This web page was produced as an assignment for an undergraduate course at Davidson College.
Expression of Yeast genes Msp1 and YGR031W in Microarrays
Msp1
As discussed in the previous web assigment (click here to view), Msp1 is a yeast gene located on chromosome 7 that is a member of the AAA ATPase family and is responsible from the translocation of proteins into the mitochondria (Gene Ontology). I plan to investigate the role(s) of Msp1 using microarray data to examine expression patterns in response to various environmental conditions.
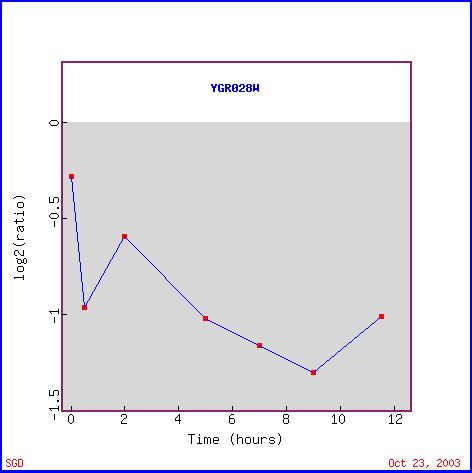
Msp1 (also called YGR028W) expression decreased significantly during sporulation. The decrease in Msp1 expression (see Figure 1), which is supposed to be linked to mitochondrial protein transport, is probably due to a lack of nutrients. Yeast produce spores when there is a lack of nutrients in the environment (Howard Hughes Medical Institute). A lack of nutrients would imply that there is a lack of nutrients for the cell to breakdown and use for energy. Since the mitochondria is used mainly for energy production, there is no need for the cell to waste it's precious energy producing mitochondrial proteins if the mitochondria cannot function. Therefore, it is logical to see the decrease in the levels of mRNA in a cell undergoing sporulation. In addition other mitochondrial proteins have reduced expression levels similar to Msp1, such as CYC2 and CAT5 (see Figure 2). Figure 2 also shows other genes with similar decreases in expression during the sporulation process. The gene right below Msp1, RIC1, is also involved in intracellular protein transport, although it functions in the nucleus. With less proteins being produced as the cell undergos sporulation, it seems logical that both of these protein transport moleculars have decreased production.

Figure 1. Expression over time of Msp1 during sporulation (photo from Expression Connection).
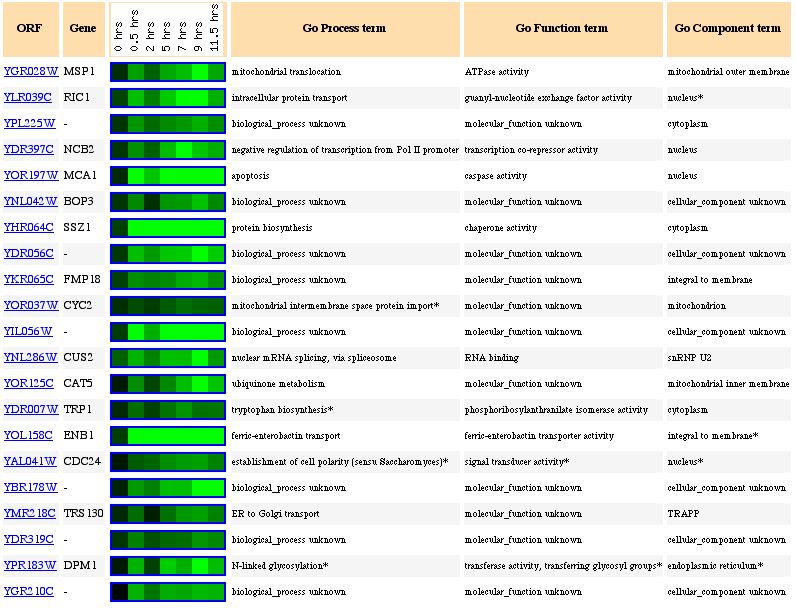

Figure 2. Genes that have similar expression patterns to Msp1 during sporulation (photo from Expression Connection).
