Scientific Literature
In a novel use of genomic-era tools, Burke et al. (2010) conducted a controlled laboratory experiment using fruit flies (Drosophila melanogaster) to resolve long-standing questions about the process of evolution. The authors sought to determine how selective pressures affect loci within the genome, and they specifically wanted to gather data on a multicellular eukaryote model system as much previous work has focused on bacteria and yeast. The "classic" (or "hard" sweep) evolutionary model suggests that novel, advantageous mutations should sweep throughout a population until they become fixed. Alternatively, the "soft" sweep model describes a situation in which selection increases fitness by acting on multiple genetic variants that are already present within the population rather than forcing newly arisen mutations to fixation.
To address these questions, Burke et al. carefully bred fruit flies for over 600 generations, selecting those flies with the shortest hatching times to start the next generation. In this way, the researchers eventually developed a colony of flies that, when compared with controls, tended to hatch up to 20% faster, hatch at smaller body sizes, and have shorter life spans. Essentially, the researchers selected for a group of flies that developed "quick" life history characteristics in comparison to their slow hatching, long-lived peers. To determine how evolution had proceeded within the fly genome, the researchers sequenced genomic DNA from both experimental and control fly groups, and then used the Drosophila reference genome to search the sequences for single nucleotide polymorphisms (SNPs). Unsuprisingly, some of the SNP regions, the potential cause for the phenotypic differences between the two populations, were located in genes integral to development. When viewed at the whole genome level, the researchers found that allele divergence between the two fly strains was not contained in a small area (i.e., did not affect only a single locus) but was spread throughout a large portion of the genome (Figure 1).
The results were also somewhat surprising because no allele fixation was observed within the fly genome whereas one might expect to observe hard sweeps that force alleles to fixation in the presence of strong artificial selective pressures (such as those employed in this study). The authors provide a number of interpretations for their findings. One possibility is that the fruit fly genome has evolved through means of soft sweeps that have not yet resulted in allele fixation because the alleles being selected for have relatively small fitness benefits. Alternatively, the authors report that their data may in fact be consistent with hard sweeps but that the newly arisen beneficial alleles may have not yet reached reached fixation because of small selection coefficients. Finally, it may be that the beneficial effects associated with certain alleles actually decrease as the alleles become more common in the population due to effects of "linked deleterious alleles or antagonistic pleiotropy" (Burke et al. 2010). This relationship could also explain the lack of allele fixation in the fly genome. In any case, this study provides evidence that in sexually reproducing populations evolution may proceed by altering (but not fixing) allelic frequency at many loci throughout the genome.
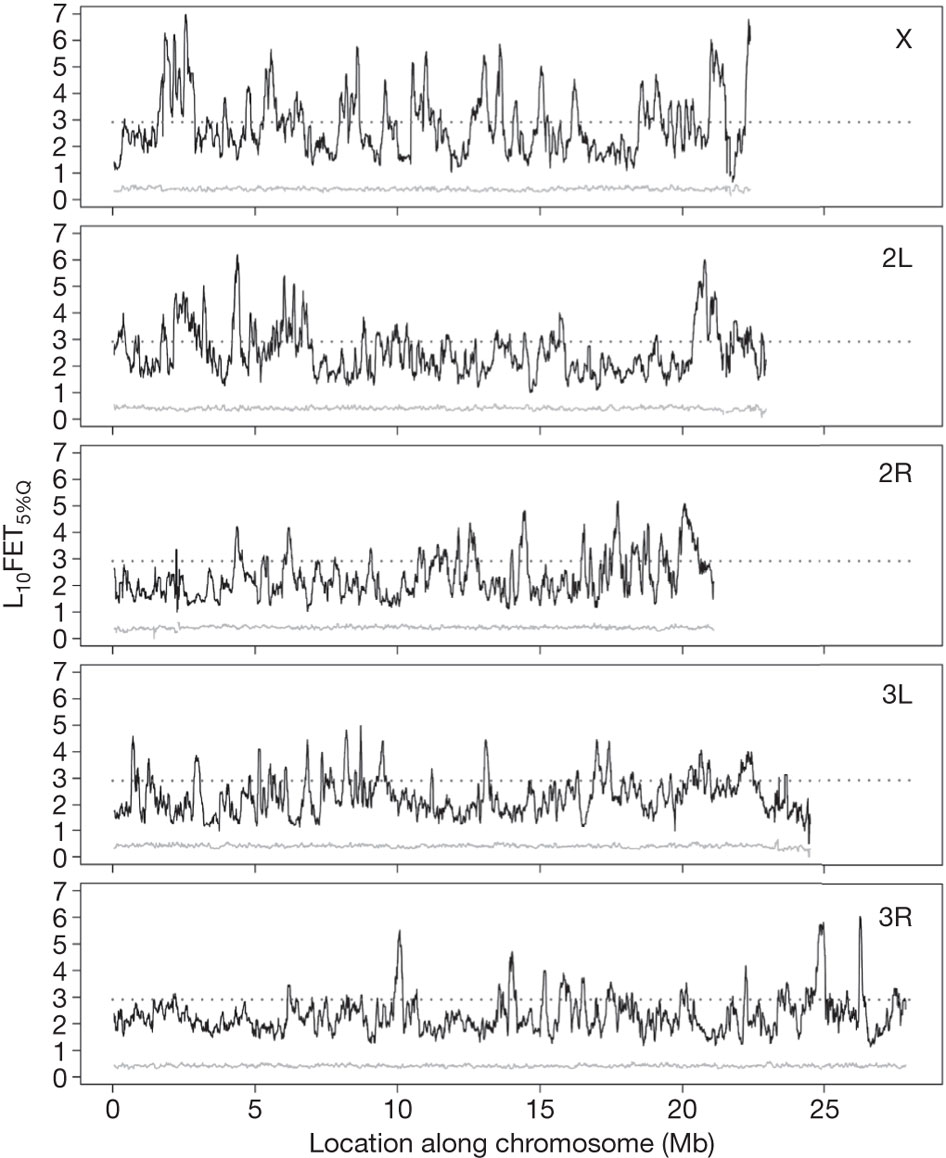
Figure 1. The solid black lines represent the degree of differentiation between experimental (artificially evolved) and control fruit fly populations. Peaks that extend above the dotted line represent areas of significant differentiation as determined by Fisher's exact test (FET) scores. Each of the five subpanels depicts a different fly chromosome. Notice that alllele differentiation has occured at many loci throughout the genome. The small grey line in each subpanel shows the differentiation scores for a single replicate population of experimental flies when compared with a pooled sample of five replicate populations of experimental flies. The low degree of differentiation seen in this comparison indicate that replicate populations within the experimental treatment are relatively similar at the genetic level. Reprinted by permission from Macmillian Publishers Ltd: Nature (Burke et al.), copyright 2010.
Popular Reporting
Popular science writers can serve as important intermediaries between research scientists and the general public. For that reason, it is all the more important that scientists ensure their work is being discussed accurately and in the proper context. Kua et al. (2004) provide useful guidelines for the evaluation of scientific writing, and here I employ those ideas in a discussion of Nicholas Wade's September 20, 2010 New York Times article describing the results of the Burke et al. 2010 study.
The first critical function of the scientific writer is to accurately communicate research findings and, if necessary, translate them into terms understandable to the general public (Kua et al. 2004). Overall, I find that Wade does very well in this regard. He effectively communicates the basic research design and evolutionary theory (i.e., the difference between "hard" and "soft" selective sweeps) necessary to understand the research without getting bogged down in scientific jargon. The article is also noteworthy for the included graphic that provides a fairly accurate visual representation of the differences between the two types of selective sweeps. Such visual aids can prove extremely useful in communicating scientific ideas to the public without resorting to long-winded verbal descriptions. Unfortunately, I believe Wade tends to simplify the conclusions reached in the study. While he flatly states that a soft sweep was responsible for the observed phenotypic differences in the fly strains, the paper authors offer alternative conclusions in their discussion, including the possibility that a classic, hard sweep may have been observed within the fly genome given more selective time. As Kua et al. (2004) mention, qualifying statements are often left out of popular scienfitic articles, and while he was obviously under space constraints, it would have been preferable if Wade had tempered his statement about the paper's ultimate implications.
In addition to accurately conveying the methods and findings of research efforts, it is equally important that the scientific writer provide enough context so that readers can effectively gauge the significance of the work being described (Kua et al. 2004). Wade begins his article by stretching the context of the research article in saying that the paper's results may impact medicine and drug development. While in a very broad sense this may be true (as Wade mentions, diseases influenced by multiple interacting genes will likely be difficult to control with drug therapy), the cited research article makes no mention at all of any potential medical implications. Aside from this reach for significance, Wade does a good job of providing context for the research being described. He quickly summarizes the historical debate about the mechanism of evolutionary sweeps which this research addresses (although does not completely resolve). He also does well to include the opinions of other evolutionary scientists not directly involved in the work. Wade includes comments from one scientist who conducts similar evolutionary research on bacteria, providing a useful contrast to the Burke et al. 2010 study which is unique precisely because it examined selective sweeps in a multicellular, eukaryotic organism.
In conclusion, this popular report concerning scientific research falls into some of the common traps associated with scientific writing, namely the tendency to use absolute rather than qualified statements concerning the results and overstating the greater implications of those findings. Overall, however, Wade effectively fulfills the two primary functions of a scientific writer in that he attempts to accurately convey the methods and data in an easy-to-understand way while also providing the contextual information necessary to place the research findings in the broader scientific landscape. I believe his work provides a good model for those seeking to translate scientific literature into popular reporting.
References
Burke, MK, Dunham, JP, Shahrestani, P, Thornton, KV, Rose, MR, and Long, AD. 2010. Genome-wide analysis of a long-term evolution experiment with Drosophila. Nature 467: 587-590.
Kua, E, Reder, M, and Grossel, MJ. 2004. Science in the news: the study of reporting genomics. Public Understanding of Science 13: 309-322.
Wade, N. 20 September 2010. Natural selection cuts broad swath through fruit fly genome. The New York Times.