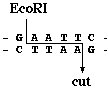
RFLP (often pronounced "rif lip", as if it were a word) is a method used by molecular biologists to follow a particular sequence of DNA as it is passed on to other cells. RFLPs can be used in many different settings to accomplish different objectives. RFLPs can be used in paternity cases or criminal cases to determine the source of a DNA sample. RFLPs can be used determine the disease status of an individual. RFLPs can be used to measure recombination rates which can lead to a genetic map with the distance between RFLP loci measured in centiMorgans.
On this web page, you can see how RFLPs are produced and then three examples of applying RFLP analysis: paternity, disease status, and genetic mapping.
Each organism inherits its DNA from its parents. Since DNA is replicated with each generation, any given sequence can be passed on to the next generation. An RFLP is a sequence of DNA that has a restriction site on each end with a "target" sequence in between. A target sequence is any segment of DNA that bind to a probe by forming complementary base pairs. A probe is a sequence of single-stranded DNA that has been tagged with radioactivity or an enzyme so that the probe can be detected. When a probe base pairs to its target, the investigator can detect this binding and know where the target sequence is since the probe is detectable. RFLP produces a series of bands when a Southern blot is performed with a particular combination of restriction enzyme and probe sequence.
For example, let's follow a particular RFLP that is defined by the restriction enzyme EcoR I and the target sequence of 20 bases GCATGCATGCATGCATGCAT. EcoR I binds to its recognition seuqence GAATTC and cuts the double-stranded DNA as shown:
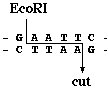
In the segement of DNA shown below, you can see the elements of an RFLP; a target sequence flanked by a pair of restriction sites. When this segment of DNA is cut by EcoR I, three restriction fragments are produced, but only one contains the target sequence which can be bound by the complementary probe sequence (purple).
Let's look at two people and the segments of DNA they carry that contain this RFLP (for clarity, we will only see one of the two stands of DNA). Since Jack and Jill are both diploid organisms, they have two copies of this RFLP. When we examine one copy from Jack and one copy from Jill, we see that they are identical:
Jack 1: -GAATTC---(8.2 kb)---GCATGCATGCATGCATGCAT---(4.2 kb)---GAATTC-
Jill 1: -GAATTC---(8.2 kb)---GCATGCATGCATGCATGCAT---(4.2 kb)---GAATTC-
When we examine their second copies of this RFLP, we see that they are not identical. Jack 2 lacks an EcoR I restriction site that Jill has 1.2 kb upstream of the target sequence (difference in italics).
Jack 2: -GAATTC--(1.8 kb)-CCCTTT--(1.2 kb)--GCATGCATGCATGCATGCAT--(1.3
kb)-GAATTC-
Jill 2: -GAATTC--(1.8 kb)-GAATTC--(1.2 kb)--GCATGCATGCATGCATGCAT--(1.3
kb)-GAATTC-
Therefore, when Jack and Jill have their DNA subject to RFLP analysis, they will have one band in common and one band that does not match the other's in molecular weight:
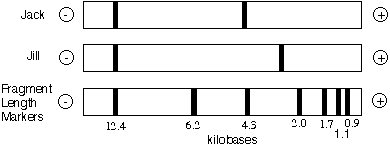
Let's use RFLP technology to determine if Jack is the father of Jill's child named Payle.
In this scenario, DNA was extracted from white blood cells from all three individuals and subjected to RFLP analysis. The results are shown below:
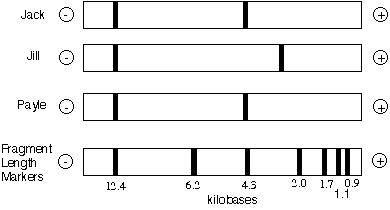
In this case, it appears that Jack could be the father, since Payle inherited the 12.4 kb fragment from Jill and the 4.3 fragment from Jack. However, it is possible that another man with similar RFLP pattern could be as well.To be certain, several more RFLP loci would be tested. It would be highly unlikely that two men (other than identical twins) would share multiple RFLP patterns and so paternity could be confirmed.
In a different scenario, DNA was extracted from white blood cells from all three individuals and subjected to RFLP analysis. The results are shown below:
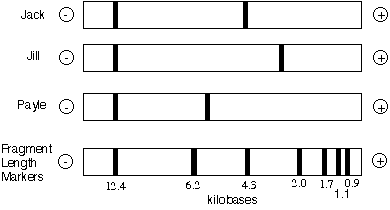
This time, it can be determined that Jack is NOT the father of Payle since Payle has a band of about 6 kb and Jack does not. Therefore, it is very probable that Payle's father is not Jack, though it is possible that Payle carries a new mutation at this locus and a different sized band was produced. What could you do as an investigator to be more certain that Jack was not the father of Payle?
In this example, we want to know if a person carries any cystic fibrosis (CF) alleles and if so, how many. Because CF is a recessive disease, anyonne with CF must be homozygous for disease alleles. From pedigree information, we can often determine who in this family is a carrier. However, if a couple comes to a genetic counselor, often an RFLP analysis is performed on the couple's DNA.
RFLPs are known for CF and so it would be easy to determine if a person were homozygous wild-type (wt), heterozygous "carrier", or homozygous disease alleles and thus have CF.
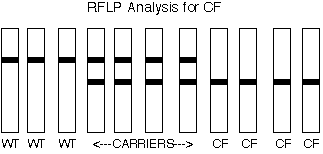
For couples expecting a child, it would be simple to test both parents and make a prediction about the eventual disease status of their fetus. For example, if both parents were homozygous wt, then all of their children would also be homozygous wt:
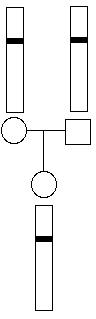
However, if both parents were heterozygous, they could have children with any of the three genotypes, though heterozygous children would be twice as likely as either of the homozygous genotypes.
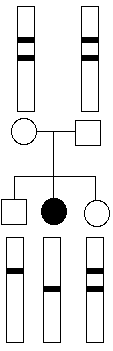
With increasing genomic sequence information, increasing numbers of genetic disease can be predicted from RFLP analyses.
To calculate the genetic distance between to loci, you need to be able to observe recombination. Traditionally, this was performed by observing phenotypes but with RFLP analysis, it is possible to measure the genetic distance between two RFLP loci whether they are a part of genes or not.
Let's look at a simple example in fruit flies. Two RFLP loci with two RFLP bands possible at each locus:
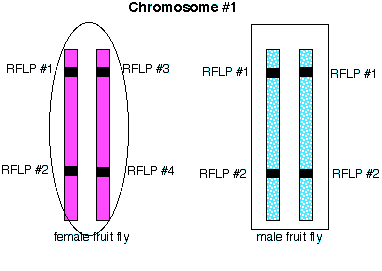
These loci are located on the same chromosome for the female (left) and the male (right). The upper locus can produce two different bands called 1 and 3. The lower locus can produce bands called 2 or 4. The male is homozygous for band 1 at the upper locus and 2 for the lower locus. The female is heterozygous at both loci. Thier RFLP banding patterns can be seen on the Southern blot below:
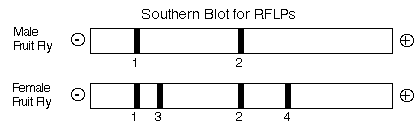
The male can only produce one type of gamete (1 and 2) but the female can produce four different gametes. Two of the possible four are called parental because they carry both RFLP bands from the same chromosome; 1 and 2 from the left chromosome or 3 and 4 from the right chromosome. The other two chromosomes are recombinant because recombination has occurred between the two loci and thus the RFLP bands are mixed so that 1 is now linked to 4 and 3 is linked to 2.

When these two flies mate, the frequency of the four possible progeny can be measured and from this information, the genetic distance between the two RFLP loci (upper and lower) can be determined.
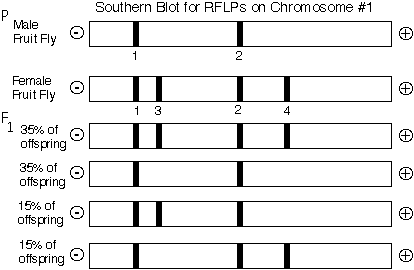
In this example, 70% of the progeny were produce from parental genotype eggs and 30% were produced by recombinant genotype eggs. Therefore, these two RFLP loci are 30 centiMorgans apart from each other.
© Copyright 2001 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu