
 Click
to see a larger version of this image.
Click
to see a larger version of this image. 2009 GCAT Microarray Workshops
Two Sessions; 20 people per session
Week of July 6 - 11
Morehouse College, Atlanta, GAThis will be the last GCAT Microarray Workshop
2009 Workshop #1 (20 participants)
2009 Workshop #2 (24 participants)
2009 Instructors
Todd Eckdahl, Laurie Heyer, Anne Rosenwald,
Consuelo Alvarez,
Charles Hauser, Malcolm Campbell, Edison Fowlks
Pre-Workshop Reading and Problem Set |
|---|
Highlights:
Free Software and Training
Interdisciplinary Team Participation Encouraged
Produce RNA; cDNA probes; Hybe; Wash; Scan; Analyze
Yeast full-genome microarrays
Computers Provided but You Can Bring Laptops
We will use Genisphere’s 3DNA method instead of direct incorporation. We give each participants a GCAT Flash Drives (1 GB each) which will allow us to deliver the workshop materials and for participants to archive all their data. We will collect all the drives during the scanning period and return them with all the tiff files the next morning in time for dry lab.
Participants will prep their own RNA. We will provide for them frozen yeast pellets and the RNA isolation kits. They will analyze the quality of their RNA by spectrophotometer only (i.e., will not run gels) and decide if they want to use their RNA or some prepared by the instructors prior to the workshop. Participant RNA isolation will take place on Day 2 for both workshops during their half day of wet lab.
Selection Criteria
Maximum Impact on African American, Hispanic, Native American, and Pacific Islander Students (e.g. Hawaii)
Previous Experience with Molecular Methods (e.g., pipetting, etc.)
OR
Mathematics/Computer Science Faculty (Math/CS + Biology teams welcomed)
Interested in Bringing Microarrays into Undergraduate Curriculum
Minimal Experience with Microarrays Preferred
Willingness to Learn and Collaborate with Other Faculty
Applied to previous Workshop but Not Accepted
(please include this information on your application)
Time of Application Submission
(used as tie breaker)
GCAT does not set quotas; we try to accomodate all qualified applicants
Purpose: Genome Consortium for Active Teaching (GCAT) offers hands-on workshops for faculty who conduct research with undergraduate students and/or teach undergraduate courses to learn about gene expression analysis via microarrays.
Workshop Organization:
Part 1 of the workshop
begins with data
analysis (see schedule below)
to introduce the microarray method and will cover data analysis using
public domain data from the literature and from GCAT. Participants
will learn public domain and open source MAGIC
Tool spot-finding and analysis software. The MIAME background
information and its use in microarray evaluation will be covered.
Participants will analyze data sets in short projects. Each of the
two workshops has room for 20 participants.
Part 2 will be a hybridization workshop which will involve hands-on preparation of fluorescently labeled probes for yeast expression microarrays, their hybridization, data acquisition and data analysis using the methods presented in part 1 of the workshop.
Part 3 concludes with data analysis of the microarrays you produced. This will also serve to reinforce what you learned about data analysis.
Where: Morehouse College, Atlanta, GA
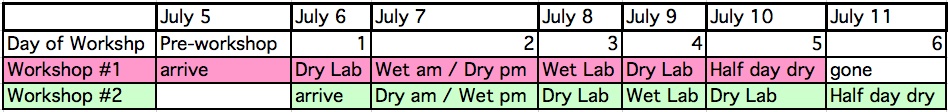
When: workshop #1: July 6 - 10 mid day; and workshop #2: 7 - 11, 2009
Costs and Scholarships: If funded by the NSF, this workshop will be completely free (housing, food, lab fees) for each participant with one catch. You pay for your airfare in advance but you will be reimbursed if you submit an action plan for implementing microarrays in your curriculum and submit receipts. For those with financial need for airfare in advance, you may petition to have this paid in advance.
Download Workshop Announcement |
Download Workshop Application |
|---|---|
Announcement
in PDF format (.pdf) |
Application
in PDF format (.pdf) |
Announcement
in RTF format (.rtf) |
Application
in RTF format (.rtf) |
Announcement
in Word format (.doc) |
Application
in Word format (.doc) |
Email Edison Fowlks or Malcolm Campbell with Questions
See photos from 2003 workshops
See photos from 2004 workshops
See photos from 2005 workshops
This material is based upon work supported by the National Science Foundation under Grant No. DBI-0627478. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
© Copyright 2009 Department of Biology, Davidson College,
Send comments, questions, and suggestions to: macampbell@davidson.edu