This web page was produced as an assignment for an undergraduate course at Davidson College.
My Favorite Yeast Genes: GSH1 and SET4
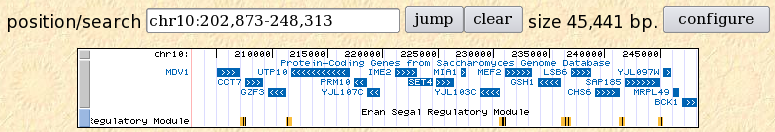
My goal in this project was to find two genes in yeast. One must have an annotated function, while the other must be an annotated ORF without any known function. I chose two genes, GSH1 and SET4, and in this web page describe them. They are both located on yeast chromosome X (10) (Figure 1).
Figure 1: Locations of GSH1 and SET4 on yeast chromosome 10 (UCSC Genome Browser v143, 2006.).
GSH1: Gamma-glutamylcysteine synthetase
Where can I find GSH1?
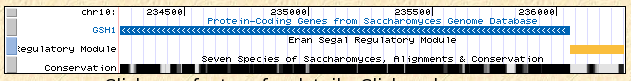
GSH1 is an annotated gene in the Saccharomyces cerevisiae (baker's yeast) genome, and is located on chromosome X (10).
Figure 2: Chromosomal location and conservation of GSH1 in S. cerevisiae (UCSC Genome Browser v143, 2006.).
What is the role of GSH1 in the life of yeast?
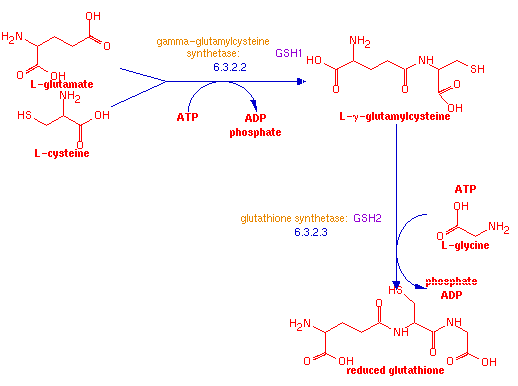
GSH1 is a yeast gene which encodes the protein gamma-glutamylcysteine synthetase. This protein synthesizes L-γ-glutamylcysteine from the amino acids L-cysteine and L-glutamate and with ATP. Another protein, glutathione synthetase, encoded by GSH2, then adds a third amino acid, glycine, to produce the tripeptide reduced glutathione.
Glutathione's primary function is to reduce cellular toxicity, and acts as an antioxidant to remove free radicals. The cysteine amino acid in glutathione contains a sulfur atom which can be reduced, and so acts as a reservoir for free radical electrons. Free radicals are often the products of cellular metabolism, especially in the mitochondria. L-γ-glutamylcysteine is usually converted into glutathionine for optimal antioxidation (Wikipedia, 2006). However, if GSH2 malfunctions, L-γ-glutamylcysteine can still function as an antioxidant.
Glutathione has also been shown to play an critical role in determining the structures of iron/sulfur proteins outside the mitochondria.
Figure 3: Schematic diagram of glutathione synthesis. (Yeast Genome Pathway Analysis, 2006.).
Mutant Alleles and Phenotypes
Determining the function of a gene can be difficult. Analysis and observation of mutant phenotypes can help clarify what roles a gene plays in its organism. Several studies on S. cerevisiae have shown some distinct mutant phenotypes for GSH1, including increased susceptibility to oxidative damage, growth termination, and defective extra-mitochondrial iron/sulfur protein formation. GSH1 is essential in eukaryotes but not in prokaryotes (Spector et al., 2001).
Growth in Reactive Oxygen Species
The antioxidant properties of glutathione become most apparent under conditions of higher than normal reactive oxygen species (ROS) concentrations. Researchers at Kyoto University in Japan wanted to know more about when glutathione becomes activated and where it is localized. They found that glutathione becomes activated in the presence of lipid hydroperoxide and other oxidative stressors. They also localized the highest concentrations of the protein, and determined that 95% of the enzyme was found inside the inner mitochondrial membrane (Inoue et al., 1995). The mitochondria are the source of many free radicals due to energy metabolism, so it would make sense to find glutathione there to a.
A different group of researchers, based in France, Germany, and Hungary, studied the phenotypes of strains with GSH1 deletions. They noticed that, upon deletion of the gene, cellular growth came to a stop within 3 days, and that glutathione was only present in minute quantities. It was also noted that, with the addition of other antioxidants, such as dithiothreitol, growth was restored. These experiments support the notion that GSH1 is a gene whose function is primarily protection against free radicals. The deletion of GSH1 leads to a rapid increase in cellular damage, but this can be offset by the introduction of antioxidants. (Sipos et al., 2002)
Maturation of Cytosolic Iron-Sulfur Proteins
Although Sipos et al. studied the growth of S. cerevisiae with GSH1 deletions, this was not their primary focus. Their goal was to investigate the effects of GSH1 on a different set of biological pathways. They found that, in addition to functioning as an antioxidant, glutathione is also involved with the correct formation of certain iron/sulfur proteins. Their study distinguished between mitochondrial and cellular proteins. While mitochondrial protein formation and function was unaffected, extra-mitochondrial iron/sulfur proteins had difficulty maturing and functioning. The study concluded that glutathione not only offers a protective role from ROS, but is critical for proper maturation of iron/sulfur proteins (Sipos et. al, 2002).
In light of the fact that glutathione is found primarily inside the mitochondria, it is surprising to see that it has a pronounced effect on extra-mitochondrial proteins. The reasons for this are unclear, but there may be much more to the roles glutathione plays than was previously imagined.
Regulation
In order for GSH1 to be expressed in yeast cells, certain regulational factors are required. Researchers in Edinburgh and Dundee, Scotland, found that the presence of heavy metals, such as cadmium, and oxidants play a major role in activating GSH1 transcription. The heavy metals are specific to binding sites in the promoter region of GSH1. In the presence of hydrogen peroxide, amino acids glutamate, glutamine, and lysine are required for proper gene regulation. In addition, the Yap1 and Yap2 proteins also affect the transcription of GSH1 (Stephen et al., 1997, Dormer et al. 2002). Thus it appears that glutathione may be produced only as much as needed, and in conditions of low cell toxicity, S. cerevisiae will not waste energy making excess glutathione.
Gene Ontology
The Gene Ontology is systematic way to define gene and protein function, and is officiated by the Gene Ontology Consortium. Gene ontology is broken down into three major categories: molecular function, biological process, and cellular component.
Molecular function is defined as the specific activities which the gene products are involved in. Biological process describes the role a gene plays in terms of larger pathways and multi-step procedures. The cellular component is a description of where in an organism the gene products operate.
The molecular function of GSH1 is categorized as "glutamate-cysteine ligase activity." This means that GSH1's product, gamma-glutamylcysteine synthetase, conjoins the two amino acids glutamate and cysteine. The biological processes are "glutathione biosynthesis" and "response to cadmium ion." Thus the ligation of glutamate and cysteine into L-γ-glutamylcysteine is the first step in the synthesis of glutathione. L-γ-glutamylcysteine also interacts with heavy metals, such as cadmium. GSH1 is an intracellular component, and so gamma-glutamylcysteine synthetase works within the cell (Gene Ontology Consortium, 2006).
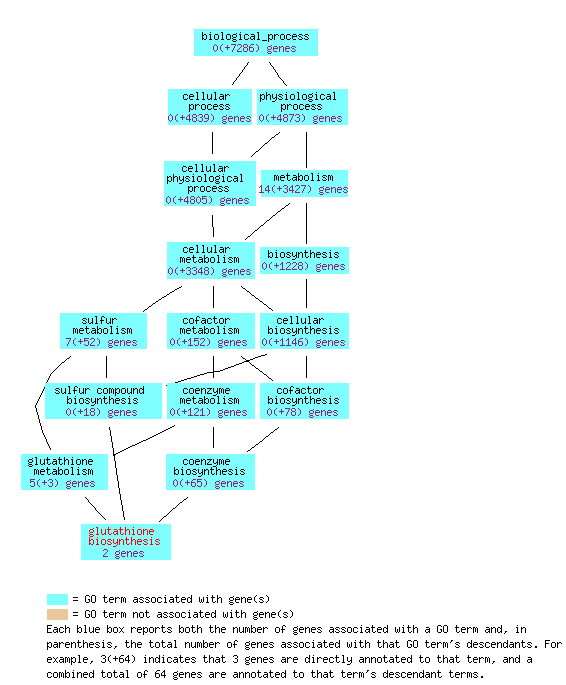
The Gene Ontology Consortium website provides some graphical visualizations of the gene ontologies:
Figure 4: The glutathione biosynthesis biological process overview for GSH1 (Saccharomyces Genome Database, 2006).
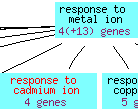
Figure 5: The cadmium ion response biological process overview for GSH1 (Saccharomyces Genome Database, 2006).
There is also a figure for glutamate-cysteine ligase activity, but it is too large to include here. You can access it here: (look to the far right of the page for the box with red lettering) http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=4357 (Saccharomyces Genome Database, 2006).
Now that I have learned about GSH1 and detoxification of yeast cells by the antioxidant glutathione, it is time to move on to my next gene, SET4.
SET4/YJL105W: A Mysterious ORF
Where can I find SET4/YJL105W?
SET4/YJL105W is a gene in S. cerevisiae which has little known about it. Whereas GSH1 has known gene ontologies, synthetic roles, and produces proteins with known functions, SET4 has none of these. SET4 is found close to GSH1 on chromosome X. It was initially predicted to encode an ORF (open reading frame) through the use of computer software. Through the use of various online analytical tools, I will attempt to make some predictions about what role this gene might play in S. cerevisiae.
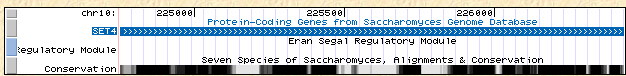
Figure 6: Location and conservation of SET4 on yeast chromosome X (UCSC Genome Browser, 2006).
Nucleotide Sequence
Here is the entire nucleotide sequence for SET4:
1 atgacttcac cggaatcact atcttctcgt catatcaggc aaggaaggac atacacaacc61 acagacaagg tcatatcgcg gtcgtcgtcg tactcatcta atagttcaat gtctaaagat
121 tacggcgatc acacaccctt gtccgtcagc agtgcagctt cagagacatt accctcacct
181 cagtatatgc cgataaggac attcaataca atgcctacag ctggcccaac gcctttacat
241 ttatttcaaa atgacagggg cattttcaac catcattctt catcaggctc atcaaaaacg
301 gcatcaacaa ataaaagagg aatagcagca gcagtagcat tggcaactgc tgccaccata
361 ccatttccac tgaaaaaaca gaatcaagat gataattcca aggtctcggt aacacacaat
421 gaatcatcga aagaaaataa aattacaccc tccatgagag cagaagataa caaacctaaa
481 aatggttgca tctgcggttc aagtgactcc aaggatgagt tgtttataca gtgtaacaaa
541 tgtaaaacgt ggcagcacaa gttatgttat gctttcaaaa aatcagatcc aataaaaaga
601 gattttgttt gcaaaagatg tgacagtgat acgaaagtgc aggttaatca agtaaaacca
661 atgatattcc ctagaaaaat gggagatgag cgattatttc aattttcatc catagtgaca
721 acttcagcat cgaacacaaa tcagcatcaa cagtctgtga ataacataga ggaacagccc
781 aagaaacgtc aacttcatta taccgcccca acaactgaaa atagcaatag tatacggaaa
841 aaattgaggc aagaaaaact ggtagtatca agccactttc tgaagccact actgaatgag
901 gtaagttctt ccaatgacac ggaattcaaa gcaataacaa tatcagagta taaggacaaa
961 tatgttaaga tgtttattga taaccattat gatgacgatt gggttgtttg ttctaactgg
1021 gaaagctcaa ggtcagctga catcgaggta agaaaatcat caaatgaaag agattttgga
1081 gtcttcgctg cagattcttg tgttaaaggt gagctaattc aagaatattt gggcaaaatt
1141 gattttcaaa aaaattatca gacagatcca aataatgact atcgtttgat gggaacgaca
1201 aaacctaaag tactttttca tccacattgg cctttatata tagactctcg agaaacaggc
1261 ggattaacaa gatacataag acggagttgt gagcccaatg tggaactagt aacggtaaga
1321 ccgcttgacg aaaaaccaag aggagataat gattgtagag ttaaatttgt tttaagggct
1381 ataagagata ttcgtaaggg agaagagata agcgtagaat ggcaatggga tttgagaaat
1441 cctatttggg agataataaa tgcatctaaa gatttggatt ccctaccgga tcccgacaag
1501 ttctggttga tggggtcaat aaagactatt ttaacaaatt gtgattgtgc atgtgggtac
1561 ttgggccata attgtccaat aactaaaatc aaaaactttt ctgaagaatt catgaggaat
1621 acgaaggaat ccctatctaa taaatcttac tttaatacaa taatgcacaa ctgtaagcca
1681 taa
Amino Acid Sequence
Here is the amino acid Refseq_012430.1 sequence for SET4:
1 mtspeslssr hirqgrtytt tdkvisrsss yssnssmskd ygdhtplsvs saasetlpsp61 qympirtfnt mptagptplh lfqndrgifn hhsssgsskt astnkrgiaa avalataati
121 pfplkkqnqd dnskvsvthn esskenkitp smraednkpk ngcicgssds kdelfiqcnk
181 cktwqhklcy afkksdpikr dfvckrcdsd tkvqvnqvkp mifprkmgde rlfqfssivt
241 tsasntnqhq qsvnnieeqp kkrqlhytap ttensnsirk klrqeklvvs shflkpllne
301 vsssndtefk aitiseykdk yvkmfidnhy dddwvvcsnw essrsadiev rkssnerdfg
361 vfaadscvkg eliqeylgki dfqknyqtdp nndyrlmgtt kpkvlfhphw plyidsretg
421 gltryirrsc epnvelvtvr pldekprgdn dcrvkfvlra irdirkgeei svewqwdlrn
481 piweiinask dldslpdpdk fwlmgsikti ltncdcacgy lghncpitki knfseefmrn
541 tkeslsnksy fntimhnckp
Predictions
To deduce SET4's function, I began with NCBI's Blast tool, one of the most useful online tools. By comparing the nucleotide or amino acid sequences with other sequences in vast databases, we can determine if similar genes or proteins have already been documented.
Here are the results I get when I do a Blast search on the amino acid sequence:
The first display of the results shows the presence of COGs (Clusters of Orthologous Groups of proteins).
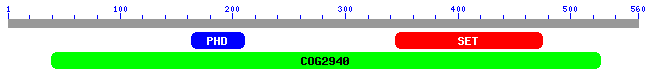
Figure 7: Clusters of Orthologous Groups of proteins matching SET4's amino acid sequence (NCBI Blast, 2006).
The blast search returns 105 matching results (not shown). NCBI also provides another useful visualization tool to see which domains of the protein are conserved. These roughly correspond to the COGs found in figure 5. Some of the results contain both PHD and SET domains, while others contain only one of the two. There are also a number of matches to COG 2940, which are probably all proteins that contain both domains. The first result is the S. cerevisiae protein encoded by SET4.
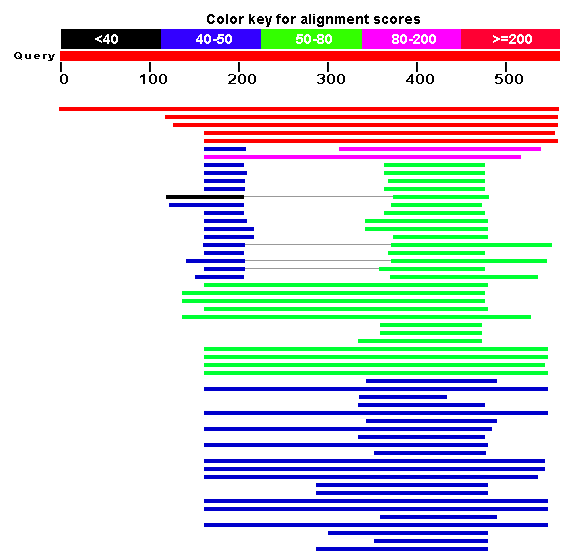
Figure 8: Conserved domains across Blast results (NCBI Blast, 2006).
Of all these results, there is one result in particular which is especially informative. The fifth result, which is colored red, indicating a bit score greater than or equal to 200, is a good place to start the analysis. The match is for another protein in S. cerevisiae! The gene is called SET3, and is on yeast chromosome XI. According to NCBI, the definition for the gene is:
"Defining member of the SET3 histone deacetylase complex which is a meiosis-specific repressor of sporulation genes; necessary for efficient transcription by RNAPII; one of two yeast proteins that contains both SET and PHD domains."
In order to understand this, I need to break it down into smaller pieces. The first part of the sentence tells me that SET3 is a member of a histone deacetylase complex. Histones are proteins that bind DNA, and so this protein must be one of several proteins which remove acetyl groups from the histones. Next, I see that this protein is useful in repressing sporulation genes during meiosis. Yeast is a fungus, and it can produce spores to reproduce. During meiosis, then, this protein must somehow repress the expression of certain sporulation genes.
The next sentence fragment tells me that SET3 is required for transcription by RNA polymerase II. It is also one of two yeast genes which contain SET and PHD domains. As I found out at the beginning of my Blast search, SET4 is the other yeast gene which contains both of these domains. Thus, through a simple online search, I have already learned a lot about SET4 that is not in its official annotation: it has high amino acid sequence conservation with SET3, and SET3 is an annotated S. cerevisiae gene with lots of known functions.
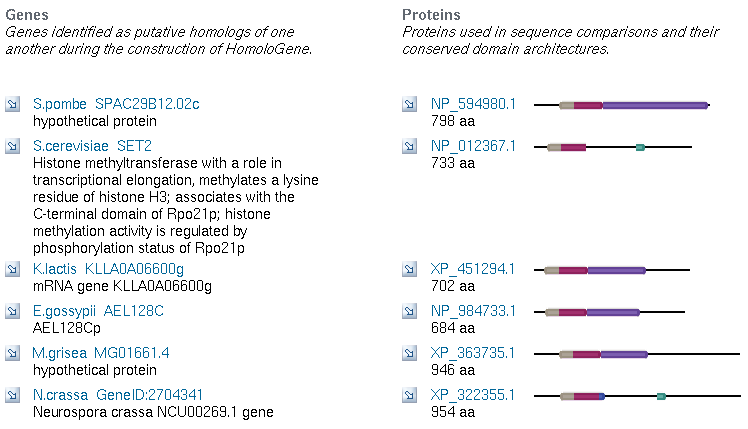
Now that I know SET4 contains some important conserved domains, a SET domain, a PHD domain, and a COG, I want to know more about these. I will start by searching for the COG, COG2940, in HomoloGene. The results give me 5 matches across different ascomycota (sac fungi). The match for S. cerevisiae is SET2, another of the SET genes in its genome. Thus the COG appears to be limited to fungi, as far as we know. The homologs in other species are not particularly informative, and are defined either as hypothetical proteins or as homologs to S. cerevisiae genes.
Figure 9: Homologous Genes for COG2940 across five ascomycota (NCBI Homologene, 2006).
Next, I investigated SET domains. A search in the HomoloGene database yields 8 results for fungi, 4 of which include S. cerevisiae. Two of the proteins are methyltransferases; one of them phosphorylates RNA polymerase II and is involved in glucose repression and telomere maintenance; and one is "involved in regulation of actin cytoskeleton, endocytosis, and viability following starvation or osmotic stress." Thus it appears that SET4 may be a methyltransferase, or it might interact with RNA polymerase II or may regulate the actin cytoskeleton or a starvation stress response. The four search results are:
- HomoloGene:40410. Gene conserved in Ascomycota
- HomoloGene:31699. Gene conserved in Ascomycota
- HomoloGene:74802. Gene conserved in Ascomycota
- HomoloGene:31563. Gene conserved in Ascomycota
Finally, the third domain to look into is the PHD. A search in the HomoloGene database for SET yields nearly 200 results, 7 of which are for fungi. Limiting the results to fungi will help make the search more informative, as I will assume that they are more pertinent to S. cerevisiae. Of the three fungal SET homologous conserved genes, 3 of them are in S. cerevisiae. All three of them involve either binding to histone complexes or gene silencing (NCBI HomoloGene, 2006). This gives a strong indication that SET4 probably binds to histones and may silence genes. The three search results are:
- HomoloGene:40420. Gene conserved in Ascomycota.
