This web page was produced as an assignment for an undergraduate course at Davidson College.
JAMES HARDEN's
Genomics Web Page
Sex Differences in the Gut Microbiome Drive Hormone-Dependent Regulation of Autoimmunity
The purpose of this article is to give insight about the effects of the microbiome on regulating hormones and protecting against autoimmunity as a result. It is relevant and significant to our growing body of knowledge on autoimmune diseases because researchers typically focus only on internal factors of pathogenesis such as disease-linked genes or loci. Therefore, there is a lot of genome-wide studies. The problem is that there is incomplete concordance in monoygotic twins, indicating the influence of environmental factors that is unaddressed with GWAS. In addition, there is a strong female bias for the prevalence and incidence of autoimmune diseases that is also unexplained by GWAS. This article addresses both the environmental factors as well as the sexual dimorphism that impact autoimmunity by studying the effects of microbiome in nonobese diabetic (NOD) mouse models. This mouse model displays spontaneous, immune-mediated pancreatic beta cell destruction, causing type 1 diabetes (T1D).
I thought that this was an extremely interesting article that highlights the importance of researching the microbiome and emphasizes the deeply intertwined interactions between bacteria and our own cells. I felt that this was definitely preliminary research that should be followed up with further studies but I believe this is a promising field that will shed light to new understandings regarding the importance of our gut biology. Throughout this discussion I will provide commentary on specific concerns, questions, and/or interesting findings.
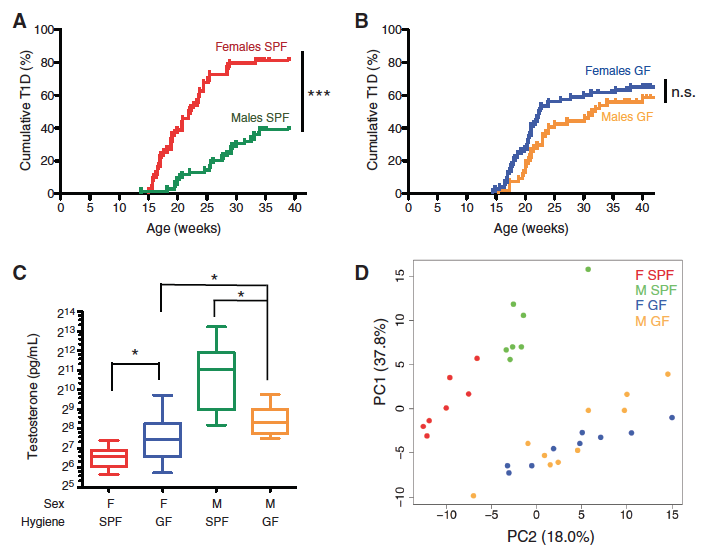
Figure 1 – Sex-biased autoimmunity in the NOD mouse depends on the microbiome.

There is evidence from previous studies that (T1D) is positively correlated with better hygiene status, so these researches are interested in researching the effect of the gut microbiome on autoimmunity. The researchers subjected NOD mice to either specific-pathogen-free (SPF) or germ-free (GF) environments, and Figure 1A and 1B show the cumulative % of T1D over a few weeks time for these two conditions. In the SPF condition (Fig. 1A), females had a significantly higher cumulative % of T1D than males, but there was no significant difference in GF condition (Fig. 1B). The relative protection of males in the SPF setting appeard to be dependent on the presence of commensal organisms.
Because of the sex-dependent trends that they found in Fig 1A and 1B, they tested to see if microbial colonization affected sex hormone levels. Figure 1C shows us that GF females displayed elevated testosterone relative to SPF females, and GF males had lower levels relative to SPF males, indicating that commensal colonization regulated testosterone production. It would be interesting to know the mechanism by which gut microbes affect testosterone production and thereby provides protection against autoimmunity, but that seems to be beyond the scope of this article.
Next, the researchers wanted to identify the bacteria in the males and females that explain the variance between the two GF and SPF NOD groups, so they used mass spectrometry-based metabolome analyses since previous studies support this method to be an accurate quantification of metabolites. The principal component analysis (PCA) in Figure 1D revealed a subset of glycerophospholipid and sphingolipid metabolites that accounted for 55.8% of the total variance between all four groups, though it did not distinguish males and females from GF condition. This further supported their claim that the microbiome regulates sex-specific features that are involved with autoimmunity protection.
Figure 2 – Sex-specific microbiome profiles emerge after puberty, and cecal microbiome transplantation can stably alter the microbiome of the host without inducing systemic immune priming.
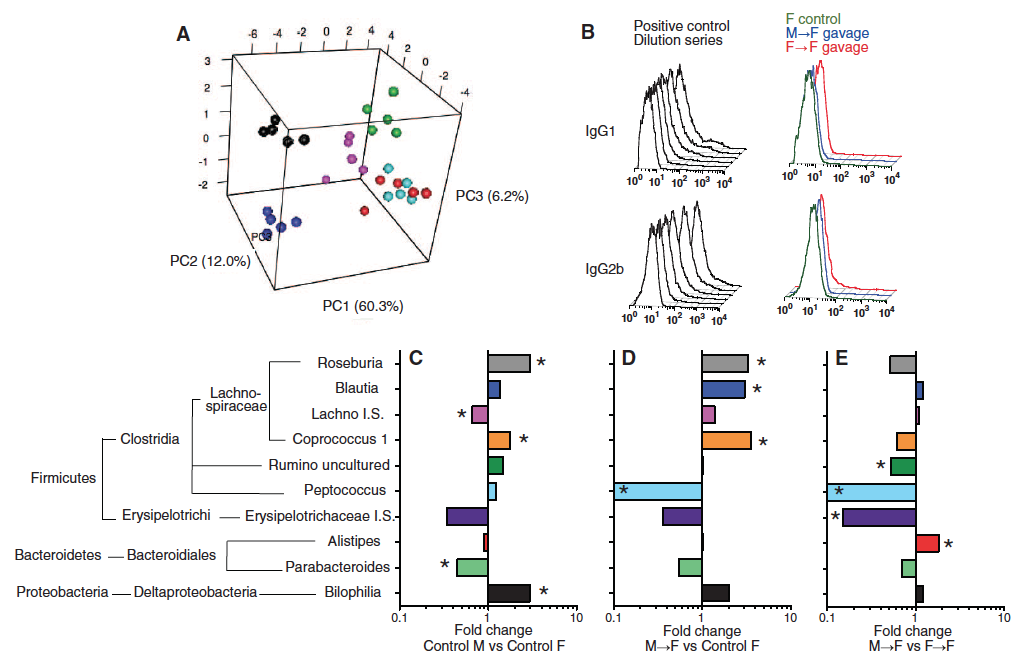
The next step was to sequence the 16S rRNA of the cecum microbiome. The researchers used PCA again to detect age- and sex-specific phenotypes. (Figure 2A: 3-week old males– turquoise; 3-week old females – red; 6-week old males – pink; 6-week old females – green; 14 week old males – blue; 14 week-old females – black). Because the red and turquoise are not separate, it appears that only mature males and females harbored distinct cecal microbiota. It is amazing to see that 30 bacterial families across only 6 different groups can explain 78% of the variance between male and female NOD mice. It is unknown why certain bacterial families prefer one sex over another. Later in this article the researchers will perform cecal transplantation from males to females to show that females have higher testosterone production upon transference (Fig. 3), but I cannot tell from their findings whether or not the females would eventually restore balance to their naturally occurring microbiome or if the new bacteria from males are self-sustaining in the females. This would be interesting to me to know if the seeming regulatory properties of microbiome on hormone production are also reversible such that the gut bacteria is actively regulated (perhaps through some positive feedback loop) to maintain a certain bacterial composition. The researchers seem to partially address this later in Figure 3D and 3G.
It was useful to know whether or not the transplantation was inducing an immune response, and whether or not the transplantation was altering the bacterial composition of recipient NOD mice. To address the first issue they measured systemic IgG1 and IgG2b for M --> F and F --> F gavage, as well as F control (Fig 2B. right histograms). They inoculated mice with bacteria for a positive control (Fig 2B. left histograms). According to these results, there was no immune response to the transplantation. To address the second issue, the researchers performed 16S rRNA sequence pairwise comparisons and showed the 10 bacterial genera that were found to be significantly different in at least one of the comparisons (Fig. 2C). This makes me wonder if there are any pathogenic bacteria genera that prefer females over males and vice versa.
Figure 3 – Transplantation of the male microbiome results in hormonal and metabolic changes in the female recipient.
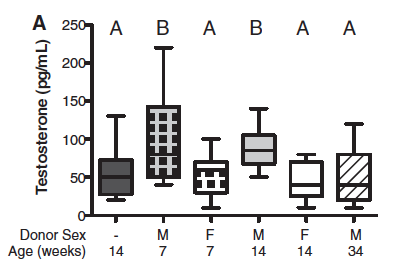
To follow up their findings correlating male microbiome with higher testosterone levels (Fig. 1C), they wanted to see if they could induce testosterone level changes through cecal transplantation from males to females. As shown in Figure 3A, females that received bacteria from age-matched males had significantly higher serum testosterone than control females, F --> F, and females with male donors 34 weeks old (letters A and B displayed above box-whisker plots indicate significance). It is peculiar to me that females with male donors 34 weeks old do not follow the same trend as females with male donors 7 and 14 weeks old. This could simply be because hormone production in general decreases with age, and so gut microbiome loses its affect over hormone production over time.
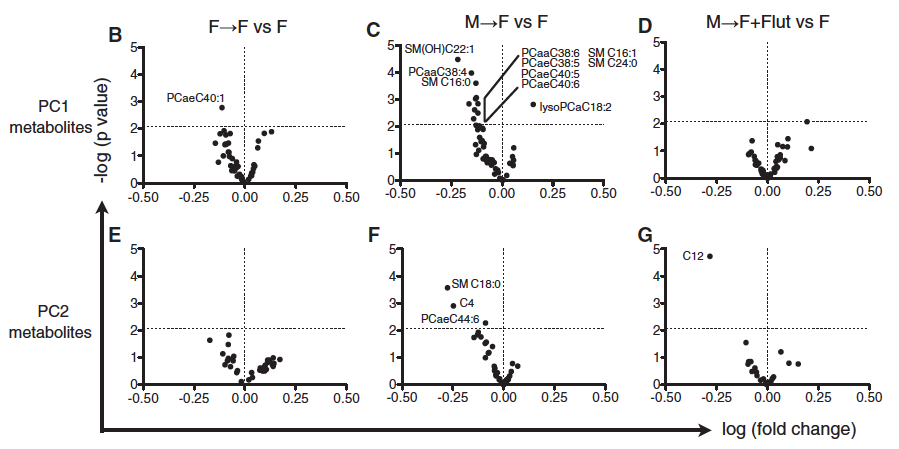
To follow up their findings of the bacterial genera that explained most of the variance between males and females (Fig. 1D), they used volcano plots to emphasize the metabolite bacteria that were significantly different after transplantation (which are labeled if they are above 2.08 p-value threshold), when comparing F --> F vs F, M --> F vs F, and M --> F + Flutamide vs F, using PC1 and PC2 metabolites from Fig. 1D (Fig. 3B-G). Interestingly, the researchers used flutamide to block androgen receptor (AR) signaling, and this attenuated all of the male microbiome-specific changes in female host metabolites. This suggests that the testosterone elevation caused by male microbiome transfer is critical for the generation of down-stream metabolomic phenotypes. It appears that blocking androgen receptors yields similar results as GF conditions. Perhaps it is the case that microbiome self-sustains by inducing a hormone like testosterone that selects for a particular composition of bacteria, facilitating a positive feedback loop that is evolutionarily advantageous for the microbiome. However, this does not answer the question of whether or not an organism can actively “correct” a microbiome after a disturbance such as transplantation. If not, then these data would suggest that the mice would simply remain hormonally challenged after the microbiome was altered.
Figure 4 – Gavage of female NOD pups with male NOD-derived intestinal microbiome results in T1D protection, decreased insulitis severity, and decreased insulin Aab titer by an androgen-dependent mechanism.
Because transfer of male microbiota altered recipient female hormonal and metabolic profiles, researchers wanted to know if these changes also altered the course of autoimmunity. This is relevant since we know that autoimmunity tends to have a strong sex bias. Figure 4A shows us the cumulative % T1D over the course of 35 weeks, and M-->F recipients have a significantly lower cumulative % T1D than control F and F-->F (Fig. 4A: unmanipulated NOD females – black; F-->F – red; M-->F – blue). More specifically, M-->F recipients were protected from invasive insulitis relative to both unmanipulated and F-->F recipients as shown in Figure 4B (Score 0-4 indicate severity of insulitis). In addition, M-->F recipients had significantly less autoantibodies (virtually none) against insulin in serum in comparison to unmanipulated F and F-->F (Fig. 4C). Insulitis and autoantibodies against insulin are both indicative of onset of T1D, and females receiving microbiota from males seem to avoid the T1D fate compared to other groups. These data demonstrate that male microbiome transfer to females drives testosterone-dependent attenuation of autoimmune phenotypes and protection from T1D.
The researchers wanted to see next if T1D-causing T cells was altered by these microbiome manipulations. They isolated splenic T cells from unmanipulated NOD mice, M-->F recipient mice, and M-->F+Flutamide recipient mice, and transferred to NOD.SCID mice through intravenous injection (Fig. 4D: unmanipulated females – black; M-->F – blue; M-->F+Flutamide – orange). The data show that it took M-->F recipient mice about 5-6 weeks longer to confer 100% cumulative T1D compared to unmanipulated F and M-->F+Flutamide recipients. Therefore, the male-specific microbiome seemed to delay the disease-fate, but did not avoid it completely. These data collectively demonstrate that male-specific microbiome protects from autoimmunity with testosterone as a key mediator. This makes me wonder, however, if testosterone induction alone could delay the onset of T1D without male-specific microbiome. We could use an AR agonist to see if this would alter the microbiome as suggested in Fig. 3D and3G, and, more importantly, if this agonist alone could alter the disease fate of the NOD mice.
Main Points
The take home message of this article is that genetics alone cannot explain the trends in T1D incidences, and studying the gut microbiome can explain sexual dimorphism and environmental influences of T1D. The gut microbiome is implicated to drive hormonal changes that ultimately plays a crucial role in autoimmunity. Thus far the data is correlational, and I am not completely convinced that the microbiome plays such a huge role in autoimmunity; however, the data showing differences in bacterial composition mostly based on sex differences is amazing to reconcile. Regardless, it would be interesting to see if normal mice can regulate their hormone levels in GF conditions, or if this phenomenon is unique to NOD mice. I would also want to see the effects of testosterone alone, since it is the “key mediator” in autoimmune protection. In general, I think the title is an overstatement and a strong claim to make. Furthermore, I am very surprised that the researchers did not characterize the T1D-causing T cells to note any phenotypic changes. I would like to have seen flow cytometry, western blots, or qPCR, just to see if there was upregulation of co-stimulatory molecules or any notable variations of the T cells between the 3 groups they used in Figure 4. I would not expect there to be T cells differences in which the majority of the variance could be explained by sex differences. T cell phenotype differences pre- and post-gavage would be nice to see too, and perhaps a functional assay to show if T cells are responding differently to self-antigen as a result of changes in microbiome or changes in hormones. I suspect that T1D is marked by Type 2 differentiation of T cells since T1D is characterized by autoantibodies against insulin. With microbiome transplantation, the foreign microbiota could induce Type 1 differentiation of T cells which would suppress antibody production to a certain extent and would at least partially explain the delay in T1D for M-->F recipients, and would otherwords not necessarily be detected by the IgG quantification in Fig. 2B which they used to check for induction of an immune response upon transplantation. We could explore this simply by analyzing the cytokine profile of the NOD mice (INF-gamma versus IL-4). Overall, though, I think this research was insightful and has opened doors for new questions to be explored and new research to be conducted.
Genomics Page
Biology Home Page
© Copyright 2014 Department of Biology, Davidson College, Davidson, NC 28035