
A large number of yeast biologists collaborated to produce a unique collection of knockout deletion yeast strains. The collaborative team systematically deleted each gene in the yeast genome. When completed, they had about 6,200 mutant strains each with a different gene deleted. However, rather than simply deleting the gene, they inserted a unique sequence of DNA for each strain and which has been called a "barcode". The analogy to barcodes found on packaging and used for checking out is a good one since a barcode should be a unique identifier for each item. For the yeast barcode deletions, the unique identifier is composed of two unique DNA sequences called "uptag" and "downtag" (figure 1). Homologous recombination was used to replace each gene with a unique barcode insert so that each strain had one gene deleted and replaced with its sequence-specific barcode insert.

Figure 1. Prototype of a barcode insert used to replace genes in the yeast genome. The two arrows represent PCR primers, the series of N's represent the unique barcode sequences and the marker gene permits selection of cells that contain the insert. The uptag consists of the 20 nucleotides on the left side and the downtag consists of the 20 nucleotides on the right. The red PCR primer sequence would be the same in every barcode insert as would the green PCR primer sequence.
Now that each strain has its gene deleted, the challenge was to determine the functional consequences of all 6,200 deletions. They could have studied each strain individually but they wanted to devise a high throughput method. To screen the mutants efficiently, they put about 500 mutants into a common flask and then asked the question, "How will each strain survive in this competitive environment?" To answer their question, they first used the two PCR primers to amplify all 500 barcode inserts from a mixture of genomic DNA extracted from an aliquot of the 500 strains. These PCR products were tagged with a red dye. The cells were allowed to grow over an extended period of time. During this time, strains with deleted genes needed to maintain normal growth rate would constitute a reduced percentage in the growing population. At various times, aliquots of cells were removed and the genomic DNA was extracted and amplified using the same two PCR primers. These PCR products were tagged with a green dye (figure 2).
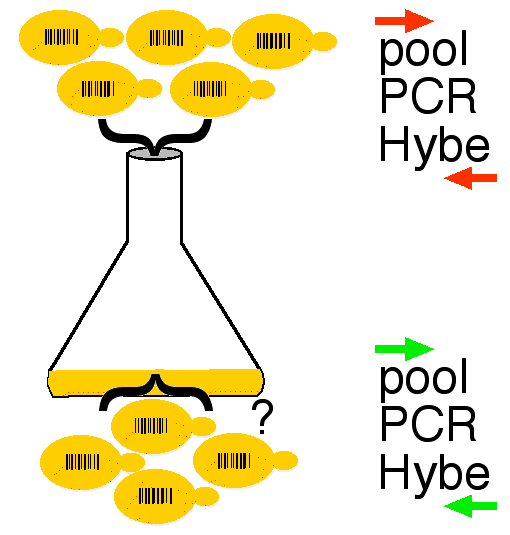
Figure 2. A mixture of different barcode-deletion strains were placed in one flask and allowed to grow. Genomic DNA from an aliquot of this initial population of cells was used as template for PCR using the two primer sequences shared by each of the deletion strains. These PCR products were labeled red. After a period of time, an aliquot of cells was again used for template of PCR using the same two PCR primers. These PCR products were colored green.
Barcode yeast experiments require a unique DNA microarray. Instead of spotting the coding DNA for each gene, the uptag and downtag for each deletion was spotted onto the glass. The two PCR products (red from time zero and green from a later time) were mixed together and incubated with the DNA microarray. For each strain there could be three possible lableing patterns for its associated uptag and downtag (figure 3).
1) If a strain grew at a rate comperable to most of the other strains, then it would appear as a yellow spot since its proportion of the initial and final total population was the same.
2) If a strain grew slower than most other cells, then its uptag and downtag spots would appear red since this strain was present at a higher concentration in the initial population than in the later population.
3) If a strain grew faster than most other cells, then its uptag and downtag spots would appear more green than yellow since its increased rate of growth resulted in an increased representation of the total population.

Figure 3. DNA microarray showing yellow, red and green spots for cells that maintained the most common growth rate, decreased growth rate or increased growth rate, respectively. The colors in this figure oversimplify reality to highlight the three extreme outcomes.
By designing two barcodes for each strain, the investigators built in a control for each strain. If a strain's growth rate were truely affected by the deletion, then both the uptag and downtag spots should exhibit similar results. If they differred, you would be advised to ignore that strain in this experiment and try again later.
© Copyright 2002 Department of Biology, Davidson College,
Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu