SER2p's interactions with within the cell can reveal a whole new world of the protein's functions. The first place to look is at other proteins in which SER2p interacts with. By knowing the proteins which SER2p interacts with can clarify the function of the gene product and show how the product is integrated into the cell's systems.
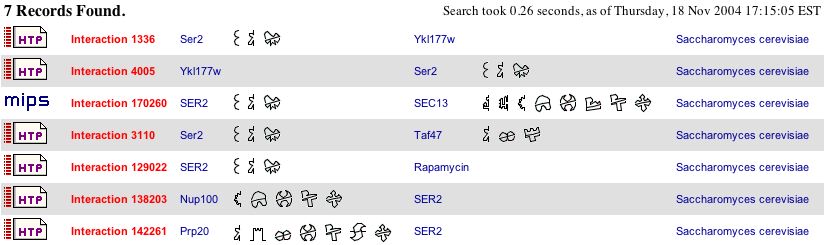
According to the BIND database , SER2p is interacts with several other proteins, shown below:

<http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR208W>
The function of these genes with which SER2p interacts is important to know, because it can show different processes SER2p is involved with. For example, SEC13 is known to be a part of a complex which releases transports vesicles from the ER to the Golgi. Taf47p is a part of a transcription factor complex; Nup100p is involved in RNA and protein transport; Prp20p is involved in DNA binding and mitosis; Ykl177wp has unknown functions. Rapamycin is a biochemical molecule which caused a moderate chemical-genetic sensitivity interaction with SER2 (Bader, Betel and Hogue, 2003, <http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR208W>).
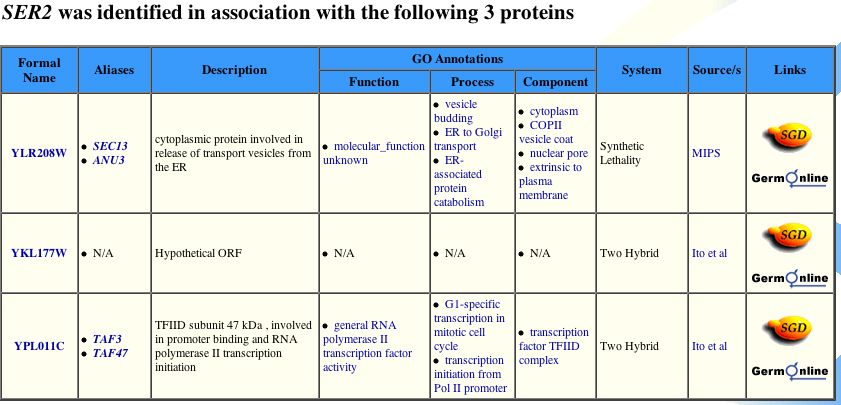
Yeast Grid Database supports the BIND data, as shown below:
<http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR208W>
| SER2's predicted function, serine and glycine synthesis, is unrelated to the predicted functions of the interacting genes. While more supporting evidence is needed, this information significantly shakes the prediction that SER2p is only involved in serine and glycine synthesis. It is possible that another version, an alternatively spliced protein, could function as a transcription factor or protein transport as well, accounting for the reported difference in function. Puzzling, is the lack of any gene product to be involved in amino acid synthesis. The significance of SER2's sensitivity to rapamycin is unclear from the BIND site, other than that mutations in the gene could exhibit phenotypes of drug sensitivity. |
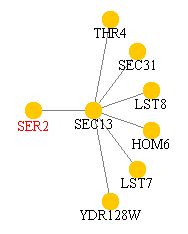
PathCalling is another protein-protein interaction web site. It lists SER2p as only interacting with SEC13p. Both BIND and Yeast Grid note the interaction.
 <http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastGene?geneIdIn=2403>
<http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastGene?geneIdIn=2403>
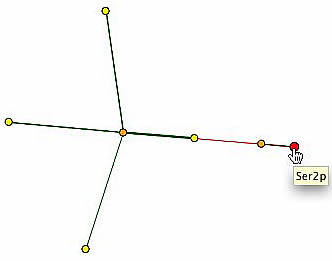
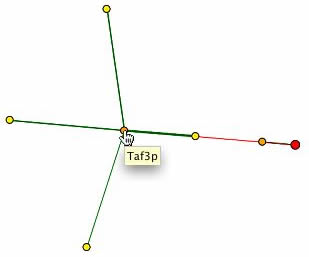
Another helpful database is the Database of Interacting Proteins [DIP]. DIP provides a graph of SER2's interacting proteins, shown below. However, even less information is available, since it has SER2 only interacting with YKL177W, a hypothetical ORF with unknown function, represented by the orange node adjacent to SER2 (left). Taf3p is the only other protein indicated on the graph whose association with SER2p was described in either the BIND or Yeast Grid database (right). Taf1p, Taf14p, Cdc39p, and Taf10p radiate from Taf3p. These gene products are associated with some variation of RNA polymerase II activity. The notable exception is Taf1p (topmost yellow node), which in addition to RNA polymerase activity, is also involved in serine/threonine kinase activity.
<http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=2085>
| The information of protein interaction on DIP does not completely verify the information found on the other databases. Two proteins are present consistently, but with one being completely unidentified and the other being unrelated to amino acid synthesis, it is difficult to say whether or not SER2p could also be involved with transcription, specifically with RNA polymerase II. The presence of Taf1, however, sheds new light on the mystery. The function of Taf1p is thought to be a serine/theonine kinase, which binds a phosphate group to an amino acid (SGD, 2004, <http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=19202>). Ser2p is though to catalyze the reaction of phosphoserine into serine and a phosphate group (SGD, 2004, <http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=4647>). Taf1 is not involved in the production of serine, according to SGD (2004, <http://db.yeastgenome.org/cgi-bin/search/textSearch?query=serine&type=pathway>), but it is plausible that serine is phosphorolated during some biological process, thereby connecting Ser2p and Taf1p, though not directly, as the graph above indicates. |
PDB does not have a known structure for SER2p. However, a search shows a series of homologs with no greater than 40% similarity. A picture of this structure can be found in the text of assignment two.
| The evidence between the interaction databases are also spotty and inconsistent.Most of the data presented does not support the hypothesis that SER2p is only involved in serine and glycine biosynthesis. The other gene products that SER2p associates with are not involved in the serine/glycine production pathway. Most are associated with transcription and protein transport. Because so many of the genes that are associated with SER2p are involved in similar pathways, it begs to ask the question why SER2p is associated with them if it does not share the same function. Two hypothesis are suggested: First, SER2p may in fact be a factor in either or both transcription and protein transport as well as serine biosynthesis, but researchers have not yet made the connections. The second hypothesis is that serine/glycine biosynthesis and transcription and/or protein transport are related in some unknown way. The first hypothesis may be the more likely of the two to be correct, with Taf1p showing how. Taf1p is involved in both RNA polymerase II activity and in serine/theonine kinase. This dual function, if accurate, illustrates how SER2p could also have two unrelated functions. |
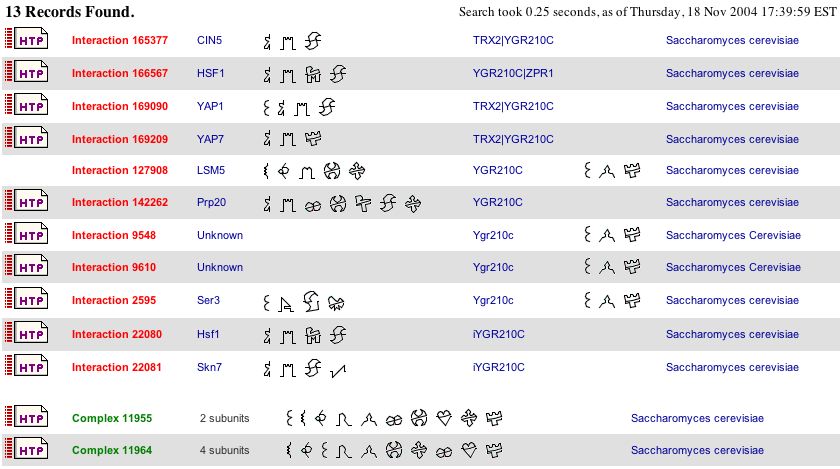
Again, interacting and associating genes are the place to look first. Below is a table of YGR210Cp's gene interactions. Several things should be clarified. "TRX2|YGR210C" indicates that the interacting gene is actually interacting with the genetic region between TRX2 and YGR210C. As seen on assignment two page, YGR210C is adjacent on the chromosome to TRX2 and ZPR1 (the first four gene interactions listed below.) The authors denote that the region regulates expression for TRX2 in the case of "TRX2|YGR210C" interactions, and that the interacting region in the case of "YGR210C|ZPR1" regulates both YGR210C and ZPR1. iYGR210C, shown in the last two gene interactions, indicates that the gene is interacting with the intergenic region which regulates expression of both YGR210C and ZPR1/YGR211W. The two complexes, shown at the very bottom of the list, indicate that YGR210C forms a protein complex. Although not noted on the table below, YGR210C forms a complex with PWP2p in Complex 11955 and with YLR222Cp, YDR449Cp, and PWP2p in Complex 11964 (Bader, Betal, and Hogue, 2003, <http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR210C>).
<http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR210C>
Yeast Grid also listed PWP2p and SER3 as having interactions with YGR210Cp ( Bader, Betel, and Hogue, 2001, <http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR210C>).
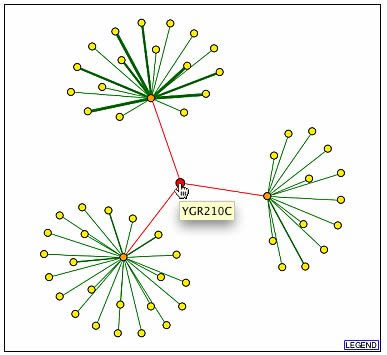
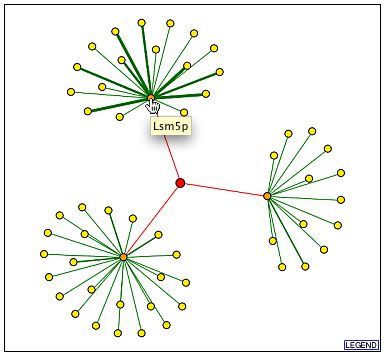
The following images are from DIP. YGR210Cp is in the middle, but note that the red edges indicted interactions found by high through-put methods without any experimental verification. All the noted genes in the three other images are found on the BIND database and were discussed above (DIP, 2001, <http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=2600>).
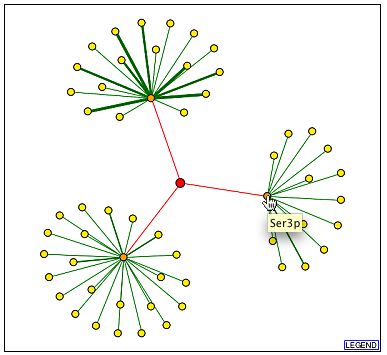
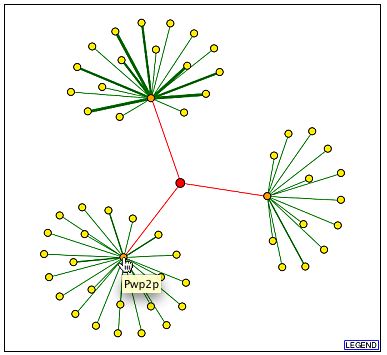
<http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=2600>
Many interesting interactions are found in the BIND database. First, consider the interactions occurring between YGR210C and its neighboring genes. Since the interactions are occurring between the genes, this might indicate that YGR210C is misclassified as an ORF and may be a promoter region instead. This is contradicted, however, by the other protein interactions with YGR210C product. HSF1p interaction with the intergenic region of YGR210C|ZPR1 is significant because they have similar functions: HSF1p is known to be involved in DNA binding, transcription regulation, protein folding, stress, and heat response. YGR210Cp is thought to be a type of zinc finger, involved in DNA binding and protein synthesis. SKN7p and iYGR210Cp interaction shows that the two proteins also share similar functions. Interactions with LSM5p further support that YGR210Cp is involved in protein synthesis or it is a part of a ribosomal subunit. LSM5p is projected to be involved in RNA binding and found within a small nucleolar ribonucleoprotein complex [SNRC]. Other functions of this protein that are loosely related to YGR210Cp's function are rRNA and mRNA processing. PRP20p is also promising. It directly binds to DNA and is involved in the export of macromolecules from the nucleus to the ribosomes. These biological process are similar and perhaps even done in conjunction with the YGR210Cp's projected function. SER3p's function is not similar to that of YGR210Cp's, but it is interesting to note that SER3p is involved in serine biosynthesis, same as SER2p. LSM5p and SER3p are also shown on the protein interaction graphs from DIP. The complexes which are formed with YGR210Cp are more complicated to interpret because there is no known function of the complex, either. PWP2p, which is listed in both BIND complexes and, according to DIP, interacts with YGR210Cp, is thought to be a part of a SNRC, as well as YLR222Cp and YDR449Cp. This suggests that if YGR210Cp also is a part of this complex, it too, would be found in a SNRC. According to AmiGO, an SNRC has a long stretch of complementary RNA found in mature mRNA that helps process ribosomal RNA (2004, <http://www.godatabase.org/cgi-bin/amigo/go.cgi?view=query&action=query&search_constraint=terms&query=5732>). This is consistent with the previously presented hypothesis that YGR210Cp is in a ribosomal subunit, and interacts directly with DNA, or now, perhaps RNA. As a side note, BIND lists YGR210Cp as having a molecular function of GTP binding and a biological process of regulation of transcription, DNA-dependent. |