Figure 3 shows the increase in expression level of Msp1 during the diauxic shift. The diauxic shift occurs when yeast cells switch from anaerobic to aerobic respiration. Mitochondria are the cell organelle that is responsible for the energy production during aerobic respiration (in other words, the majority of the ATP from aerobic respiration comes from the mitochondria). Therefore, the increase in Msp1 expression during the diauxic shift is expected. When the cell needs to use the mitochondria to produce ATP, production of mitochondrial related proteins is expected to increase, as we see in Figure 3. Figure 4 shows other genes with similar expression patterns and several are related to the mitochondria and the formation of ATP, as we might have predicted, such as IDH2, LSC2, LPD1, COX4 and ATP5.
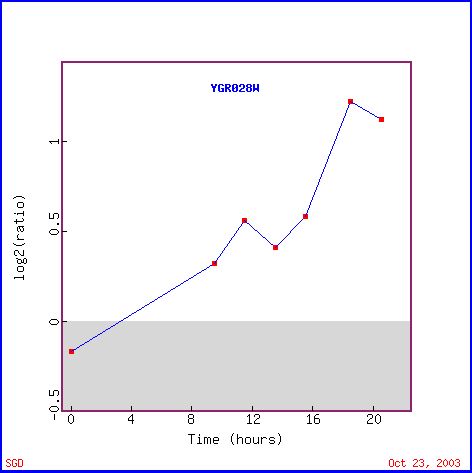
Figure 3. Expression of Msp1 during the diauxic shift (photo from Expression connection).
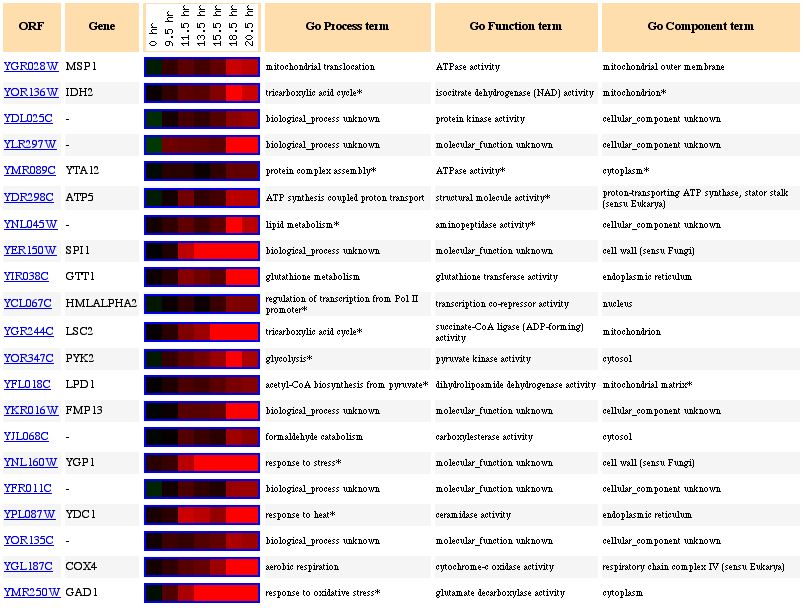

Figure 4. Expression of Msp1 and genes of similar expression during the diauxic shift (photo from Expression connection).
Based on the information regarding the changes in Msp1 expression from microarrays, I can conclude that Msp1 is a mitochondrial protein that is involved in the cell's response to aerobic conditions, as is indicated by the gene's gene ontology annotations.
YGR031W
In my previous webpage, I asserted that YGR031W was a yeast gene that was a lysophospholipase embedded across a membrane in the cell, perhaps the Golgi or the ER. I will try to use evidence from changes in YGR031W expression patterns in yeast to verify my hypothesis.
There is a high number of genes with similar expression patterns to YGR031W during sporulation (see Figure 4) that are located in the cytoplasm, nucleus, and plasma membranes. Since I previously saw that YGR031W is predicted to have a transmembrane domain, I may be able to assume that YGR031W is associated with a plasma membrane and interacts mainly with the cytoplasm. During sporulation, the activity of YGR031W breifly increases and then decreases (see Figure 3). A lysophospholipase is an enzyme that catalyses the hydrolysis of a single fatty acid ester bond in lysoglycerophosphatidates with the formation of glyceryl phosphatidates and a fatty acid. Since lysophospholipases catalize the formation of fatty acids from larger lysoglycerophosphatidates, the inital increase in expression after induction of sporation could be a way for the cell to gain a small amount of energy from it's own stores since there is a lack of nutrients in the environment. The subsequent decrease in expression may indicate that the cell has enough energy to produce spores and has entered a dormant stage where the production of YGR031W is not needed and is therefore suppressed to save energy.
