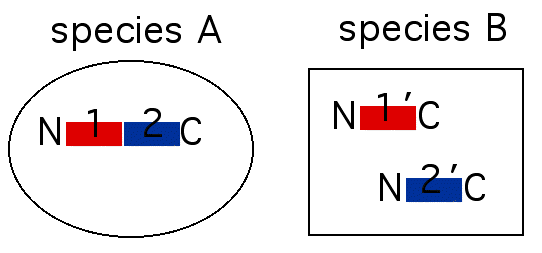
About 40% of all genes have no known function. To reduce the number of "unknown genes", it is possible to perform a sequence analysis looking for functional domains. To perform this sequence analysis, it is necessary to compare the genomes of different species. In species A, a protein exists that has two functional domains (labeled 1 and 2 in figure 1). In species B, there may be two orthologs of the two domains but in species B, the orthologs are found in separate genes (labeled 1' and 2' in figure 1).
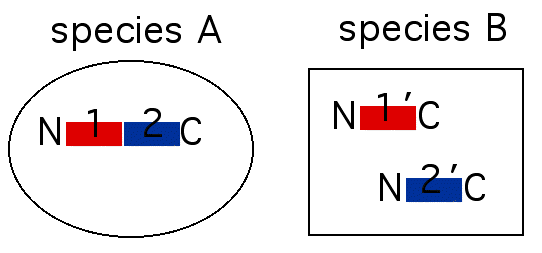
Figure 1. Two speices with conserved functions. Species A performs a particular task using two domains in a single protein. Species B performs the same task but utilizes two genes encoding similar domains.
Let's consider an hypothetical example. Kinases must have at least two domains to perform their task of phosphorylating a substrate. One domain binds the substrate and the other binds ATP and transfers the terminal phosphate onto the substrate. It is easy to image that in species A, this task could be performed by a single protein while in species B, two genes form a heterodimer to accomplish the same task.
Using this type of conservation of sequence but divergence of gene number and size, investigators have been able to mine genome sequences to determine the functional roles of previously "unknown" genes.
© Copyright 2002 Department of Biology, Davidson College,
Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu