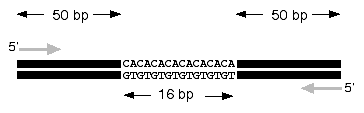
Microsatellites (sometimes referred to as a variable number of tandem repeats or VNTRs) are short segments of DNA that have a repeated sequence such as CACACACA, and they tend to occur in non-coding DNA. In some microsatellites, the repeated unit (e.g. CA) may occur four times, in others it may be seven, or two, or thirty. In diploid organisms such as elephants, each individual animal will have two copies of any particular microsatellite segment. For example, a father might have a genotype of 12 repeats and 19 repeats, a mother might have 18 repeats and 15 repeats while their first born might have repeats of 12 and 15. On rare occasions, microsatellites can cause the DNA polymerase to make an extra copy of CA similar to the way we find it difficult to say łtoy boat˛ several times in a row with consistent accuracy. If an individualąs DNA polymerase adds to the repeated sequence, then this slightly larger version can be passed on to offspring who will usually replicate it accurately. Over time, as animals in a population breed, they will recombine their microsatellites during sexual reproduction and the population will maintain a variety of microsatellites that is characteristic for that population and distinct from other populations which do not interbreed.
The most common way to detect microsatellites is to design PCR primers that are unique to one locus in the genome and that base pair on either side of the repeated portion (figure 1). Therefore, a single pair of PCR primers will work for every individual in the species and produce different sized products for each of the different length microsatellites.
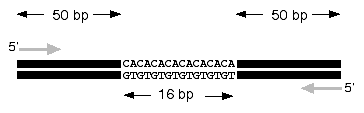
Figure 1. Detecting microsatellites from genomic DNA. Two PCR primers (forward and reverse gray arrows) are designed to flank the microsatellite region. If there were zero repeats, the PCR product would be 100 bp in length. Therefore, by determining the size of each PCR product (in this case 116 bp), you can calculate how many CA repeats are present in each microsatellite (8 CA repeats in this example).
The PCR products are then separated by either gel electrophoresis or capillary electrophoresis. Either way, the investigator can determine the size of the PCR product and thus how many times the dinucleotide "CA" was repeated for each allele (figure 2). It would be nice if microsatellite data produced only two bands but often there are minor bands in addition to the major bands; they are called stutter bands and they usually differ from the major bands by two nucleotides.
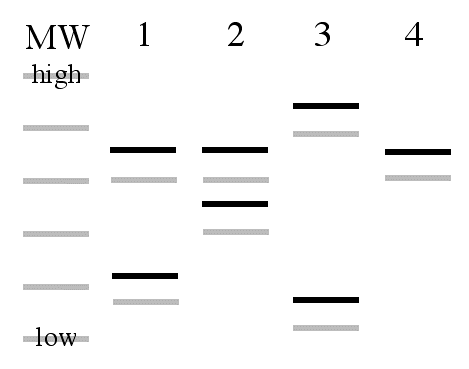
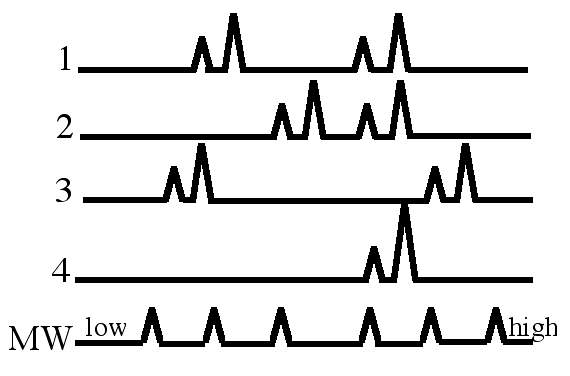
Figure 2. Stylized examples of microsatellite data. Left half: four sets of data were produced by gel electrophoresis and so you can see the major (black) and stutter (gray) bands. MW; molecular weight standards. Right half: These data were produced by analysis on an automated capillary electrophoresis-based DNA sequencer. The data are line graphs with the location of each peak on the X-axis representing a different sized PCR product and the height of each peak indicates the amount of PCR product. The major bands produce higher peaks than the stutter peaks.
Thought Questions:
1) Why do you think microsatellites with repeating units of two nucleotides are usually located in non-coding DNA?
2) Why might a stutter band differ from a major band by two nucleotides instead of one?
3) In the figure above, one animal only has a single band. How can this be?
4) Compare the two forms of data in figure 2 above? Are the data identical or not?
© Copyright 2001 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu