This web page was produced as an assigment for an undergraduate course at
Davidson College.
The pET Expression System
Introduction
Expression systems are designed to produce many copies of a desired protein
within a host cell. In order to accomplish this, an expression vector is inserted
into a host cell. This vector contains all of the genetic coding necessary
to produce the protein, including a promoter appropriate to the host cell,
a sequence which terminates transcription, and a sequence which codes for
ribosome binding (Purves et al., 2001). One expression system was developed
in 1986 by W. F. Studier and B. A. Moffatt, who created an RNA polymerase
expression system which was highly selective for bacteriophage T7 RNA polymerase.
The initial system involved two different methods of maintaining T7 RNA polymerase
into the cell - in one method, a lambda bacteriophage was used to insert the
gene which codes for T7 RNA polymerase, and in the other, the gene for T7
RNA polymerase was inserted into the host chromosome (Studier et al, 1986).
This expression system has become known as the pET Expression System, and
is now widely used because of its ability to mass-produce proteins, the specificity
involved in the T7 promoter which only binds T7 RNA polymerase, and also the
design of the system which allows for the easy manipulation of how much of
the desired protein is expressed and when that expression occurs. (Unger,
1997; Novagen, 2002-2003).
Producing Proteins with the pET Expression
System
Designing the pET Plasmid:
A pET vector is a bacterial plasmid designed to enable the quick
production of a large quantity of any desired protein when activated. This
plasmid (pictured below) contains several important elements - a lacI
gene which codes for the lac repressor protein, a T7 promoter which
is specific to only T7 RNA polymerase (not bacterial RNA polymerase) and also
does not occur anywhere in the prokaryotic genome, a lac operator
which can block transcription, a polylinker, an f1 origin of replication (so
that a single-stranded plasmid can be produced when co-infected with M13 helper
phage), an ampicillin resistance gene, and a ColE1 origin of replication (Blaber,
1998).
To start the process, your favorite gene (YFG) is cloned into
a pET plasmid at the polylinker site. Both the T7 promoter and the lac
operator are located 5' to YFG. When the T7 RNA polymerase is present and
the lac operator is not repressed, the transcription of YFG proceeds
rapidly. Because T7 is a viral promoter, it transcribes rapidly and profusely
for as long as the T7 RNA polymerase is present (Campbell, 2003). The expression
of your favorite protein (YFP) increases rapidly as the amount of mRNA transcribed
from YFG increases. Within a few hours, YFP is one of the most prevalent components
of the cell. (Unger, 1997).

Figure 1: The pET Vector. This plasmid contains
a drug resistant marker for ampicillin resistance (green), the lacI
gene (blue), the T7 transcription promoter (red), the lac operator region
(pale green) 3' to the T7 promoter, and a polylinker region (black). Also,
there are two origins of replication - one is the f1 origin which enables
the production of a single stranded vector under appropriate conditions, and
the other is the conventional origin of replication. This image was used with
permission from Dr. Michael Blaber, of Florida State University. The original
reference can be found here.
Introducing T7 RNA Polymerase to the
Host Cell:
One of the most important parts of the pET Expression System
involves the fact that YFG is not transcribed unless the T7 RNA polymerase
is present. Prokaryotic cells do not produce this type of RNA, and therefore
the T7 RNA polymerase must be added. Usually, the host cell for this expression
system is a bacteria which has been genetically engineered to incorporate
the gene for T7 RNA polymerase, the lac promoter and the lac
operator in it's genome. When lactose or a molecule similar to lactose is
present inside the cell, transcription of the T7 RNA polymerase is activated..
Typically, the host cell used is E. coli strain BL(DE3)
(Blaber, 1998). The diagram below shows the genome of the host cell.
