Spring 1999 Molecular Biology Exam #1 - Learning the Tools
There is no time limit on this test, though I have tried to design one that you should be able to complete within 2.5 hours, except for typing. You are not allowed to use your notes, or any books, nor are you allowed to discuss the test with anyone until Wednesday Feb. 3, 1999. EXAMS ARE DUE AT 10:30 ON WEDNESDAY, FEBRUARY 3. You may use a calculator and/or ruler. The answers to the questions must be typed on a separate sheet of paper unless the question specifically says to write the answer in the space provided. If you do not write your answers on the appropriate pages, I may not find them unless you have indicated where the answers are.
Please do not write or type your name on any page other than this cover page. Staple all your pages (INCLUDING THE TEST PAGES) together when finished with the exam.
Name (please print here):
Write out the full pledge and sign:
Here is the honor code
http://www.davidson.edu/student/redbook/honorgeneral.html#honorcode
"On my honor I have neither given nor received unauthorized information regarding this work, I have followed and will continue to observe all regulations regarding it, and I am unaware of any violation of the Honor Code by others."
How long did this exam take you to complete (excluding typing)?
Average for this test in 1999 was 85% (range 100% to 60%)
6 pts.
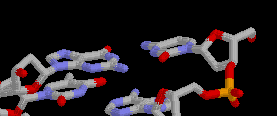
1. You have been asked by Kenneth Starr (no relation to Ringo) to identify this molecule. Of course you recognize it to be a piece of double stranded DNA and you even notice that the bases shown probably have 3 H-bonds between them.
a. After stating your name for the record, tell the
grand jury which base appears on the left and which one on the
right (typical partisan politics).
guanine on the left
cytosine on the right
b. As if this were not enough, now the "special
prosecutor" wants to know which base is on the 5' end and
which one is on the 3' end of each strand. Please tell him the
correct answer or you will be prosecuted.
cytosine on the 5 ' end
guanine on the 3 ' end

6 pts.
2. DNA replication can be a political hot potato(e). But there
are some very useful enzymes for us to use. Tell me which enzyme
we have used in lab and why we used it.
DNA ligase was used to clone the PCR product
into the pNoTA plasmid.
16 pts.
3. The following notes were found in the kitchen of a work study student and a certain teacher has been accused of leaking classified information to students who wanted to take his class but were obstructed from doing so. Please help clarify the situation by showing that this course is not so hard and telling the congress how to do these calculations:
a. Make a 500 mL solution that is 5 mM NaCl, 15% SDS
(v/v if the stock solution of SDS is 50% w/v), 0.01 M Tris-HCl,
pH 7.5. (FW NaCl = 58.44; FW Tris = 121; FW SDS = 288.4)
0.146 g NaCl + 150mL 50% SDS + 0.61 g Tris
Put about 200 mL of water in a flask, add the above and dissolve.
pH to 7.5 with HCl, then add water to 500 mL.
b. How many microliters (µL) would you need to
pipet if you wanted to deliver 750 ng of a dsDNA to a new tube?
The OD260 of the solution you are to pipet read 0.084.
(Remember the formula for conversion: 1 OD 260 unit
has a concentration of 50 µg/mL dsDNA.)
178.6 µL of this stock (which has
a concentration of 4.2 ng/µL).
c. What does OD 260 mean anyway??
This means that we measured the absorbance
of light (260 nm wavelength) of a solution. If this were a DNA
solution, we could determine the concentration of the DNA, as
in problem "b" above.
d. Make a 150 mL solution that is 1200 µM NaCl,
and 0.05 mM KOH (stock solution of KOH is 5 M ), and 0.1 M SDS
(start with the dry stuff).
0.01 g NaCl + 1.5 µL KOH + 4.32 g
SDS
Put about 100 mL of water in flask. Dissolve the above and then
bring the volume up to 150 mL with water.
18 pts.
4. Using the DNA sequencing gel below, please help us identify the owner of this DNA. The dideoxynucleotides that were used were loaded in the order of (left to right) C G A T.

a. Record for the senate the sequence of this DNA (not
the template DNA) from 5' to 3'. You should be able to read at
least 45 bases. Please record the first 45 you can read.
CC ATG TCC AAT CGG TCA GTC CGT CAT TGC TAC
GCG ATA TTA GCC C....
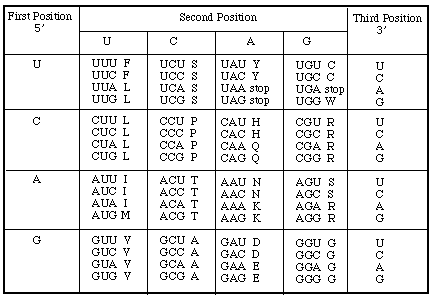
b. Look for the first start codon and translate as much
as you can using the codon table below. You must use the full
names of the amino acids.
methionine serine, asparagine, arginine,
serine, valine, arginine, histidine, cysteine, tyrosine, alanine,
isoleucine, leucine, alanine, ...proline....
6 pts.
5. If this piece of DNA were a cDNA, explain to those non-science
members of the House what is 5' to a start codon and downstream
of a stop codon. Also explain the function of these features.
5 ' untranslated region (UT)
3 ' UT
Both of these provide stability for the mRNA in the cytoplasm
and can influence translation too.
6 pts.
6. The House managers have determined that a certain protein
has been identified that was not where it should be. They claim
it should have been in the cytoplasm but was instead found located
in the Golgi body! (Gasp!) Explain to this one-minded group what
one would have to do to the gene of a person to convert a Golgi-localized
protein into a cytoplasmic protein.
To convert a Golgi-localized protein to
a cytoplasmic one, the gene sequence would have to be altered.
