Physical and in silico Approaches Identify DNA-PK in a Tax DNA-damage Response Interactome
Summary/Figure Explanations
It has been determined that Adult T-cell Leukemia (ATL) is caused by the Human T-cell Leukemia Virus type 1 (HTLV-1). Previous research has indicated that the HTLV-1 protein Tax may play a pivotal role in the development of ATL. This role that Tax seems to play in the development of disease, along with the myriad of functions this protein has been observed to perform, made it an ideal protein to study. A number of previous experiments have attempted to identify proteins that interact with Tax, using several methods including yeast two-hybrid, immunoprecipitation, western analysis, and 2-D gel analysis coupled with mass spectrometry.
The researchers began by performing a literature search for proteins that had been observed to interact with Tax, and then narrowed those some 67 proteins down further by limiting their analysis to only proteins that were involved in DNA damage repair. Out of this new list, they chose four proteins that had known functions and that could possibly affect a cell's DNA repair response: Rad51, TOP1, Chk2, and 53BP1.
In addition to the literature search, the researchers also produced their own physical Tax interactome. They accomplished this by using a S-Tax-GFP vector, which is a Tax protein with S-tags on the amino terminus and a GFP protein attached to the carboxyl end (It had previously been determined that this tagged version of Tax did not afffect protein binding). They limited their results to only nuclear binding proteins, and these proteins were then analyzed using LC-MS/MS (liquid chromatography and mass spectrometry). This was done three separate times, and 86% percent of the proteins were present on every run. The resulting proteins were then ranked according to their unique peptides, protein score, sequence converage, and relative abundance of predicted peptides from a protein to determine their relative binding strength to Tax.
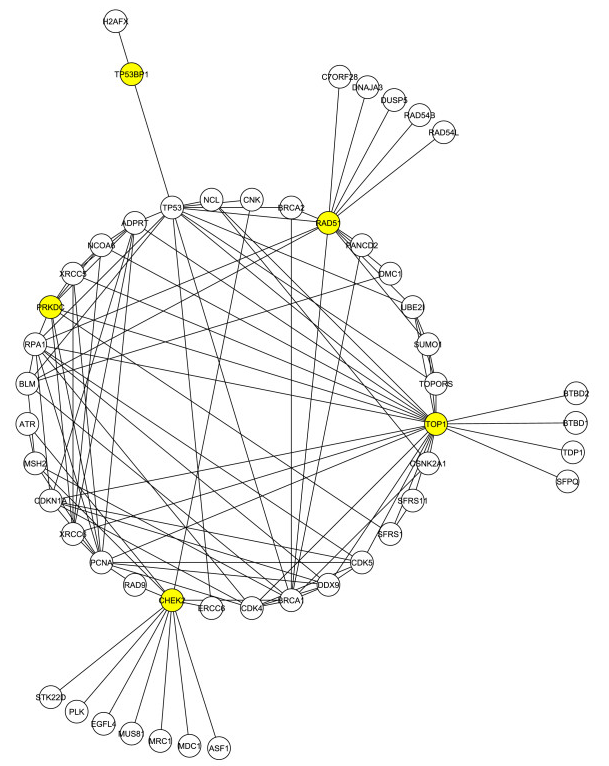
They used these four proteins and the Human Protein Reference Database to construct a network of first neighbors, which is shown in Figure 1 and referred to by the researchers as G1. G1 consists of 50 proteins involved in 112 interactions.

Figure 1. The first neighbor network of the 4 proteins identified from the literature. Highlighted in yellow are the 4 original proteins used to create the network, along with DNA-PKcs.
In this network, there is present a core of 12 proteins associated with DNA repair that highly interact with each other. Several of these proteins were also isolated in the mass spectrometry analysis mentioned above. Among this group of proteins, one novel protein in particular, DNA-PKcs, seemed to play a central role. Upon comparison to the rankings constructed from the physical Tax interactome, the researchers saw that DNA-PK was one of the highest ranking proteins that interacted with Tax.