- HomoloGene:35593. Gene conserved in Ascomycota.
- HomoloGene:40555. Gene conserved in Ascomycota.
Of course, there are also many other homologous matches to the SET and PHD domains, but it would take a long time to go through them all. I investigated the ones that S. cerevisiae has, assuming that the odds are greater that they are more similar to SET4.
A quick scan down the remaining Blast results does not show much useful information. Most results yield hypothetical proteins, telling me that there is still a lot of research to be done and that it is possible this gene encodes some widespread, fundamentally important protein. I did find a gene in humans, which encodes an "MLL5 protein." The only information of interest this page gives me is that the gene is prominent in placental tissue, and is a putative methyltransferase. Fungi and humans are not closely related, yet both contain homology in their methyltransferases. This is another good indication that SET4 contains a methyltransferase.
Another useful tool at my disposal is the kyte-doolittle hydropathy predictor. This computational tool makes guesses as to whether or not a protein contains transmembrane regions. Using the default settings (window size 9), I find that there is only one small potential transmembrane region. Changing the window size to 12 moves the peak below the threshold level. Thus the protein most likely is not transmembrane, which agrees with the idea that it is found around the chromosome.
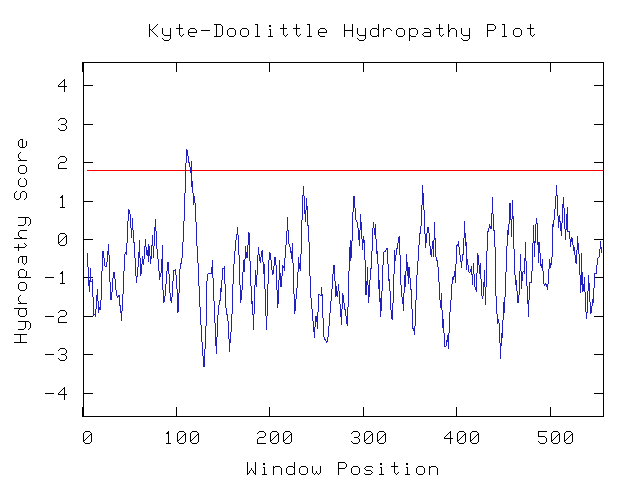
Figure 10: Kyte-Doolittle hydropathy plot (Genomics Consortium at Davidson, 2006).
Another nice tool, called Predator, allows us to predict the 3-d structure of a protein in terms of whether or not it contains α-helices, β-pleated sheets, and so on. This allows us to gain some understanding of the tertiary structure and gives us a feel for qualities such as rigidity and compactness. Here is what Predator predicts for SET4:
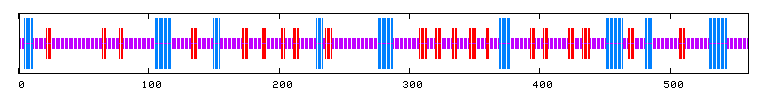
Figure 11: Predator Predicted 3-d structure (Pôle BioInformatique Lyonnais, 2006). 15.00% of the structures are predicted to be alpha helices, 16.07% extended strands, and 68.93% random coils.
At this point it is still difficult to propose a very precise role for SET4. Nevertheless, I will conjecture that it encodes a protein which binds to histone complexes and modifies the presence of either methyl or acetyl groups. SET4 may also serve as a gene silencer. It may have one transmembrane region, and is largely composed of random coils. It is hard to guess beyond this what function this may serve, although epigenetic modifications can have a large impact on processes at the genomic level.
Summary
Through my investigations into yeast gene GSH1, I have learned about S. cerevisiae's mechanism for removing free radicals using glutathione. The cysteine atom in the tripeptide acts as an antioxidant, protecting cells from oxidative damage. GSH1 also modifies cytosolic iron/sulfur proteins post-translationally.