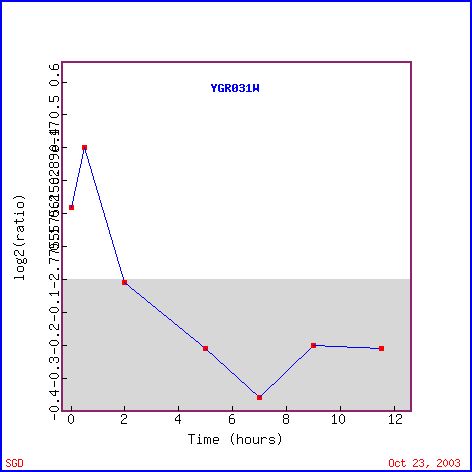
Figure 5. Expression of YGR031W during sporulation (photo from Expression connection).

Figure 6. Expression of YGR031W and genes with similar expression patterns during sporulation (photo from Expression connection).
The expression of YGR031W decreased in response to histone deletion (see Figure 7). Since histones are thought to indirectly regulate gene control through access to the gene, the deletion of histones causing a suppression of YGR031W is significant. Only 10% of yeast genes had decreased expression levels compared to wildtype yeast (Wyrick et al, 1999). Since there is a decreased level of expression of YGR031W in response to decreased histone levels, we can guess that expression of YGR031W is physically affected by the lack of histones and is induced through the presence of histones (ie the folding around the histone actually induces expression), or that YGR031W is repressed by gene whose regulation is increased in the absence of histones. Or we could assume that there is some functional relationship between the lack of histones and the repression of YGR031W. However, the genes with similar expression patterns (see Figure 8) do not appear to have anything in common and therefore, it is possible that the expression of YGR031W is physically linked (ie induced) in the presence of histones.
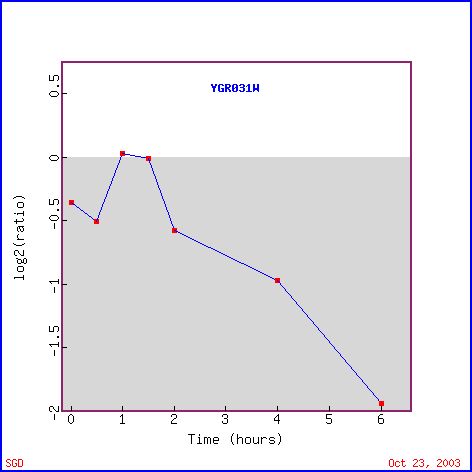
Figure 7. Expression of YGR031W during histone depletion (photo from Expression connection).
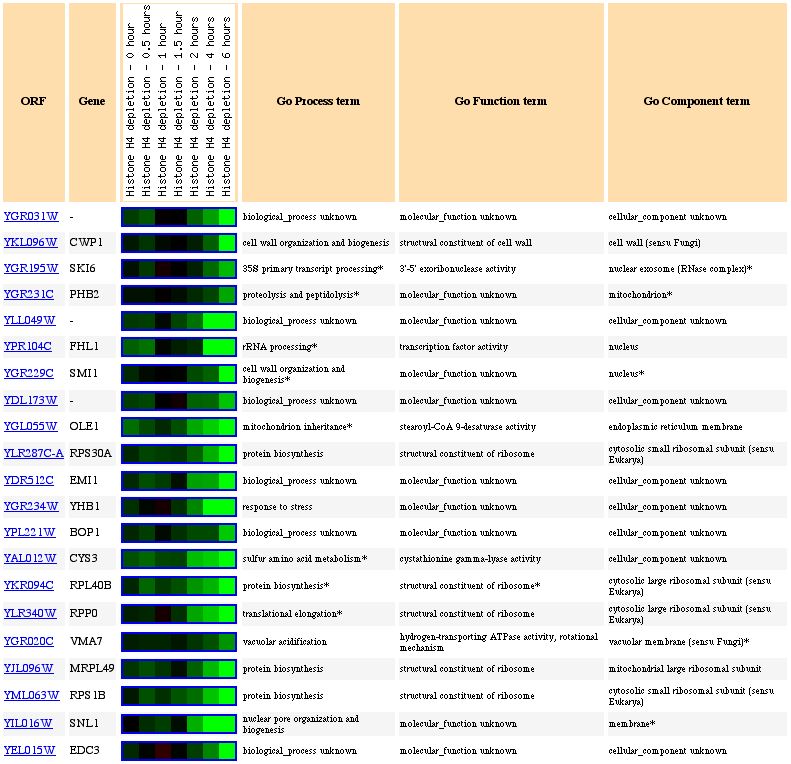

Figure 8. Expression of YGR031W and genes with similar expression during histone depletion (photo from Expression connection).