The researchers then decided to look at the structure of G1 without the 4 original proteins used to create the network, in order to determine how the first neighbors interacted without interference from the original 4 proteins. This resulted in a new network consisting of 29 proteins and 60 interactions, which is shown in Figure 2.
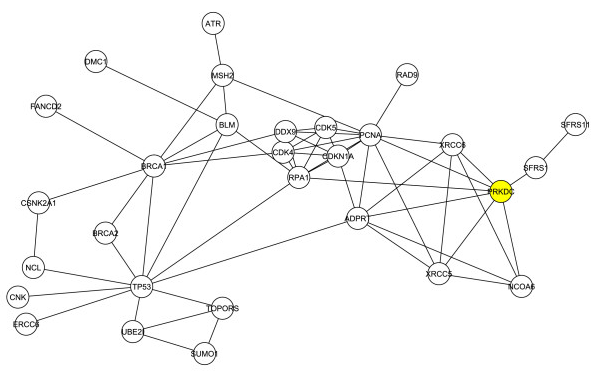
Figure 2. DNA-PKcs is highlighted in yellow.
DNA-PKcs (highlighted in yellow) was again shown to have a critical role within the network, and this network determines that DNA-PKcs's importance is largely independent of the original 4 proteins.
Next, a new network was created using G1, but this time limiting it only to the interactions involved in DNA repair functions. This new network (shown in Figure 3) was labeled G1*, and was made up of 26 different proteins with 42 interactions among them.
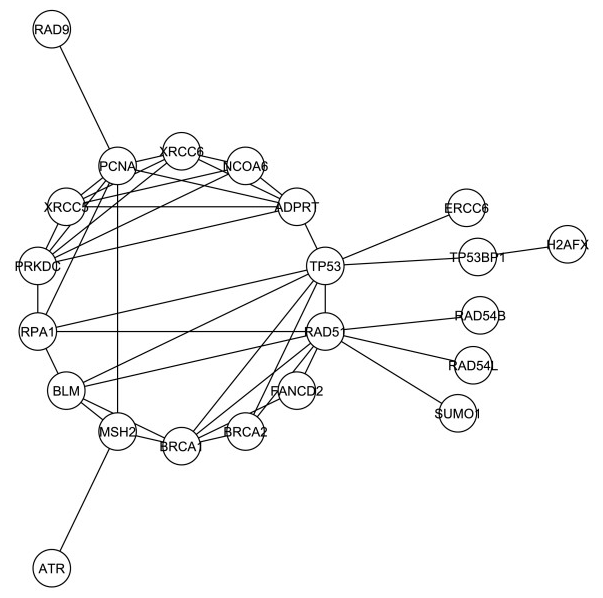
Figure 3.
In this new network that considers only the DNA damage repair response, DNA-PKcs (labeled in the figure as PRKDC) was found to have a central role once again, and in comparison to the other proteins actually rose in rank of degree and betweenness centrality, which are techniques to determine the importance of a protein in a network.
The next step in the experiment was to provide a broader context in which these proteins interact. A new network of proteins that interact with Tax was constructed that included second neighbors. This network consisted of 667 proteins and 3827 interactions. They then restricted this network to only include proteins involved in DNA damage repair. They refer to this new network as G2*, and it contained 114 proteins. Clustering analysis was done on the G2* network in order to determine the most integral proteins involved in the DNA repair response associated with Tax. This newly contructed network based on clustering in shown in Figure 4.
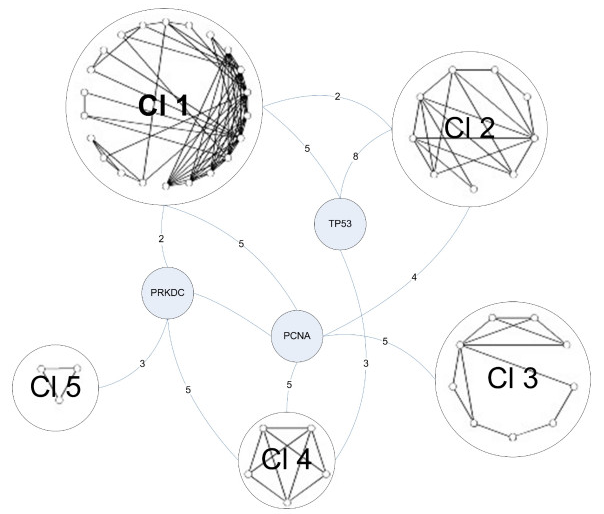
Figure 4.
