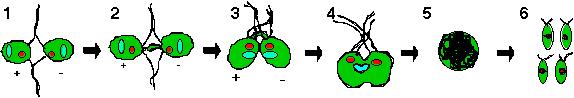
Chalamydomonas reinhardtii is a eukaryotic,
unicellular, biflagellated, haploid green alga that can be found in the
soil and from fresh water habitats (Goodenough, 1995). It is often touted
as the "green yeast" because it is flexible and suitable for carrying out
classical genetic studies. Many researchers have chosen Chlamydomonas
as a model organism to study signal transduction and transmembrane signalling
(Quarmby and Hartzell, 1994), flagellar structure (Hunnicutt et al.,
1990; Goodenough, 1985), flagellar interaction (Goodenough, 1989) and others.
Wild type Chlamydomonas spends most
of its life cycle as a vegetative cell. However, when subjected to nitrogen
deprivation and in the presence of light, Chlamydomonas can quickly
adapt to the new environment by differentiating into gametic cells capable
of sexual reproduction. Formation of these gametes is associated with alteration
of the cellís biochemistry and subcellular morphology, including expression
of several gamete-specific genes (Gromoff et al., 1993). Chlamydomonas
is heterothallic with two mating types called the mating type plus (mt+)
and mating type minus (mt-). These two mating types, though similar
in morphology, differ in their ultrastructure. The locus that codes for
the mating types behaves like a single gene, but it is found to have different
DNA sequences (Ferris et al., 1987).
The Chlamydomonas flagella have multiple
functions. They are involved in signal transduction, cell-substrate interactions,
gliding motility (Bloodgood, 1988) and play an important role in gamete
recognition during sexual reproduction. During mating, flagellar agglutination
occurs between the two mating types. Agglutination is a process where the
flagella from one Chlamydomonas mating type adheres to the flagella
of the opposite mating type (Figure 1-1). The agglutination process
is mediated by flagellar glycoproteins called agglutinins. Once the gametes
agglutinate, an intracellular signaling cascade triggers the rise of cAMP
within the gametes which initiates several mating responses (Pasquale and
Goodenough, 1987). First, a change in the morphology results in flagellar
tip activation. This is characterize by tip enlargement and accumulation
of fibrous proteins in specific regions near the terminal membrane (Figure
1-2; Harris, 1989). Next, autolysin is secreted by the gametes to facilitate
the shedding of cell walls so that fusion of the gametes can proceed. Activation
of the mating structures also occur. For the plus gamete, this involves
an extension of a fertilization tubule that is ultimately linked with the
minus mating structure (Figure 1-3). It has also been reported that cAMP
also enhances sexual agglutinability of the Chlamydomonas flagellar
by the dynamic recruitment of agglutinins to the flagellar surface (Goodenough,
1989). All of these mating responses eventually lead to fusion of the gametes
to form a heterokaryotic, quadriflagellated zygote (Figure 1-4). Shortly
after fusion, adhesion between the flagellar is rapidly lost. Subsequent
maturation of all the zygotes causes the withdrawal of the flagellar prior
to spore formation (Figure 1-5; Goodenough et al., 1995). The diploid
spores will remain dormant until favorable conditions allow the spore to
undergo meiosis to form four haploid progeny, two of each mating type (Figure
1-6).
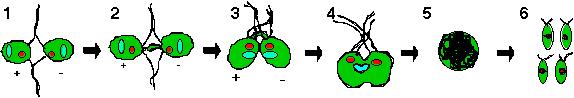
Figure 1: Chlamydomonas mating cycle involves mating type recognition via flagellar agglutination (1-1). The cAMP released will then induce elongation of mating structures (1-2) which leads to the formation of a fertilization tubule that interconnects the two cells (1-3). Fusion of both cells occur to form a quadriflagellated heterokaryon (1-4) which matures into a diploid zygote (1-5). Formation of the four haploid progeny will happen only under favorable environmental conditions.
