This web page was produced as an assignment for an undergraduate
course at Davidson College.
Review of "Evolutionarily Assembled cis-Regulatory Module at a Human Ciliopathy Locus"
by Lee et al. (2012)
Background
The neurodevelopmental disorder, Joubert syndrome, is one type of many cilliopathy disorders. These disorders correspond to faulty structure or function of primary cilia. Lee and colleagues studied Joubert syndrome previously and found the gene TMEM216 for a transmembrane protein linked to half of a subset of Joubert syndrome patients. However, the other half of these patients had normal TMEM216 genes and gene products. These patients had neither DNA mutations, nor abnormal mRNA or proteins. Thus, Lee et al. (2012) decided to continue investigation. Their 2012 study, “Evolutionarily Assembled cis-Regulatory Module at a Human Ciliopathy Locus,” examines what causes the other half of the Joubert syndrome subset and the interplay between the two causes.
Methods
Initially, Lee et al. sequenced all of the known and predicted exons and promoters in an interval the questioned patients had in common, which was near TMEM216. Sequencing allowed Lee et al. to narrow down possible mutations that contributed to the disease state. From this analysis, they looked for what gene existed near the mutations.
Once they determined TMEM138 as the link between all the questioned Joubert syndrome patients, they sequenced and analyzed the gene and protein product. Comparing TMEM138 to TMEM216 at this level led Lee et al. to preform phylogenetic analysis. They analyzed the genes and the intergenic regions for their evolutionary conservation.
To test how these genes both relate to Joubert syndrome, Lee et al. began asking questions about the function of both genes and their products. They looked at database protein microarray data and in situ hybridization of human embryos to examine coordinated expression of the gene products. In examining the role of the genes’ neighboring locations in expression, they used their evolutionary data. They compared the mRNA expression patterns in zebrafish and mice, which have separated and clustering of the genes, respectively. To do so, they used quantitative PCR of mRNA from various tissues.
Once Lee et al. understood that the location of the genes next to each other in high vertebrates has significance, they questioned what within the evolutionarily conserved intergenic region could cause the effect. After determining that a link between the expression of both genes does exist, Lee et al. could continue asking questions about the connections between the gene products.
Lee et al. tested whether or not the protein products share a function by examining the phenotypes of mutant and knockdown cells for each gene. Furthermore, they looked at the cellular location of each protein using immunostaining. Similarly, they used time-lapse fluorescence to study the position of the proteins in more detail. From this, they analyzed what the result of the protein’s function included.
At this level, Lee et al. preformed knockdowns of genes using siRNA to test the requirements of vesicular movement. Knowing the relationship between the two proteins in preforming their task, they looked for a protein that physically linked the two (by linking the vesicles). To confirm their predicted protein (TRAPII), they preformed a knockdown of a subunit of TRAPPII and used live-cell imaging to view the state of the target vesicles.
Now understanding the interaction of TMEM216 and TMEM138, Lee et al. decided to question how the interaction came to be that way. Thus, they tested whether the protein localization and function came before the coordinated expression and regulation. They knew that expression was not coordinated in the lower vertebrates from their phylogenetic analysis, so Lee et al. studied these aspects in zebrafish. Moreover, Lee et al. wanted to examine if the genetic function of the genes was shared between the proteins in the lower vertebrates, knowing that the proteins localized to the same place as in humans. To test this, they preformed more knockdowns of the gens and looked at the phenotypes of the zebrafish.
Results
Lee et al. found four missense and one deletion mutation common in the questioned patients near an unclassified gene, TMEM138. Moreover, they saw that the mutations behaved together in a simple recessive form that likely illustrated a loss-of-function pathway. This discovery fit with their understanding of Joubert syndrome phenotypes. When looking at TMEM216 and TMEM138 together, their data showed that while both are transmembrane proteins, they have no sequence homology or shared protein domains.
From their phylogenetic analysis, Lee et al. found that the differences between the two genes occur because they are from different protein families that convergently evolved. They also found that the genes were conserved in all the higher vertebrates along with their location and orientation. TMEM216 and TMEM138 in higher vertebrates are in a head to tail orientation, and their intgergenic region was also conserved. Lee et al. found that this conservation occurred after the amphibian and reptile split.
