This
web page was produced as an assignment for an undergraduate course at
Davidson College
Back to homepage
Killifish Adaptation in Response to Complex
Pollution

What's
the research project?
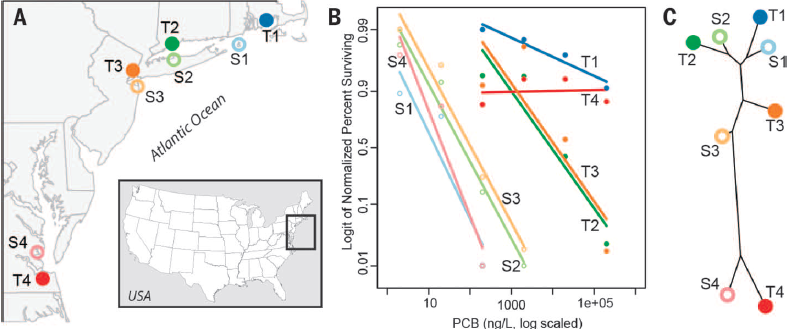
Figure 1.
A. The geographic locations of their Tolerant and Susceptible killifish
populations. C. A phylogenetic tree created by examining SNP
frequencies, showing that geographic location remains the greatest
determinant of genetic similarity between killifish populations. B.
Larval survival of susceptible and tolerant subpopulations when
introduced to increasing concentrations of PCB 126, a toxic pollutant
impacting the AHR pathway. Killifish were kept in identical conditions
for two generations preceding hatching these larvae, showing that genes,
not environment, determines larval resistance to PCB 126. (Replicated
Reid et al. 2016)
Is this hypothesis testing or discovery science?
Reid et al. used a combination of hypothesis testing and
discovery science: while they set out with the intention of determining if
tolerant populations share adaptations, they analyzed mass amounts of
killifish genetic and transcriptomic data to find candidate genes.
What genomic technology was used in the project?
The researchers fully sequences 43 to 50 individuals from
each of the eight sample populations, though the intensity varied: T1 and
S1 genomes were sequenced an average of 7x/individual, and the others were
only 0.6x/individual. Reid et al. split the sequenced genome into 5-kb
chunks, and searched for regions that showed less than 0.1% chance of
having evolved randomly. They found strong signals of recent convergent
evolution in the pollution-tolerant populations, particularly in genes
involved in the aryl hydrocarbon receptor signaling pathway (AHR).
Fig 2A. The difference between allele frequencies (top) and
nucleotide diversity (bottom) between
tolerant and susceptible population pairs. Areas outside the red
dashed lines indicate outliers, including the five AHR-implicated genes
indicated at the top of the figure. (Replicated
Reid et al. 2016)
They also RNA-seq data to determine which gene products differed between
the tolerant and susceptible populations. Reid et al. found that 70 genes
regulated by the AHR pathway were up-regulated in susceptible fish
populations introduced to polychlorinated biphenyl , but not in tolerant
populations. This is also indicative of convergent natural selection to
survive human pollution as two AHR-interfering pollutants (halogenated
aromatic hydrocarbons (HAHs), and polycyclic aromatic hydrocarbons (PAHs))
dominated at tolerant sites.
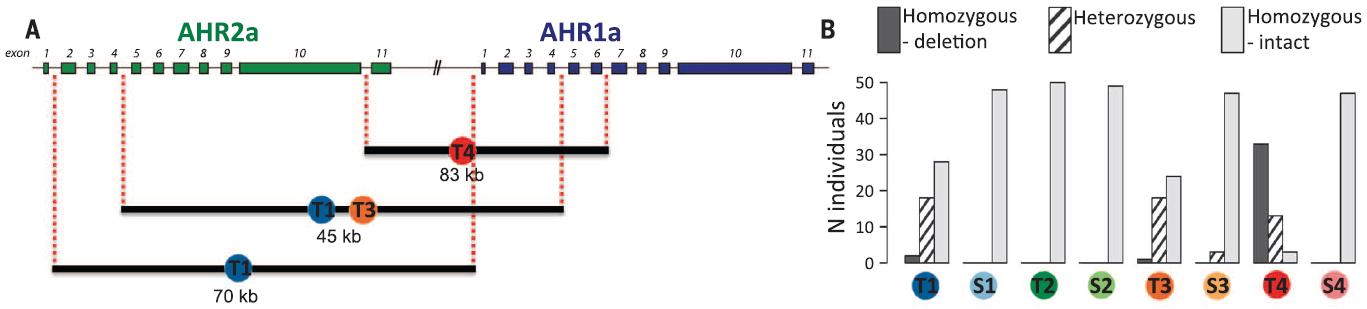
Fig 3A. Reid et al.'s gene model
of AHR pathway-related genes, with black bars indicating regions deleted
from indicated tolerant populations. 3B. The number of individuals found
to have homozygous deletions (black), heterozygous deletions (striped),
and no deletions (grey). Notice that S3 is the only susceptible population
to display a deletion genotype, and T2 is the only tolerant population to
display no deletion genotype. (Replicated
Reid et al. 2016)
The researchers found that tolerant fish had common and
population-specific adaptations. For example, in sites 1, 2, and 3 a
transcriptional target of AHR, the gene CYP1A showed consistent
duplication. However, site 4 was
mainly contaminated with PAHs and lacked this duplication. This
indicates that the tolerant fish are not only evolving tolerance to
complex chemical pollutants, but compensatory measures to survive evolving
that tolerance.
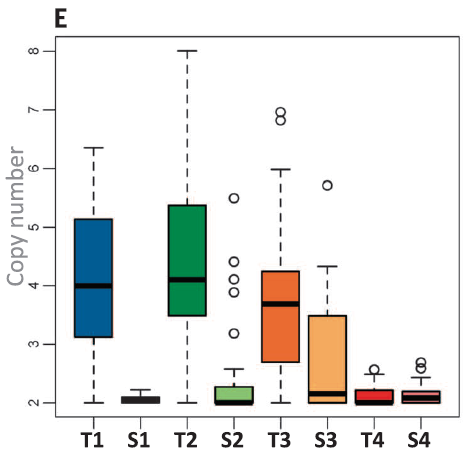
Fig 3E. T1, T2, and T3 show duplication of the CPY1A gene. (Replicated
Reid et al. 2016)
Take Home Message
The main takeaway from this
paper was that while natural selection targeted the aryl hydrocarbon
receptor signaling pathway (AHR) in all populations, the mechanism
varied between populations. Additionally, mechanisms varied based on
the chemical makeup of the pollutants affecting the region, suggesting
that killifish populations evolving complex adaptations in response
to unique and complex chemical mixtures. (Reid et al. 2016)
Additionally, AHR
interacts with a variety of signaling pathways, from immune
signaling to cell cycle regulation. A signaling pathway so integral to
so many physiological functions can't be tamped down without side
effects, and killifish may be evolving compensatory adaptations to
mitigate the effects of interfering with AHR while evolving the
pollution tolerance that's knocking it down.
Evaluation
On one hand, this research
is encouraging as it suggests species are capable of evolving and
therefore surviving human-caused
environmental degradation. While geographic T-S pairs
remained more genetically similar than any other pairing, tolerant
populations showed analogous patterns of adaptation facing complex
mixtures of pollution. However, strong selection pressure, wherein
unfit individuals are incapable of survival and or reproduction,
decreases genetic variation in a population. In fact, all tolerant
populations showed a smaller effective population size, meaning that
there is less variation per fish a worrying development in terms of
long-term species survival in a quickly changing world. Reid et al end
their paper by reminding the reader that killifish started out as a
widespread and genetically diverse species, and so were an ideal
candidate for studying adaptation in a variety of polluted habitats.
Being widespread and abundant puts killifish at little risk of
extinction regardless of their capacity to evolve.
Nevertheless, it's important to keep in mind the fact that most
species are rare. In my opinion choosing killifish makes this paper a
good precursor for genomic studies of evolution in endemic species
rather than a conclusive answer to the question of if threatened
organisms will be capable of adapting to humanity's destructive effect
on their habitats.
Bibliography
Reid, Noah M. et al.
2016. The genomic landscape of rapid repeated evolutionary adaptation
to toxic pollution in wild fish.
Science [Internet]. [cited 5
Feb 2017];354(6317) : 1305. Available from
http://science.sciencemag.org/content/354/6317/1305
Genomics
Page
Biology Home Page
Email Questions or Comments:
jaburtonakright@davidson.edu
© Copyright 2017 Department of Biology,
Davidson College, Davidson, NC 28035

