
GCAT Print Your Own Limited Gene Set Microarrays
If you want your students to produce their own mini-DNA microarrays, you can use this method. One significant benefit of this method is you do not need to isolate any RNA or DNA from biological sources. It is a fast and nearly fool-proof method to use if you want your students to understand the overall method of DNA microarrays.

Davidson logo designed, printed, probed and scanned by Mac Cowell, Kristen DeCelle, and Matt Gemberling (2006).
The goal is to spot DNA onto the class in a predetermined pattern, then use oligonucleotides as probes to hybridize to the spotted DNA. You do not need a robot to use this method, though this page provides details including robot printing. Much of this work was developed by a series of undergraduates at Davidson College (Emily Oldham, Dan Pierce;Danielle Choi, Peter Lowry, Jennifer Madden; and the 2005 "Genome Crew".
You would think this method would produce predictable ratios, but your students will quickly learn that DNA microarrays can produce ratios with little variation, but these ratios do not always match the ratio of probes you pipet! This opens a large area for reading literature and discussing the best uses of microarray data. You can get the latest information from the Davidson GCAT wiki page.
Source of target DNA
You can use any piece of DNA as the target, but we have found cloned PCR products work especially well. By using PCR products, you can target any gene or cDNA. Ideally, you will want to balance the GC content and size of the PCR products. Furthermore, but cloning the PCR products into a plasmid, you can produce very large quantities of target DNA for very little expense. You can also use empty plasmid as negative control DNA.
We chose 10 yeast genes and produced PCR products 500 bp long:
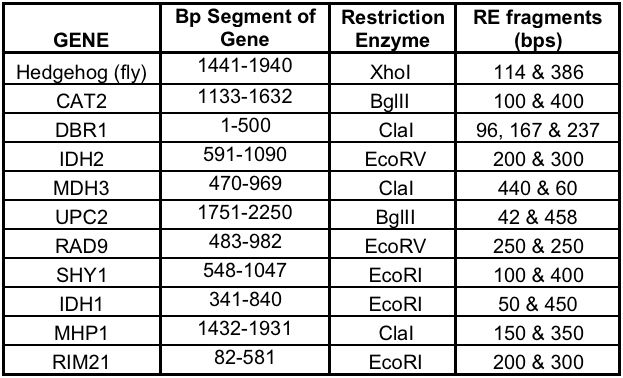
Full 61mers used to produce the Davidson Logo above

Nucleotide sequences of probes for Cy3 and Cy5
for Mhp, Shy, and Idh1.
31mer dendrimer capture sequences are in italics and 30mer
DNA probes are in bold.
Printing microarrays
You can hand-spot the DNA using a pipet. We have done this for quick and dirty mini-arrays. You can purchase a hand-printer or you can go high-tech and purchase a spotting robot. We have a developed an extensive protocol that works very well with our robot (download PDF or Word version). A comparison of printing methods, as well as scanners and dyes can be found at this web page.
Probes for hybridization
We developed a series of oligonucleotides (61mers) that do not form hairpins, do not bind to themselves, do not bind to each other, and only bind to one of the 10 different PCR products. The 61mers also have similar melting temperatures when bound to target DNA.

Detection of probes
We use the 3DNA method to detect the bound oligos because the signal amplification is so great. This method works very well with a very high success rate, even for beginners.
Click here for a step-by-step protocol of the entire process (9 MB Word File)
© Copyright 2006
Department of Biology, Davidson College, Davidson, NC 28035
Send comments, questions, and suggestions to: macampbell@davidson.edu