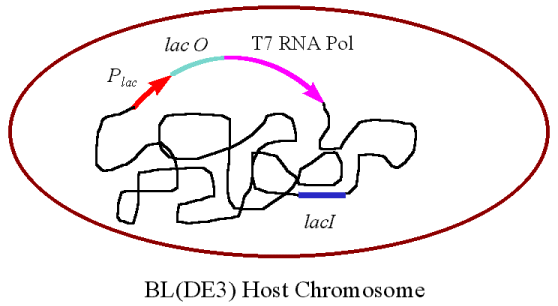
Figure 2: The Host Chromosome. The lac
promoter is in red, the lac operator is green, the gene which encodes
the T7 RNA polymerase is pink, and the lac inducer is blue. The host
cell genome is represented by the black line, and the brown line represents
the cell membrane. This image was used with permission from Dr. Michael Blaber,
of Florida State University. The original reference can be found here.
T7 RNA polymerase can be introduced to the cell through methods
other than incorporation a gene in the host chromosome. It can be introduced
through infection of the original host cell with lambda CE6 (Novagen 2003-2003).
Controlling the pET Expression System
Control of the pET expression system is accomplished through
the lac promoter and operator. Before YFG can be transcribed, T7
polymerase must be present. The gene on the host cell chromosome usually has
an inducible promoter which is activated by IPTG. This molecule, IPTG, displaces
the repressor from the lac operator. Since there are lac
operators on both the gene encoding T7 polymerase and YFG, IPTG activates
both genes (Science Advisory Board, 2003; Campbell, 2003). Therefore, when
IPTG is added to the cell, the T7 polymerase is expressed, and quickly begins
to transcribe YFG which is then translated as YFP.
IPTG works to displace a lac repressor since IPTG is
an analogue of lactose (Blaber, 1998). The lac genes express enzymes
which are involved in the breaking down of lactose, and therefore, the presence
of lactose (or it's analogue) would trigger the initiation of transcription
of lac genes.
References
Blaber M. 1998 Spring. Web Page for Lecture 25 of Molecular
Biology and Biotechnology Course: Prokaryotic Expression Vectors. <http://wine1.sb.fsu.edu/bch5425/lect25/lect25.htm>.
Accessed 2003 Feb17.
Campbell M.. 2003 Jan 31. Class Lecture.
Heller HC, Orians GH, Purves WK, Sadava D. 2001. Life: The Science
of Biology. 6th ed. Sunderland (MA): Sinauer Associates, Inc.; p 251, p 322-323
Novagen.. 2002-2003. Protein Expression: Prokaryotic Expression:
pETBlue and pET System Overview. Novagen 2002-2003 Catalog. p 84-91. <http://www.novagen.com/SharedImages/Novagen/05_PROEXP.pdf>.
Accessed 2003 Feb 17.
Novagen. 2001. Protein Expression: Prokaryotic Expression: pET
System Overview. 2001 Novagen Catalog. p 68-72. <http://www.novagen.com/docs/ndis/C183-001.pdf>.
Accessed 2003 Feb 17.
Socalsmithers of the Science Advisory Board. (Date not specified,
but site is copyrighted 1997-2003). Product Reviews: Novagen pET Vector Expression
System. <http://www.scienceboard.net/resources/prodreviews.asp?cat=9&product=339>.
Accessed 2003 Feb 17.
Moffatt BA, Studier FW. 1986. Use of Bacteriophage T-7 RNA Polymerase
to Direct Selective High-Level Expression of Cloned Genes [abstract]. In J
Molecular Biology 189(1):113-130. WebSPIRS database can be accessed under
"Science and Technology" at <http://www.davidson.edu/administrative/library/refer/indexes.asp?dbtype=Science&search=Submit+Query>.
Accessed 2003 Feb 17.
Unger TF. 1997. Show Me The Money: Prokaryotic Expression Vectors
and Purification Systems. The Scientist 11(17). <http://www.the-scientist.com/yr1997/sept/profile2_970901.html>.
Accessed 2003 Feb 17.
Send questions, comments, and suggestions
to Jamie Causey.

