
This website was produced as an assignment for an undergratuate course at Davidson College
Home safsaCurly Hair Gene dsafPaper Review 1sdaf Paper Review 2

Movember safsaScientific Article safsWhat's Next?
What Did Garraway et al. Do?
Garraway et al. sequenced the entire genome of seven men with prostate tumors of stage T2c or greater. Some elements they looked for were point mutations (in particular non-silent ones) as well as rearrangements (Fig. 2). The compared the mean mutation frequency of 0.9 per megabase to the mutation rates found in other cancers. 0.9 is similar to the rate found in myeloid leukaemia and breast cancer, but quite a bit lower (7-15 fold) than small cell lung cancer and melanoma. The paper focuses more on rearrangements (red arrow), but I find it interresting that there does not appear to be a correlation between mutation rate and the number of rearrangements.
Figure 2: The landscape of somatic alterations in primary human prostate cancers. Tumors with TMPRSS2-ERG gene fusions are starred.
They obtained approximately 30 fold sequence coverage for each genome, and used this data to produce Circos plots (Fig. 3). Circos plots take the 3 billion letters that make up our DNA and depict the information circularly. Their primary function is to allow easy visualization of relationships within the genome. The outer ring is divided and labeled by chromosome. The inner ring shows chromosomal copy number, copy gains are red, copy losses are blue. The three patients on the top row exhibit a TMPRSS2-ERG gene fusion, while the four below do not. The green arcs within the Circos plots show interchromosomal translocations, or areas that have rearranged within a chromosome. The purple arcs represent intrachromosomal rearrangements, or areas where DNA has moved from one chromosome to another.
Figure 3: Circos plots of the seven prostate tumors. The top three (PR-0581, PR-1701, and PR-2832) have a TMPRSS2-ERG gene fusion, while the four on the bottom are fusion negative.
Unfortunately the red and blue on the inner ring is difficult to see unless the figure is enlarged (Fig. 4, the arrows are colored to match the colors indicating copy number change). However despite problems visualizing copy number changes, Circos plots are an effective way to present the data Garraway et al. produced. The numerous purple arcs foreshadow the upcoming discussion of chromosomal rearrangements. One of the limitations of Circos plots is that the purple arcs do not provide much information about what type of loci were exchanged or whether the exchanges were balanced. Patient PR-2832 has noticeably more purple arcs than PR-0508, but without knowing what the rearrangements interrupt, it is impossible to know whether the rearrangements actually cause any problems.
Figure 4: Detail of a circos plot. The arrows highlight areas of copy gain (red) or copy loss (blue).
Why Sequence The Whole Genome?
An interview with the lead investigators of this paper can be found on the Prostate Cancer Foundation's website. In the future the Prostate Cancer Foundation hopes that the cost of sequencing the genome ($20,000 or less) will be less than the cost of overtreating prostate cancer. The foundation estimates that $1.5 billion is spent each year on unnecessary treatments. Despite advances in the treatment of prostate cancer, it is still largely unknown what actually causes the disease. There seems to be a genetic component, as prostate cancer can run in families, but correlations have also been noted between prostate cancer and a multitude of different lifestyle factors.
What Kinds of Rearrangments Were Observed In The Genome?
Garraway et. al found a median of 90 rearrangements per genome, which is similar to the rearrangement number found in breast cancer. Interestingly, they found that in many cases the chromosomal rearrangements were balanced, meaning that there was no overall loss of genetic material (Fig. 5).
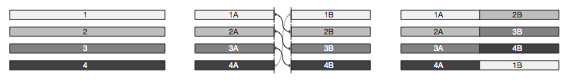
Figure 5: Theoretical balanced rearrangement. Although multiple loci are interrupted there is no overall loss of genetic material.
One such example is the TMPRSS2-ERG gene fusion, found in three of the prostate cancers (Fig. 6).
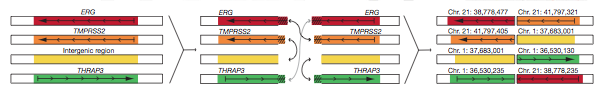
Figure 6: TMPRSS2-ERG gene fusion, found in three of the seven prostate cancer tumors. This particular fusion is from tumor PR-1701, but is representative of the other two.
The TMPRSS2-ERG gene fusion is a closed quartet of balanced rearrangements (Fig. 7). I found Fig. 7 to be relatively hard to decipher. They are attempting to show the rearrangement of two chromosomal regions on chromosome 21 and 1. The whole chromosome is shown above, with the four zoomed in regions shown below. The blue and purple arcs connect discordant read pairs (purple indicates interchromosomal, blue indicates intrachromosomal). Although the arcs connect colored bars, it is unclear what the colors mean, as they do not match the colors which represent each loci shown in Fig. 6. In addition they state that the directionality of the thin gray bars represents mapping orientation, but the bars are so tiny, it is impossible to see without zooming in a lot (Fig. 8). In Fig. 8 a thin gray bar with directionality is indicated with a green arrow. Strangely some of the bars have no directionality (red arrow). There are also several small colored lines, not connected by blue or purple arcs (indicated with black arrows) that I could not find an explanation of.
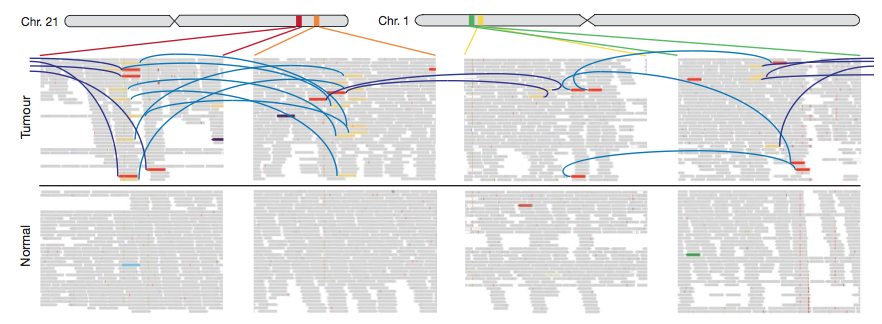
Figure 7: Rearrangment of four loci on two chromosomes in PR-1701.
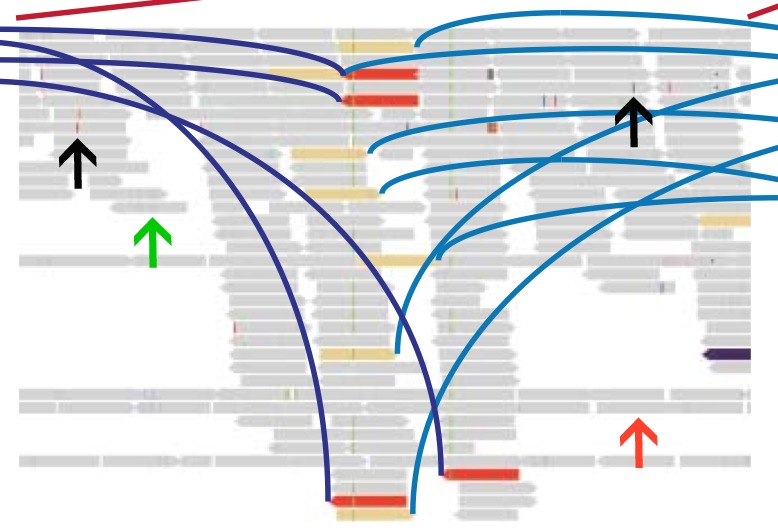
Figure 8: Detail from Figure 7. The green arrow indicates a thin gray line with directionality, the red arrow indicates one without directionality. The two black arrows indicate features for which I could find no explanation.
In addition to the relatively simple rearrangement shown in Fig. 6, some patients had far more complicated breakage and fusion (Fig. 9). Fig. 9 illustrates a rearrangement that involved nine loci on four different chromosomes. A general problem with these rearrangement figures is that they really only give a good idea of where the genes came from in the normal genome, not where they are now localized in the tumor genome. The labels above each loci provide the origin, not the destination.
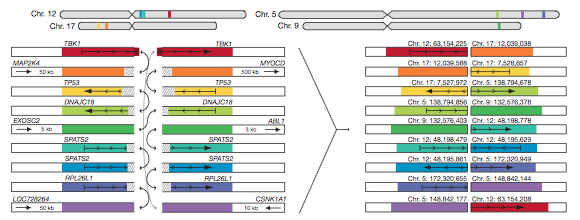
Figure 9: Complex rearrangement observed in PR-2832. Breakpoints and fusions occur on four different chromosomes and involved nine different loci. Like in Figure 7, some rearrangements are interchromosomal while others are intrachromosomal.
Were There Any Patterns to The Rearrangements?
Garraway et al. suspected that there might be factors that lead to different genomic regions becoming co-localized before rearrangement. It had been shown previously (Lin et al., 2009) that androgen signaling could induce the co-localization of the TMPRSS2 and ERG genes (the same genes shown to be fused in Figs. 6 and 7). Garraway et al. proposed that migration might occur to transcription factories. Transcription factories are discrete sites within the nucleus that contain RNA polymerase II holoenzyme. To test this theory they looked at the genomic regions involved in the rearrangements and characterized transcriptional patterns and chromatin marks. ChIP-Sequencing uses chromatin immunoprecipitation and parallel DNA sequencing to determine how proteins interact with DNA (Fig. 10).
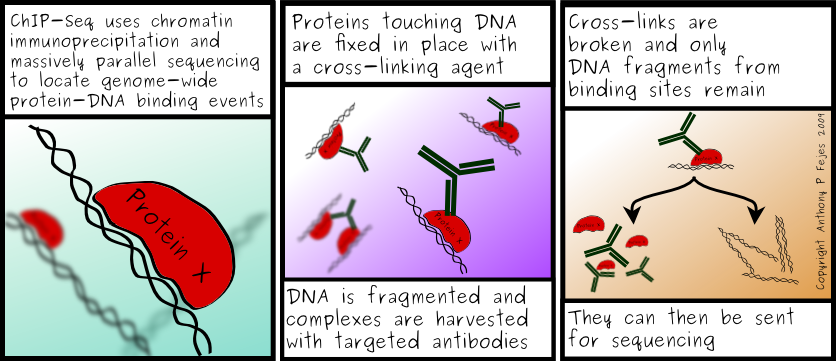
Image from Fejes.ca
Figure 10: Cartoon explaining ChIP-Seq.
Using the ChIP-seq data from a prostate cancer cell line they found significant spatial correlation with various marks of open chromatin (Fig. 11). TMPRSS2-ERG fusion tumors are shown in black, in general near marks of active transcription (green arrows) rearrangements are enriched and near a mark of closed chromatin (red arrow) rearrangement is depleted. The opposite is true of TMPRSS2-ERG fusion negative tumors. Enrichment versus depletion was determined by calculating the log of the ratio of observed breakpoint rates to background rate around ChIP-seq peaks. This means that areas of high enrichment have lots of rearrangements and areas of high depletion are very stable.
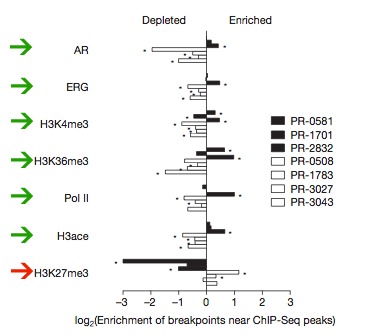
Figure 11: Association between rearrangement breakpoints and genome- wide transcriptional/histone marks in prostate cancer. Green arrows indicate marks of active transcription, the red arrow highlights a mark of closed chromatin.
Because they only used black and white in this figure, it becomes a bit harder to read. It would have been easier to color code each patient, so you did not have to compare the order in the legend to the order next to each ChIP-seq peak. However the advantage of using black and white is that the pattern of TMPRSS2-ERG fusion positive genes versus fusion negative genes is easier to visualize.
Were Any Other Genes Preferentially Affected Like TMPRSS2 and ERG?
In addition to the three patients found to harbor TMPRSS2-ERG gene fusions, three patients were also found to contain rearrangements of CSMD3 and CADM2. These rearrangements occurred at frequencies higher than simply chance would explain. CSMD3 is a large gene that has multiple CUB and sushi repeats. Despite the presence of CSMD3 rearrangements in three of these patients, upon more analysis of ninety-four more prostate tumors no other CSMD3 rearrangements were found. CADM2 encodes a nectin-like cell adhesion molecule. Some nectin-like proteins have shown tumor suppression properties in other contexts, thus the CADM2 rearrangement could certainly be involved in prostate cancer. Breakpoints in CADM2 were found to be intragenic by both sequencing and FISH (Fig 12). While the FISH is relatively convincing, it does appear that the gene has broken apart and now two regions are located further apart, the arrangement could also be due to twisting of the chromosomes, which is not visible.
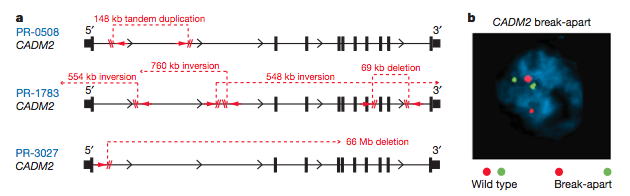
Figure 12: Diagram of breakpoints in CADM2 in three different prostate cancer tumors. FISH analysis confirmed the break-apart of CADM2. The two flourescent markers clearly appear to be further apart on one chromosome than on the other (wild-type).
The PTEN tumor suppressor gene was also found to contain breakpoints in two of the prostate tumors (Fig 13).
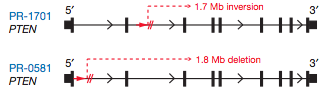
Figure 13: Intragenic breakpoints in PTEN.
Two other tumors had rearrangements in the MAGI2 gene (Fig. 14). MAGI2 encodes a protein which interacts with PTEN, the tumor suppressor gene (Fig. 13). Like in Fig. 12, Garraway et al. used FISH to confirm the inversion. You can see that on one chromosome (wild type) the order appears to be red, green, green, while on the other inverted one the order is green, red, green.
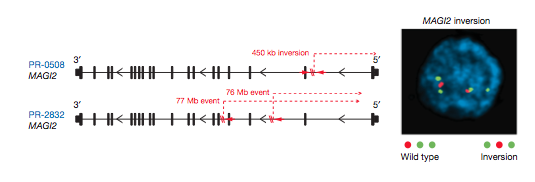
Figure 14: Intragenic breakpoints in MAGI2 plus FISH confirmation of the inversion found in another prostate tumor.