In the testing of the protein expression patterns, Lee et al. found that the proteins had coordinated expression in kidney, cerebellar buds, and telencephalon tissue in human embryos between four and eight weeks old. From their test for the importance of their chromosomal locations so near eachother, they found that while mice did have coordinated expression, zebrafish with separated TMEM216 and TMEM138 did not. When they looked for what could cause the coordinated expression, they found a transcription factor that binds within the intergenic region correlated with the expression. This regulatory protein was X4, and its binding in the DNA nearby both genes is an example of a cis-regulatory unit that works on both genes.
Looking at the possibility of the proteins functioning together when expressed together, the phenotypes of the mutants and knockdowns showed short or no cilia, mutated fibroblasts, and defects in cilliogenesis from loss of either gene. Thus, Lee et al. found that both genes are necessary for the development of healthy cilia.
When the interaction of the proteins was examined further, Lee et al. found that the cellular location of both proteins was at the base of cilia and in (separate) nearby vesicles. In addition, they found that the vesicles with TMEM216 went to the centrosome while the TMEM138 vesicles tethered to the TMEM216 vesicles. They also found that these vesicles contain proteins targeted for cilia. For example, the vesicles carry protein CEP290.
While the vesicles of each protein work together, Lee et al. found that the movement of the vesicles depended on the proteins. The TMEM216 vesicles functioned independently despite lacking TMEM138 protein; however, TMEM138 vesicles could not continue to function in the TMEM216 knockdowns. Thus, TMEM138 function was dependent on TMEM216, which means the movement of the CEP290 protein was dependent as well. This dependency based on the vesicles linking together was confirmed when Lee et al. showed that TRAPII (tethering protein) knockdowns separated the vesicles.
In examining the evolutionary significance of these interactions, Lee et al. found that the coordinated vesicular movement with the proteins preceded the coordination of gene expression. In contrast, they found that the when the genes were faulty in lower vertebrates each protein shared some similar phenotypes, but TMEM216 mutants had extras that TMEM138 mutants did not exhibit. Thus, they showed that the genes originally did not have such similar function, and that the switch to clustering the genes likely made that leap.
Figures analysis (Author's permission to reproduce figures pending)
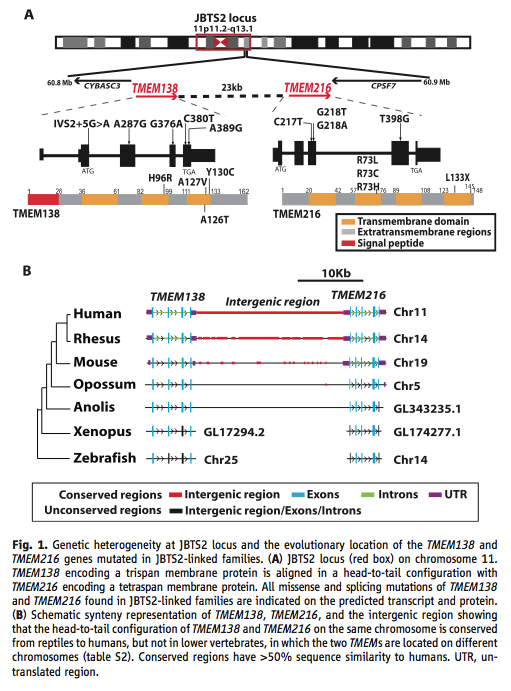
Figure 1. Schematics of the genes, the proteins, and the conservation throughout evolution. (A) This panel shows the location of the linked genes, their orientation, and the mutations as they affect the mRNA and proteins. All occur within the realm known for Joubert syndrome. It makes the visualization of the neighboring aspect of the gene easier to follow. (B) This panel shows the synteny and conservation of the two genes. The similarity of the exons and the intergenic regions decreases with distance from humans. Lower vertebrates have the genes, but they are not on the same chromosome. They base conservation on 50 percent similarity in sequence compared to humans.