Lastly, changes in the environmental levels of zinc repress the expression of YGR031W. Of the genes whose gene ontology annotations are known, many are integral to membranes such as the vacular membrane and the cell wall. This data supports my hypothesis that YGR031W is a transmembrane spanning protein.
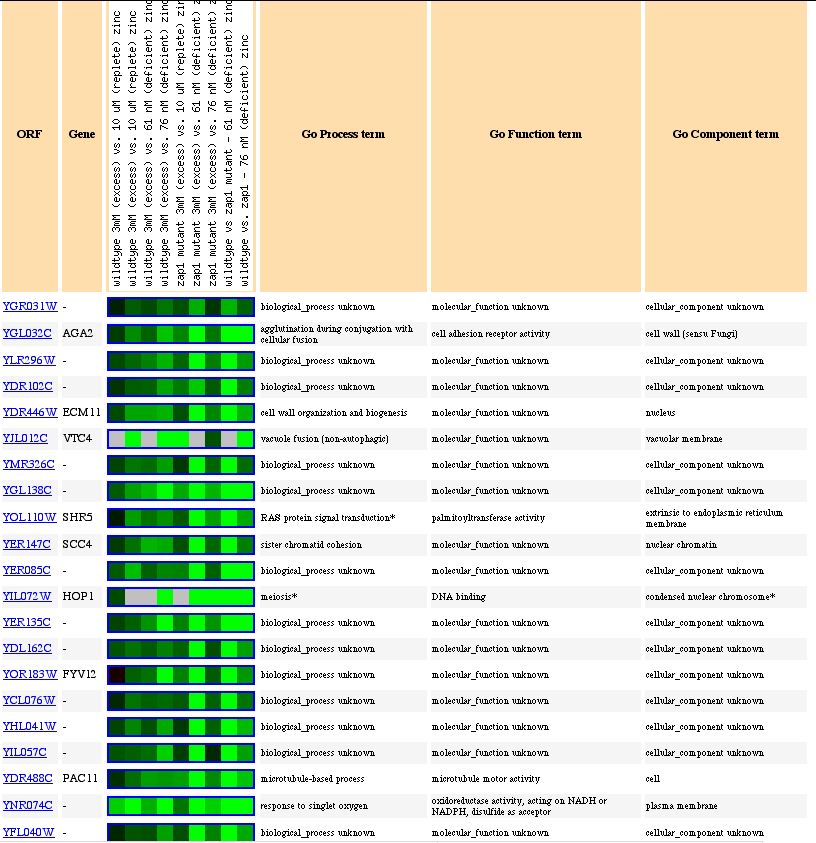
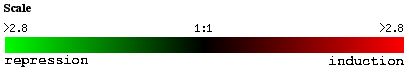
Figure 9. Expression of YGR031W and genes with similar expression patterns in response to varying levels of zinc (photo from Expression connection).
In conclusion, the evidence from microarray expression supports my hypothesis (or at least, it doesn't disprove it) that YGR031W is a lypophospholipase that is integral to a membrane, such as the nuclear or ER, and seems to interact with cytoplasm.
References
Chu, S, J DeRisi, M Eisen, J Mulholland, D Botstein, P Brown, and I Herskowitz. 1998. The transcriptional program of sporulation in budding yeast. Science 282(5389):699-705.
DeRisi, J, V Iyer, and P Brown. 1997. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278(5338):680-6.
Expression Connection. <http://genome-www4.stanford.edu/cgi-bin/SGD/expression/expressionConnection.pl>
Lyons, T, A Gasch, L Gaither, D Botstein, P Brown, and D Eide. 2000. Genome-wide characterization of the Zap1p zinc-responsive region in yeast. Proceedings of the National Acadamy of the Sciences 97(14):7957-62.
Wyrick, J, F Holstege, E Jennings, H Causton, D Shore, M Grunstein, E Lander, and R Young. 1999. Chromosomal landscape of nucleosome-dependent gene expression and silencing in yeast. Nature 402(6760):418-21.
Questions or comments?: e-mail Sarah Baxley