Ideal experiments to verify the functional hypotheses of YGR210Cp would include all aspects of gene ontology. Here are a few examples of how to test the predicted gene ontologies: Cellular Component: The location of YGR210Cp in the cell might be the easiest place to start. It is predicted that YGR210Cp is a part of SNRCs. To test this, first isolate wt SNRC and create a knockout strain of YGR210C and isolate this strain's SNRC. Perform a western blot and compare the sizes of the protein complexes. If the knockout complex is smaller in size, assume that YGR210Cp is a part of the complex. If there is no difference in complex size, it can be assumed that YGR210Cp is not a part of the complex. Another experiment that can be done is to probe a cell with tagged YGR210Cp antibodies. This method will reveal where the protein is located. Variations on this method can be used to discover if the protein is always present in one part of the cell of if it moves under varying circumstances and stresses. Biological Process: If YGR210Cp is a part of the SNRC, then it is important to find the biological pathways which SNRCs are involved in. Examining gene expression data in under many circumstances and stresses of all the components of the complex could provide clues to find the associated pathways. Determining the 3D structure of both YGR210C and the complex could provide valuable clues to its function as well. A great experiment (but perhaps and idealistic one) would be to create a microarray-like assay specifically for YGR210Cp and SNRC with all the cellular components present, including organelles, inorganic molecules - anything that is present in a living yeast cell. Probing with marked YGR210Cp and SNRC would show not only the proteins with which they interacted, but also if there were other factors as well, such as ligands, DNA or RNA attachments, and the like. Again, this may be an impossible experiment to perform, but it would reveal huge amounts of information that after analysis would help piece together the pathway puzzle. Molecular Function: An experiment determining the molecular function of YGR210Cp could be done using three testing probes marked differently (e.g. immunoflourescence at three different wavelengths.) The three probes would be YGR210Cp, a functional wt SNRC, and a mutated SNRC missing the YGR210C protein. Each probe would test a number of different functional variables. By comparing wt SNRC's functions to SNRC(YGR210Cp) and YGR210Cp, the function of both YGR210Cp and SNRCs could be inferred. Examples of functional tests would be whether or not the molecule bound to DNA or RNA, whether it assisted in protein folding, whether it affected transcription rates, or any other function that could be associated with the proposed pathway. |
AmiGO. 2004, Nov. Small nucleolar ribonucleoprotein complex. <http://www.godatabase.org/cgi-bin/amigo/go.cgi?view=query&action=query&search_constraint=terms&query=5732>. Accessed 2004 Nov 18.
Bader GD, Betel D, Hogue CW. 2003. BIND: the Biomolecular Interaction Network Database. Nucleic Acids Res. 31(1):248-50 <http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR208W>. Accessed 2004 Nov 18.
Bader GD, Betel D, Hogue CW. 2003. BIND: the Biomolecular Interaction Network Database. Nucleic Acids Res. 31(1):248-50 <http://bind.ca/Action?textquery=RecordType:%20(interaction%20complex%20pathway%20)+AND+YGR210C>. Accessed 2004 Nov 18.
Breitkreutz, BJ and Stark C. 2001. Yeast Grid. <http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR208W>. Accessed 2004 Nov 18.
Breitkreutz, BJ and Stark C. 2001. Yeast Grid. <http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR210C>. Accessed 2004 Nov 19.
Database of Interacting Proteins [DIP]. 2001 Dec 31. DIP:2085N. <http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=2085>. Accessed 2004 Nov 18.
Database of Interacting Proteins [DIP]. 2001 Dec 31. DIP:2600N. <http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=2600>. Accessed 2004 Nov 19.
Saccharomyces Genome Database [SGD]. 2004. <http://www.yeastgenome.org/>. Accessed 2004 Nov 18.
Databases searched, but with no results of either gene: Swiss-2D Database, ExPASy, PDB, PROWL, TRIPLES, Schwikowski PDF Files, and PDB and PathCalling (YGR210C, only).
Contact Megan at mecastle@davidson.edu