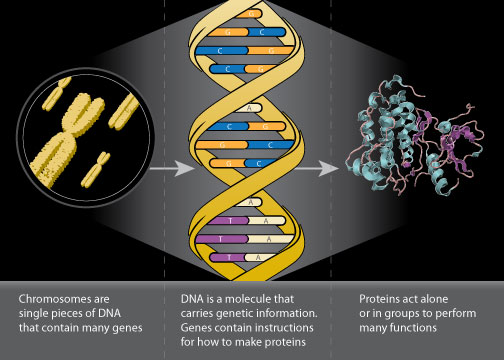
From Chromosomes to Proteins. Image from National Cancer Institute
This web page was produced as an assignment for an undergraduate course at Davidson College.

From Chromosomes to Proteins. Image from National Cancer Institute
Consider how a fresh biology student might respond when asked to define the protein content in a human body. Thinking back to their first intro bio course, the student might respond with the adage One gene, one enzyme. That is to say for every gene in the human genome, a single polypeptide is formed and shaped into an enzyme. While not accurate, our novice biologist is on the right track towards describing the expansive field of proteomics.
A proteome is the total set of expressed proteins contained in a cell at a given moment. This definition accounts for the vast potential of proteins that can occur within a single cell as well as whole organisms. Through alternate splicing genes can produce multiple variants of the same protein while posttranscriptional modification can alter the function of a single protein in variety of ways. Furthermore cells express different proteins through their various life cycles, while tissues are known for producing proteins unique to the tissue type. The vast data generated from studying the proteomes of a cell, let alone a whole organism makes the study of proteomes, or proteomics a dynamic and expansive field. Proteomics is the study, indexing, and documenting of proteomes in order to determine the responsible elements for key biological functions.
Many biological processes can be controlled by posttranscriptional modification of proteins and alternate splicing. Measuring and determining the proteome of cells and organisms can offer hints towards how basic biological systems operate and are controlled.
In addition, proteomics can assist in identifying subtle biomarkers, or indicators of a particular biological state. Biomarker discovery through proteomics research can identify the onset of diseases that are difficult to recognize. One application of clinical proteomics involves the detection of cancer through unique biomarkers found in blood plasma.
Gel-based proteomics are a common method to both isolate and quantify proteins in a sample. Using a two-dimensional gel electrophoresis, proteins are identified through banding patterns. In cases of samples with many proteins, chromatography and electrophoresis based techniques can be used to identify specific proteins.
Mass spectrometry is used in proteomics in order to identify individual proteins. Using mass spec, the exact molecular composition of an individual protein can be determined and used in order to determine the peptide sequence of the identified protein. While the technology is powerful, complex mixtures of protein can hinder the use of mass spec in determining individual proteins.
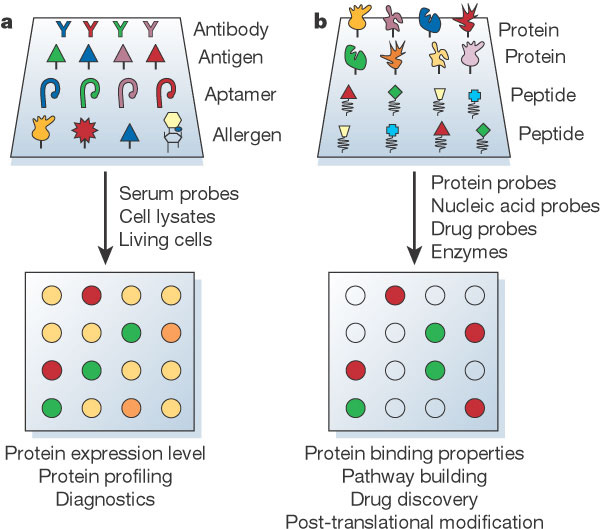
Analytical protein microarray. Image from Nature.
Protein microarrays are another versatile proteomics tool capable of identifying and measuring proteins. Different microarrays are capable of detecting a wide range of proteins and data pertaining to protein function and expression. Chips can monitor the expression level of a protein and also can be used to identify critical biomarkers.
Proteomics is a critical area of research in the post-genomic era based on the potential applications of the field. Research into clinical proteomics can assist with the identification of the onset of diseases, not limited to cancer. Continued improvements in proteomic related technology will usher in a new revolution for biology similar to the developments that led to the massive progress in the field of genomics.
Chevalier, F. 2010. Highlights on the capacities of Gel-based proteomics. Proteome Science 8:23.
Office of Cancer Clinical Proteomics Research [NCI] National Cancer Institute. Accessed 2014 Jan 25.
Phi. E. et al. 2003. Protein analysis on a proteomic scale. Nature 422.
Andrew Liu's Homepage
Genomics Page
Biology Home Page
© Copyright 2014 Department of Biology, Davidson College, Davidson, NC 28035