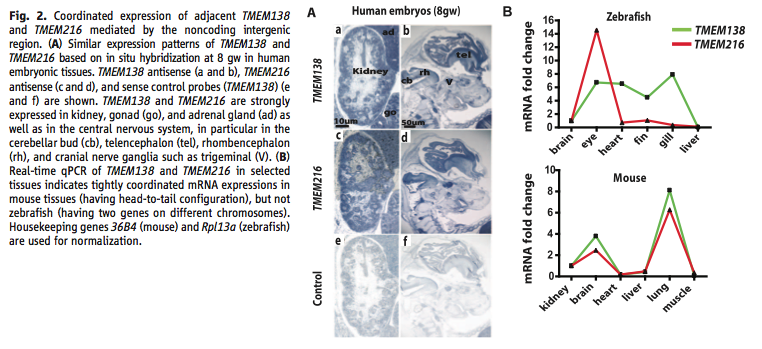
Figure 2. Coordinated expression tests. (A) This panel shows the location of each gene’s mRNA using in situ hybridization of eight-week human embryos. They show a control for their method. It shows the tissues with gene expression. (B) This panel shows the data from real-time qPCR of mRNA for each gene in zebrafish and mice. It shows that the separated genes in zebrafish do not show coordination in various tissues, but that mice do. The scale for the zebrafish plot diminishes the difference that they found.
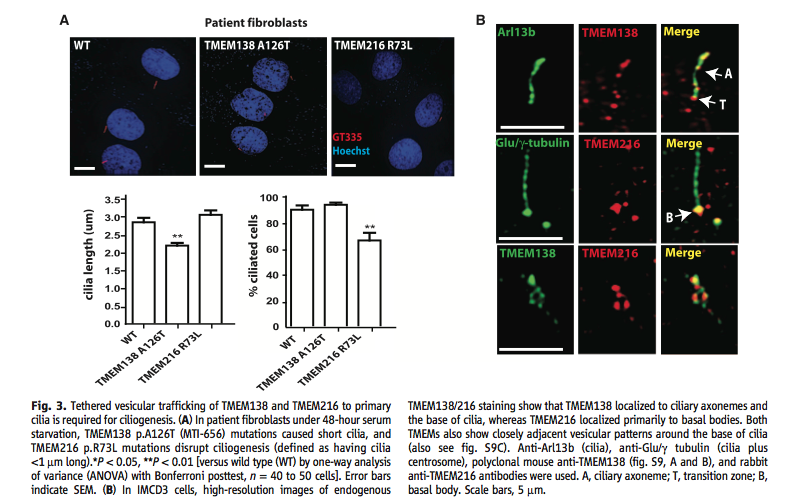
Figure 3. Examination of phenotype and cellular location of proteins. (A) The top part of this panel shows the effect that mutations of TMEM138 and TMEM216 have on cilia. Shortened and non-existent cilia occur in the mutated fibroblasts. The bottom part of the panel quantifies the image. (B) This panel shows the cellular location of the two proteins. They show where the genes localize individually. The Arl13b and Glu/γ-tubulin images show the parts of the cilia and area the proteins could go, while the merged individual images show what part of this system the proteins fit in. The bottom row shows how close the two proteins are to each other.
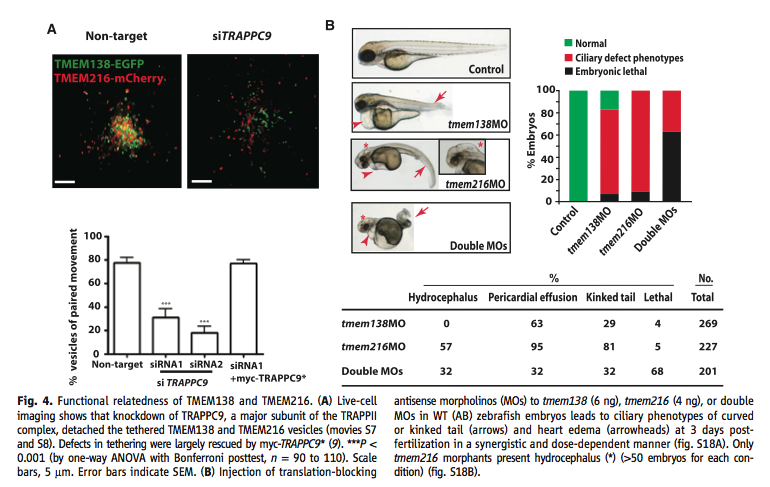
Figure 4. Testing the combined function of the genes and the presence of this coordination before clustering. (A) This panel shows live-cell images of normal cells and ones with the vesicle tethering protein knocked down with siRNA. They show both qualitative and quantitative representations of their results. In addition, they show that the effect of knockdown can be reversed when a tagged version of the tethering protein was introduced. (B) This panel shows the effect of knockdowns of each gene in zebrafish. Top left shows the phenotype, while top right quantifies the phenotype. It shows that each gene causes the effect even though on separate chromosomes. It also shows that having neither has an extreme result of lethality. The bottom table shows the number of embryos with specific defects based on each knockdown. Hydrocephalus only occurs when TMEM216 is mutated, showing that some difference in the functionality of the proteins before linkage, clustering, and co-regulation evolved.