One would need to modify the encoding nucleotides of the first
exon so that the portion which encoded the signal sequence was
deleted. This would cause the protein to be translated on free
ribosomes and stay in the cytoplasm.
6 pts.
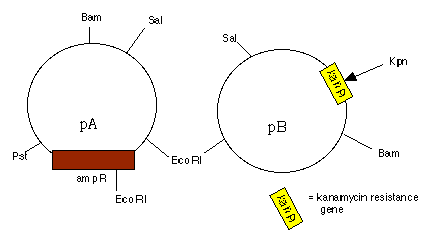
7. Tell me how you would clone the ampicillin resistance gene
from plasmid A into plasmid B (both of which are popular plasmids
in political circles, and completely sequenced) at the Kpn site
indicated by an arrow. To receive full credit, you must tell me
how you would know your cloning attempt was successful or not.
pA is 3.6 kb and pB is 3.0 kb in size.
I would use PCR with primers that amplified
the ampR gene but also contained Kpn linkers. The PCR product
and pB would be digested with Kpn, and then ligated together.
The ligation reaction would be transformed into E. coli cells
and plated on amp containing LB. If the cells grew, they must
have contained the ampR gene. I would then either test them on
Kan plates and look for cells that failed to grow (negative result)
or digest minipreped DNA with Kpn and look for the appropriate
banding pattern.
8 pts.
8. Let's make up a completely fictitious situation. Let's say a very famous person was involved in a sexual affair with someone half his (or her) age and knew it was wrong. But the younger adult wanted to take advantage of the situation and saved some evidence of the affair for later use. How would this younger person prove that she (or he) possessed a piece of clothing that was stained by the older adult? (Does everyone understand the scenario here? If not, call me to clarify what we are talking about.)
a. What method would you recommend that the younger
one (she or he) use and why?
RFLP analysis would be the most obvious
choice. DNA would be extracted from the cloth and then subjected
to analyses at several loci (different restriction enzyme and
probe combinations).
b. What controls would be necessary when performing
your recommended experiment?
One should test the two consenting adults'
DNA, as well as the technicians' DNA. A positive control (known
standards) for each locus would be nice too.
14 pts.
9. Let's imagine there is a genetic disease that forces a person to do dumb things every 4 months. And let's imagine that this gene is expressed in the big toe. Let's also imagine that people afflicted with this disease have very large big toes and the dumb-gene protein is easily purified. The next three questions do not have to have related answers so don't let "a" stop you from being creative with "b" or "c", for example.
a. Explain to me how you would go about deducing the
amino acid sequence of this dumb protein in the fastest way possible
(your country's next official dumb act is only a couple days away).
This gene has never been cloned before in any species.
Either sequence some of the protein and
design a probe to screen a genomic library, or use a mAb and an
expression library. If you do the latter, you would have to use
the resulting cDNA to screen a genomic library, so plan A seems
to be the better choice (one screening).
b. How would you determine what tissues, if any, the
mouse homologous protein is expressed in mice?
If you had a mAb, you could see if it bound
the mouse homolog. If you get no signal, then you need to determine
if mice make said dumb protein by screening a mouse library using
the human sequence as a probe. Eventually, you could perform an
RNA blot using RNA from a range of tissues to see where the gene
is transcribed. If the original mAb works, then you could do a
protein blot on a variety of mouse tissues. This mAb approach
would detect protein expression directly while the Northern blot
would be an indirect measurement.
c. How would you design a plasmid that would act as
an early warning system in humans when this gene is getting ready
to be transcribed again so you could prevent the next outbreak
of dumbness in humans with said disease?
I would put the human DUMB promoter upstream
of GFP. Then I would secretly get this construct into the muscles
of the big toe by injecting the DNA (not noticed by someone so
dumb) into the toe. THen I would examine the toes daily with a
UV lamp to find out the level of GFP expression.
4 pts.
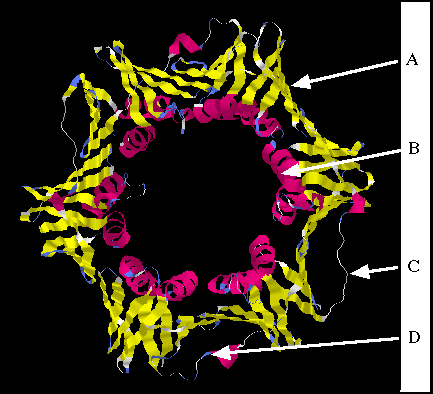
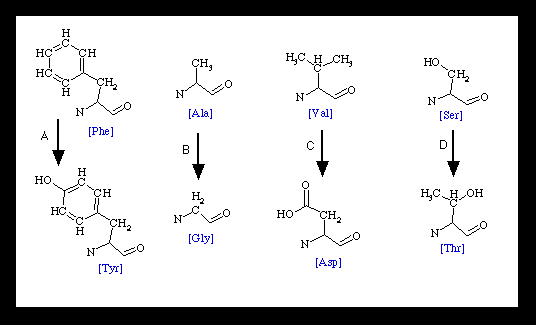
10. For the molecule depicted, tell me which amino acid substitution
(multiple choice from below) might cause the greatest functional
change and why you think so.
A seems to be the most problematic since
a hydrophobic R group has been changed to a hydrophillic one,
and one that could be phosphorylated even. This occurs within
a ß-pleated sheet. The substantial change in amino acid
at position C is not in an area of certain structure, so it does
not seem as critical.
10 pts.
11. Tell me how you could make a monoclonal antibody against
the mouse act-dumb protein and then find out if it bound to the
human homolog or not.
This would follow the traditional route
with the main exception that you cannot make a mAb against a mouse
protein in a mouse. You must use another organism. Hybridomas
would have to be screened to see which ones actually make antibodies
to the DUMB protein - you cannot guess which ones will.