Agglutinins are mating type-specific adhesion molecules that reside on the surface of the flagellar membrane and they mediate adhesion between the mating types in a "velcro-like" manner. The agglutinins found on both mating types are homologous glycoproteins rich in hydroxyproline (~12%) and serine(~10%) residues (Cooper et al., 1983). Mutagenesis studies have shown that the carbohydrate domains in agglutinins are O-linked glycoproteins (Vallon et al., 1995). Agglutinins and the cell walls of Chlamydomonas and higher plants have very similar amino acid composition (Catt et al., 1976; Vallon et al., 1995). Rapid freeze deep-etch technique revealed that the structural domains of both plus and minus agglutinins are highly homologous (Heuser, 1983). There are three distinct structural domains on agglutinins which are categorized as the head, shaft and hook (Figure 2). Compared to the mt+ agglutinin, the mt- agglutinin is found to have a larger globular head and a gentle bend at the distal portion of the shaft (Goodenough et al., 1985). Several other agglutinin-related proteins have been found on the flagellar surface and they are non-adhesive. These agglutinin-related proteins which are identified as short canes, loops and crescents have no known biological significance (Goodenough et al., 1985). During flagellar agglutination, agglutinins are not fixed in a rigid position. Instead, there is a dynamic flow of agglutinins from a resevoir in the cell body to the surface of the flagellar (Hunnicutt et al.,1990). Since flagellar agglutination is a key event that initiates sexual reproduction, any defects in agglutinins could give rise to a failure of the sexual recognition mechanism.
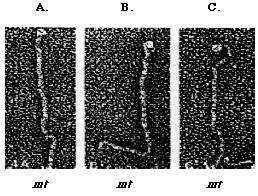
Figure 2: The Heuser rapid-freeze deep etch method (1983) is used to study the structure of Chlamydomonas mt+ (A and B) and mt- (C) flagellar agglutinins. Agglutinin's structure comprise of the distal head, the shaft (top), and the hook (bottom). The shephard's crook-shaped head is unique to mt- agglutinin (Adair, 1985).
The main objective of this research project was to characterize pmh1, a C. reinhardtii mating mutant. pmh1 is a mt- mutant isolated by Dr. Virginia Armbrust while she was a postdoctoral fellow at Washington University. Chlamydomonas mutant strains can be rapidly generated via insertional mutagenesis and plasmid rescue (Quarmby and Hartzell, 1994). To generate mating mutants, Dr. Armbrust transformed a circular plasmid pArg7.8 which codes for argininosuccinate lyase into an auxotrophic mt-Arg7- mutant. Since the auxotrophic mutants cannot survive in an arginine-deficient medium, only those cells that have successfully taken up and integrated the linearized plasmid into the host genome, and thus express the argininosuccinate lyase gene, would grow on the minimal media. The integration of the plasmid into the host genome is random and thus, mutations will occur at the site of insertion. Approximately 1100 transformed cells were screened for mutant phenotypes. In this case, the transformed mt- cells were mated with mt+ gametes and various mutant phenotypes were observed and characterized. pmh1 cells were isolated as one of the mating mutants that could not agglutinate with mt+ cells. However, fusion and zygote formation can be induced if extracellular cAMP is introduced. This suggested that agglutinins might have failed to mediate adhesion since the cell fusion mechanism appeared to be functioning normally (Russo, personal communication).
To further characterize pmh1 at the molecular level, plasmid rescue was carried out to retrieve the inserted plasmid. Using the Sal I restriction enzyme, the plasmid that contained the ampicilin resistance gene (AmpR) and the E. coli origin of replication (OriC) was extracted from the Chlamydomonas genome, recircularized and transformed in competent E. coli cells. Selection for amphicilin resistance was then carried out. Since some of the transformed E. coli contained both the AmpR gene as well as a portion of the Chlamydomonas DNA, the Chlamydomonas DNA flanking the original insertion was cloned (Quarmby and Hartzell, 1994). The Chlamydomonas DNA fragment obtained from the plasmid rescue was used as a probe and a Chlamydomonas wild type genomic library was screened to isolate the putative pmh1 gene. Restriction mapping of the isolated putative pmh1 gene was then performed (Figure 3).
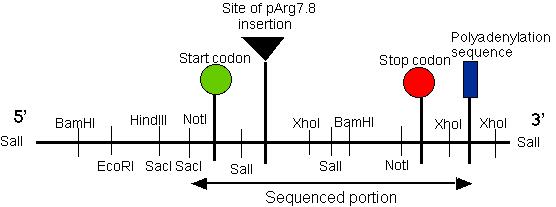
Figure 3: Restriction map of the putative pmh1 gene. The start, stop codons for the largest open reading frame on the cDNA of pmh1 are indicated on the map. The plasmid insertion site is also labeled on this diagram. The sequenced portion of pmh1 gene spans from the Not I/Sac I restriction site to the polyadenylation signal. (Restriction mapping done by E.M. Long in Dr. Armbrust's lab).