Conclusions
Overall, Lee et al. concluded that evolution assembled genes with similar function and rearranged them in order to organize them by that function. Moreover, they found that evolution can create regulatory systems for clusters of genes that further organize and coordinate the similar function. Thus, the act of clustering genes can lead to more similarity in the genes’ functions. In this case, Lee et al. found that such changes and regulation can occur even with nonparalogous genes. In relation to the disease state in question, Joubert syndrome, the interaction of the two gene products and the coordinated regulation that Lee et al. found highlights why the subset can come from faultiness in either one of the genes. Along the way, Lee et al. were able to make conclusions about the interactions of two genes and their gene products. They figured out the regulation that linked the two genes and the cell location, physical interaction, dependency, and function of the proteins.
Opinion
In general, this paper does show satisfactory evidence for what it concludes. The paper methodically determines the interactions and functions of the genes and product, with supportive results. In addition, I like that Lee et al. examine the protein and gene interactions in an evolutionary context. This aspect illustrates the significance of having such interactions while it also helps support their conclusions about the function of the genes and gene products. However, I do find some faults in the paper, mostly within the figures.
Starting with the figures: First, in Figure 1 (A) the representation crowds the information to a point that makes it hard to follow. Also in Figure 1, they do not explain much of how they got the data in the representations within the figure. While some of the process is explained in the text, the reasoning and methods are still unclear. Second, in Figure 2 (B) the plot does not show the stats that Lee et al. provide in the text, which seems unnecessary given the space they have. In addition, they do not say what time point they used for this part of the figure and they assume that any differences are due to the genes' location and orientation. Given that the species are so distant, many other complexities may exist. However, all this questionability only adds to the fact that they connected their expression data with lines. This suggests that the expression levels slowly changed between tissues, when that data was not taken. It seems that this data would have been better represented with a clustered bar graph. Third, in Figure 4 (B) it seems questionable to assume that all the effects shown come from the location of the genes and not at all from other differences between the distant species. The data do seem to show a large enough change to believe. However, without statistics and a discussion of why they think they can assume location was the only factor the result can still be questioned. Looking at the care put into this figure, the bottom table is poorly made—the lines are not centered. While unimportant, this sloppiness highlights the nature of the real issues in the figures and data.
In the text of the article, there are some other areas of concern. At the beginning of their exploration Lee et al. limited their sequencing to known and predicted exons and promoters in the disease-linked region of patients. This limits their findings to what the Human Genome Project found, but there could be other factors involved that sequencing techniques do not pick up. In addition, they assume that the causes could only be within exons and promoters. Thus, there could have been other mutations in the region that affected another gene nearby, complicating the interactions beyond just TMEM216 and TMEM138. In the rest of their discussion, Lee et al. refer to data that they do not show and methods they do not explain. In two cases Lee et al. write, “We further examined several experimental features” (2012). While supplemental material could resolve this issue, and it likely stems from limited space, the conclusions they reach after these tests seem important enough to present in at least a limited form within the published article.
References
Lee, J. H., Silhav, J. L., Lee, K. E., Al-Gazali, L., Thomas, S., Davis, E. E., Bielas, S. L., Hill, K. J., Iannicelli, M., Brancati, F., Gabriel, S. B., Russ, C., Logan, C. V., Sharif, S. M., Bennett, C. P., Abe, M., Hildebrandt, F., Diplas, B., Attie-Bitach, T., Katsanis, N., Rajab, A., Koul, R., Sztriha, L., Waters, E. R., Ferro-Novick, S., Woods, C. G., Johnson, C. A., Valente, E. M., Zaki, M. S., Gleeson, J. G. 2012. Evolutionarily assembled cis-regulatory module at a human ciliopathy locus. Science 335: 966-969.
Return to Main Page
Genomics Page
Biology Home Page
Email Questions or Comments
© Copyright 2012 Department of Biology, Davidson College,
Davidson, NC 28035



