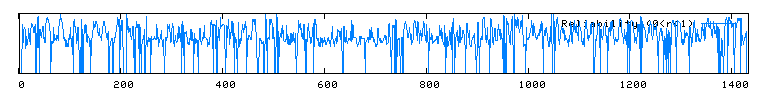 |
YMR279C |
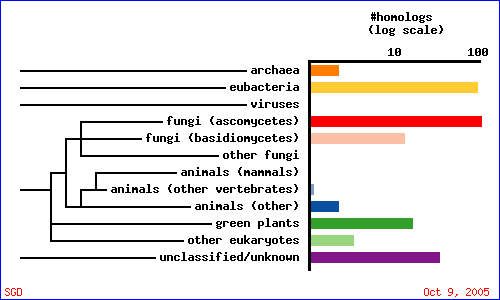
As indicated in the overview section, there currently is no accepted gene ontology information for YMR279C. What can we find out using the awesome array of public databases online? Let's start with a PSI-BLAST query (Fig. 9 & Table 3). Interestingly, YMR279C seems to be much more conserved than CAT8. Perhaps this is because CAT8 is nearly 3 times as large as YMR279C and hence, from a purely statistical perspective, less likely to retain its integrity over time. On the other hand, CAT8 may just be a very specific regulatory gene for yeast, and YMR279C of more general function. The BLAST results indicate YMR279C may be potentially have a role as a transporter of some kind, possibly a drug resistance transporter, in which case it would probably have a number of transmembrane domains. Let's investigate the domains conserved in our potential transporter, and then judge its potential to be embedded in a phosopholipid bilayer by calculating a Kyte-Doolittle plot.
|
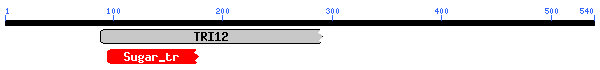
| 1. pfam06609, TRI12, Fungal trichothecene efflux pump (TRI12). Only 36.0% aligned, e-value=0.001 2. pfam00083, Sugar_tr, Sugar (and other) transporter.. Only 18.8% aligned, e-value=0.008. |
|
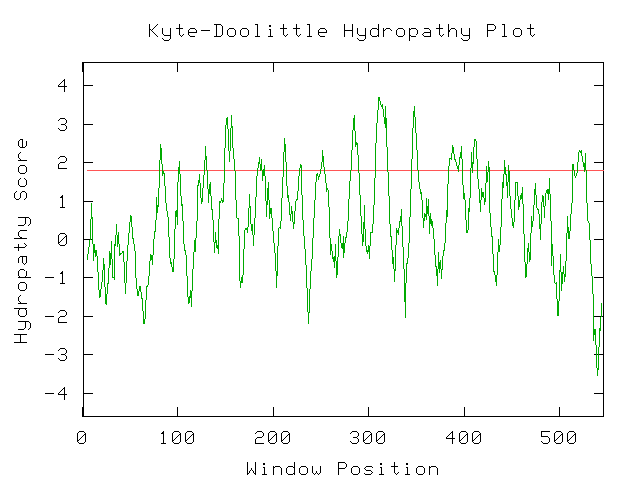
The conserved domain search (fig. 10) presents interesting results, correlating YMR0279C with two other transport proteins, one apparently involved in pumping toxins. However, neither have enough sequence in common with YMR0279C to generate significant e-values. The Kyte-Doolittle hydropathy plot (fig. 11) indicates the protein is predicted to have a substantial number (at least 7) of transmembrane domains, which supports the hypothesis that YMR0279 is a transmembrane protein.
PREDATOR : Alpha helix (Hh) : 138 is 25.56% 310 helix (Gg) : 0 is 0.00% Pi helix (Ii) : 0 is 0.00% Beta bridge (Bb) : 0 is 0.00% Extended strand (Ee) : 145 is 26.85% Beta turn (Tt) : 0 is 0.00% Bend region (Ss) : 0 is 0.00% Random coil (Cc) : 257 is 47.59% Ambigous states (?) : 0 is 0.00% Other states : 0 is 0.00 |
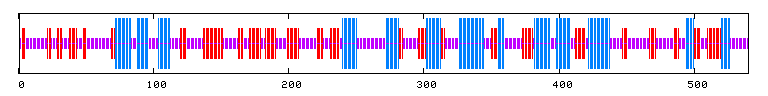 |
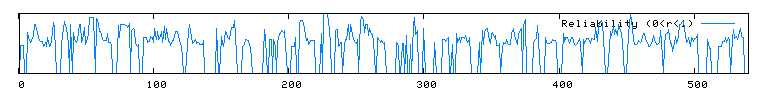 |
Conclusion |
Almost no direct data is available for the non-verified yeast protein YMR279C, and nothing can be certain where expirimental evidence is lacking. Nonetheless, by searching for other, known genes that encode similar amino acid sequences, it was possible to generate some preliminary data. It appears as if YMR279C is a transmembrane protein, possibly involved in the transport of a moderately complex molecule. The evidence for this comes from its conserved domains, which were also present in a protein that was involved in transporting sugar, and another that was involved in transporting a toxin.
CAT8, on the other hand, has been rather well studied, and besides its actual structure, of which there is no definitive information, its function is understood rather well. While not the direct initiator of the diauxic shift, CAT8 becomes active rather early as an essential translation regulator and is responsible for derepressing at least 34 genes as it participates in the expression cascade that ultimately reverses yeast's metabolism.
References |
1Wikipedia. 2005 3 Oct. Fermentation#History.<http://en.wikipedia.org/wiki/Fermentation#History>. Accessed 2005 8 Oct.
2NCBI. 2003. National Center for Biotechnology Information. <http://www.ncbi.nih.gov/>. Accessed 2005 8 Oct.
3Saccharomyces Genome Database. 2005. <http://www.yeastgenome.org/cache/genomeSnapshot.html>. Accessed 2005 8 Oct.
4Randez-Gil, F., Bojunga, N., Proft, M., & Entian, K-D. 1997. Glucose Derepression of Gluconeogenic Enzymes in Saccharomyces cerevisiae Correlates with Phosphorylation of the Gene Activator Cat8p. Molecular and Cellular Biology 17(5):2502-2510. Freetext .pdf available at http://mcb.asm.org/cgi/reprint/17/5/2502?view=reprint&pmid=9111319. PMID:9111319. Accessed 10/8/2005.
5Haurie, V. et al. 2001.The Transcriptional Activator Cat8p Provides a Major Contribution to the Reprogramming of Carbon Metabolism during the Diauxic Shift in Saccharomyces cerevisiae. The Journal of Biological Chemistry 276(1):76-85. Freetext .pdf available at http://www.jbc.org/cgi/content/full/276/1/76. PMID:11024040. Accessed 10/8/2005.
6Marchler-Bauer A, Bryant SH (2004), "CD-Search: protein domain annotations on the fly.", Nucleic Acids Res. 32:W327-331.
Kyte-Doolite Hydropathy Plot. 2003. <http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/activities/kd/kyte-doolittle.htm>. Accessed 2005 8 Oct.
[PREDATOR] <http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_preda.html>. Accessed 2005 8 Oct.
© Copyright 2005 Department of Biology, Davidson College, Davidson, NC 28035
Send comments, questions, and suggestions to: macowell "at" davidson.edu