Figure 4 consists of 5 separate clusters of proteins interconnected by only 3 individual proteins. One of these 3 linking proteins is DNA-PKcs. If Figure 4 is an accurate representation of what truly happens within a cell, then DNA-PKcs plays a central role in cellular response pathways regarding DNA damage repair.
To verify that Tax and DNA-PKcs do in fact bind together, the researchers performed an affinity pull-down analysis (much like immunoprecipitation) with S-Tax, using S-GFP as a control. Proteins that were found to bind to these proteins were isolated and immunoblotted with anti-DNA-PKcs, and the results are shown in Figure 5.
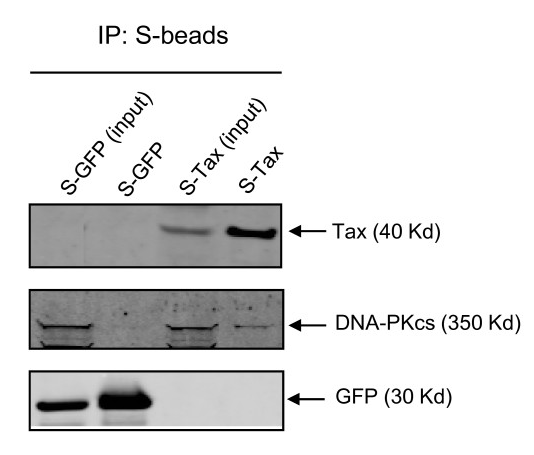
Figure 5.
The "input" columns 1 and 3 show the proteins extracted from each respective experimental transfection (S-GFP and S-Tax). Columns 2 and 4 show the results for each respective input after affinity purification. GFP and Tax were only found in the experiments where each of those proteins were transfected, but after purification, DNA-PKcs was found to only bind to S-Tax, providing conclusive evidence that DNA-PKcs is in fact is a protein that binds to Tax.
Opinion
This article has both good and bad points within it. The researchers did a good job explaining to the reader the steps they went through and establishing that DNA-PKcs plays an important role in the cellular DNA damage repair response. This is shown espeically nicely and conclusively in Figure 4. Also, throughout the paper the writers do a good job of remaining open to new possibilities, and even take a few lines throughout the article to give examples of other possible explanations. They make a convincing case, as most of aspects of their experiment were well controlled for. Good reasons and explanations are given regarding how clustering was done or how certain networks were created.
One problem I had when reading the paper was discerning the difference between DNA-PK and DNA-PKcs. This is never really addressed clearly in the article and I think that is important information the reader needs to understand. Also, referring to DNA-PKcs in the figures as PRKDC was confusing initially. Another issue I had with this paper is the importance of using both physical and in silico interactomes. The researchers do not do a very good job explaining the significance and usefulness of the two, or really even why they used both. There are little bits of information and explanation here and there, but no clear statements. The few references and little information that is given serves only to confuse the reader further.
Perhaps the biggest issue I have with this paper is the questions that burn in the back of my mind while reading it through: What is the significance of this experiment? Why is this important or worth reading? The writers explain in the beginnning the role Tax plays in the development of ATL, and also state that this experiment is for discovery purposes. This is all well and good, but what use is the determination of DNA-PKcs as an important protein? How can this knowledge be applied? The writers address none of these questions. Can this new information lead to treatment of ATL, or a better understanding of the disease? Perhaps, but the researchers never touch on the subject.
References
Ramadan, E, et al. (2008). "Physical and in silico approaches identify DNA-PK in a Tax DNA-damage response interactome". Retrovirology 5:92.
Return To Biology Course Materials
Davidson College Biology Department