The putative pmh1 gene restriction map
allowed Dr. Armbrust and her lab to construct several DNA probes using
various restriction enzymes. Probes A, B, C, D and E spanned the
entire length of the putative gene (Figure 4). These probes were used to
hybridize RNA blots which contained total mRNA derived from wild type mt-,
mt+ cells and the pmh1 mutant. Probes A, D and E bound to a
4.4 kb message, probe C bound to a 1.8 kb message while probe B bound to
both messages (data not shown; Figure 4). Thus, our putative gene must
contain two separate coding regions corresponding to a 1.8 kb and a 4.4
kb message.
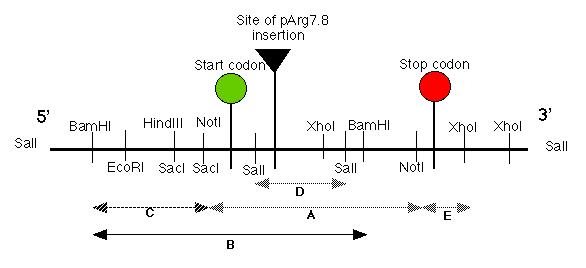
Figure 4: The five probes (A-E) were created by Dr. Armbrust's lab. The probes spanned almost the entire length of the putative pmh1 gene. Probes A, D and E bound to the 4.4 kb message while probe C bound to the 1.8 kb message. Meanwhile, the largest probe, probe B bound to both the 4.4 kb message and the 1.8 kb message. This suggests that the gene that codes for the 1.8 kb message is located upstream from our pmh1 gene that codes for the 4.4 kb message (diagram redrawn from Dr. Virginia Armbrust).
Dr. Armbrust decided to determine the location
and extent of the mutation in the pmh1 gene by doing another RNA
blot to detect the presence of both the mRNA messages in the mutant cell.
Since probe B bound to both the 1.8 kb and 4.4 kb mRNA messages, it was
used to probe the pmh1 RNA blot. The results revealed that only
the 1.8 kb mRNA message was detected. The absence of the 4.4 kb message
suggested that the insertion of the pArg7.8 linearized plasmid must have
occured somewhere in the pmh1 gene that coded for the 4.4 kb message.
Thus, probe B was unable to hybridize with the 4.4 kb message since it
was either no longer being transcribed or is unstable due to the presence
of the inserted plasmid. These results was reconfirmed by using probe E
on a mRNA blot analysis (Figure 5). Probe E bound exclusively to the 4.4
kb transcript. As expected, no bands were detected in both the pmh1
vegetative cells and gametes thus confirming that the pArg7.8 insertion
site occurred in the pmh1 gene.
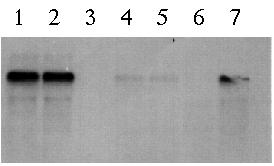
Figure 5: Northern blot using probe E to hybridize total cellular RNA from wild type mt+ vegetative cells (lane 1), wild type mt- vegetative cells (lane 2), pmh1 vegetative cells (lane 3), wild type mt+ gametes (lane 4), wild type mt- gametes (lane 5), pmh1 gametes (lane 6) and 24-hour wild type zygotes (lane 7). Based on the gel, the 4.4 kb message is strongly detected in wild type vegetative cells, faintly detected in wild type gametes and absent in pmh1 gametes and vegetative cells (Northern blot photo from Dr. Virginia Armbrust).
A wild type mt- cDNA library was screened using probe E and four cDNA fragments that bound to the probe were isolated. The lengths of these cDNA fragments were 700bp, 900bp, 1200bp and 1900bp. Dr. Armbrust then carried out sequencing on these cDNA fragments. Shortly after I started working on my project, Dr. Armbrust managed to sequence a partial genomic DNA sequence which we believe contains the putative pmh1 gene. The sequencing data allowed the identification of the coding regions and the polyadenylation signal of the gene. In addition, the location of the start codon and stop codon of the largest open reading frame in the sequence was elucidated (Figure 3).
I hope to characterize pmh1 by cloning the gene and then attempt to rescue the phenotype via complementation. Since the pmh1 cDNA had been sequenced, I also hope to identify the function of the pmh1 gene in Chlamydomonas through sequence analysis. Once the function is determined, a logical hypothesis can be proposed to account for the failure of agglutination in the pmh1 mating mutants.