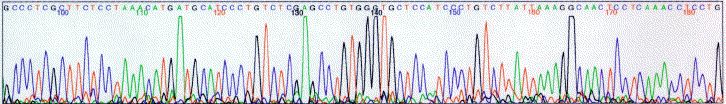
Background
In lecture, we have discused the utility of DNA sequence comparisons in determining the phylogeny of organisms. Nucleic acid sequence similarity is an indication of organism relatedness. In addition, sequence analysis can be useful in identifying microbes. It now is quite simple to isolate DNA from an unknown organism, sequence the 16S rRNA gene and determine the identity of that organism. Not only can such a sequence analysis help to elucidate the identity of an unknown microbe, but it also can provide important information about the phylogeny of the organism and maybe provide additional clues regarding the physiology of the organism.
The most commonly used method for obtaining rRNA sequences from an organism involves the use of the polymerase chain reaction (PCR) to amplify rRNA gene sequences. In the polymerase chain reaction, a pair of short synthetic DNAs that can base pair with opposite ends of the gene of interest (and on opposite strands) are mixed with a DNA sample from the organism of interest along with dNTPs and a DNA polymerase. After heating to make the sample DNA single-stranded, the synthetic DNAs, called primers, can base pair with the sample DNA and be used as primers by the DNA polymerase. After one such reaction, if you began with one copy of your gene of interest, you will now have two. If the reaction is repeated, you will have four, and so on. In general, the reaction is repeated twenty to thirty times. For organism identification, the primers used are derived from opposite ends of the 16S rRNA gene. Because the sequence of the 16S rRNA gene is highly conserved in many different organisms, investigators use "universal primers" that can base pair with any 16S rRNA sequence well enough to allow PCR amplification.
Once the gene of interest has been amplified, it can be sequenced. DNA
sequencing involves the use of dideoxynucleotide triphosphates (ddNTPs), which
also are referred to as chain terminators. Unlike deoxynucleotide triphosphates,
these molecules lack an -OH group on the 3' carbon atom of the ribose sugar.
In automated sequencing, each specific ddNTP (ddATP, ddTTP, ddCTP, ddGTP) is
labeled with a fluorescent dye. As the products of the sequencing reaction
are separated by electrophoresis, a laser "reads" the order of the fluorescent
dyes. This information, then, is transformed into an easy-to-read printout,
such as the one shown here.

For more information about dideoxy sequencing, click here.
So, now that you have sequenced the 16SrRNA gene of an organism, what
can you do with it? Most likely, you want to compare it to other
16S rRNA sequences. Such a sequence comparison will allow you to
identify your organism or, perhaps, reveal that you have isolated a previously
uncharacterized organism (maybe the elusive Davidsonnus wildcatii
bacterium!). Luckily, you don't have to pour over thousands of pages
of sequence data to make such a comparison. Virtually all published
DNA sequences are deposited in a publicly available database known as GenBank.
This sequence database is maintained by the National Center for Biotechnology
Information (NCBI), which is part of the NIH. The European Molecular
Biology Laboratory (EMBL) and the DNA DataBank of Japan (DDB) also maintian
DNA databases. These sites contain sequence data from thousands of
species - and the amount of information is growing at a phenomenal rate.
Software at these sites is available to perform sequence comparisons between
your sequence of interest and the entire database.
Assignment
Each of you will receive an email that contains a DNA sequence. Your job, then, is to determine from what species this sequence was obtained. After determining the identity of your mystery microbe, write a short paper (4 page maximum, double-spaced) that describes your microbe. This paper should not be merely a textbook description. Rather, uncover some interesting or unique feature of your microbe. Each paper must include at least one reference from a primary research paper. The papers will be due by 9:30 AM on Monday, February 4, 2013. In addition, each of you will give a 5 minute talk about your microbe during the laboratory periods on Feb. 5 or Feb. 6, 2013.
When you receive your email, copy the DNA sequence. Then, go to the following URL:
http://www.ncbi.nlm.nih.gov/blast/
In the 'Basic BLAST' section , click on 'nucleotide blast'
You should see the following page:
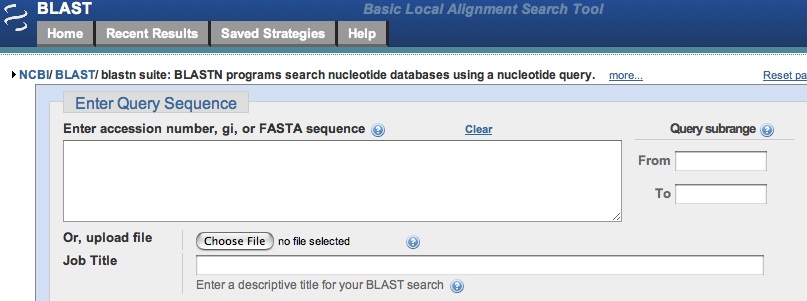
Paste your sequence into the box marked "Enter Query Sequence ". For "Database,"
select "Nucleotide collection". Hit <BLAST>. The following screens should
appear. Note: it may take as long as
ten minutes for your search to be completed, especially if you are conducting
your search during the day. When your search has been completed, you will see a screen like this one:
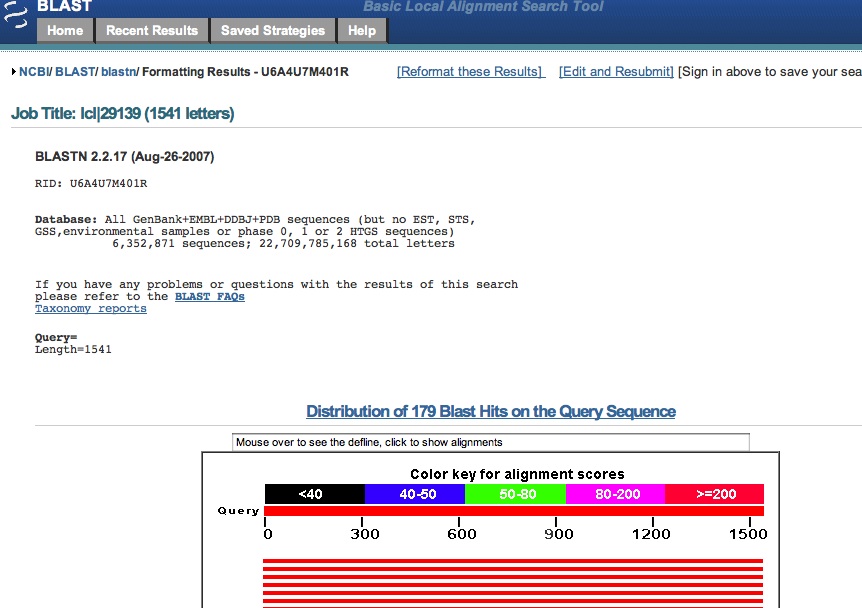
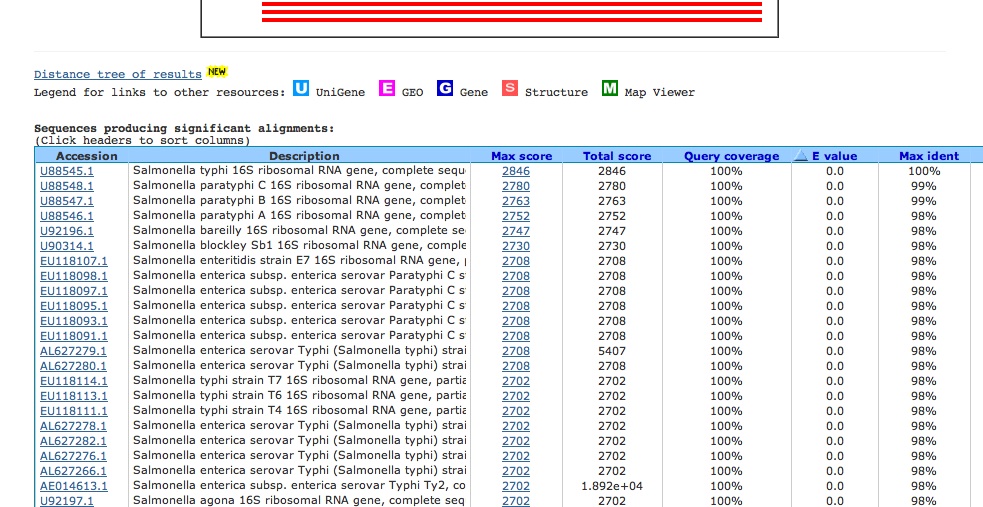
The "Distribution of Blast Hits" panel shows how similar the matching sequences are to your input sequence. As the color key indicates, red indicates high similarity, while black indicates relatively low similarity. The "Sequences producing significant alignments" panel provides you with a list of sequences in the GenBank database that display some similarity to your sequence. They are arranged from most similar to least similar. The "Score" represents a numerical assessment of the similarity two sequences show. The "E value" is a measure of how likely the identified similarity is simply due to chance. The greater the E value is, the less likely it is that the observed similarity is significant. The identification number preceding each entry is a link to more in depth information about that particular sequence, including citation information and the actual DNA sequence. This information should allow you to determine the identity of your mystery microbe and allow you to begin researching this organism.
This exercise has been adapted from "Identification of bacterial unknowns by rRNA sequence similarity," Richard J. Roller (Department of Microbiology, Univeristy of Iowa) author. Licensed for use, ASM MicrobeLibrary