I have also learned about a gene with an unknown function, SET4. Through the use of online tools, such as NCBI Blast, NCBI HomoloGene, and the Kyte-Doolittle hydropathy plot, I have developed a putative function for the protein it encodes. It will take more investigation into S. cerevisiae's chromosomal methylation and histone complexes to test my predictions.
References
Journal Articles
Sipos, Katalin, Heike Lange, Zsuzsanna Fekete, Pascaline Ullmann, Roland Lill, and Gyulla Kispal. Maturation of Cytosolic Iron-Sulfur Proteins Requires Glutathione. Journal of Biological Chemistry. Vol 277, No. 30. pp26944-26949. 2002.
Spector, Daniel, Jean Labarre, and Michael B. Toledano. A Genetic Investigation of the Essential Role of Glutathione: Mutations in the Proline Biosynthesis Pathway are the Only Suppressors of Glutathione Auxotrophy in Yeast. Journal of Biological Chemistry. Vol. 276, No. 10. 2001.
Wheeler, Glen L., Kathryn A. Quinn, Gabriel Perrone, Ian W. Dawes, and Chris M. Grant. Glutathione regulates the expression of γ-glutamylcysteine synthetase via the Met4 transcription factor. Molecular Microbiology. pp545-556. 2002.
Online Abstracts
Dormer UH, Westwater J, Stephen DW, Jamieson DJ. 2002 Jun. Oxidant regulation of the Saccharomyces cerevisiae GSH1 gene [abstract]. In Biochim Biophys Acta. NCBI PubMed database. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=12031480&query_hl=9&itool=pubmed_docsum> Accessed 2006 Oct 5.
Inoue Y, Tran LT, Kamakura M, Izawa S, Miki T, Tsujimoto Y, Kimura A. 1995 Dec. Oxidative stress response in yeast: glutathione peroxidase of Hansenula mrakii is bound to the membrane of both mitochondria and cytoplasm [abstract]. In Biochim Biophys Acta. NCBI PubMed database. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=8541308&query_hl=12&itool=pubmed_DocSum> Accessed 2006 Oct 5.
Amino acid-dependent regulation of the Saccharomyces cerevisiae GSH1 gene by hydrogen peroxide. 1997 Jan. Amino acid-dependent regulation of the Saccharomyces cerevisiae GSH1 gene by hydrogen peroxide [abstract]. In Mol Microbiol. NCBI PubMed Database. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=9044254&query_hl=9&itool=pubmed_docsum> Accessed 2006 Oct 4.
Online References
Genomics Consortium at Davidson. 2006. kyte-doolittle entry form <http://gcat.davidson.edu/DGPB/kd/kyte-doolittle.htm> Accessed 2006 Oct 5
NCBI Blast. 2006. NCBI BLAST. <href="http://ncbi.nlm.nih.gov/blast/> Accessed 2006 Oct 5
NCBI Homologene. 2006. HomoloGene:40410. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=31699> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:31699. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=31699> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:74802. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=74802> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:31563. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=31563> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:40420. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=40420> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:35593. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=35593> Accessed 2006 Oct 5.
NCBI Homologene. 2006. HomoloGene:40555. Gene conserved in Ascomycota. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=homologene&dopt=HomoloGene&list_uids=40555> Accessed 2006 Oct 5.
Pôle BioInformatique Lyonnais. 2006. NPS@ : PREDATOR secondary structure prediction <http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_preda.html> Accessed 2006 Oct 5.
Saccharomyces Genome Database. 2006. Saccharomyces cerevisiae glutathione biosynthesis. <http://pathway.yeastgenome.org:8555/> Accessed 2006 Oct 4.
Saccharomyces Genome Database. 2006. GO term: glutathione biosyntehsis <http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=6750> Accessed 2006 Oct 5.
Saccharomyces Genome Database. 2006. GO term: response to cadmium ion <http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=46686> Accessed 2006 Oct 5.
Saccharomyces Genome Database. 2006. GO term: glutamate-cysteine ligase activity <http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=4357> Accessed 2006 Oct 5.
UCSC Genome Browser v143. 2006. S. Cerevisiae chr10:157,432-293,754. <http://genome.ucsc.edu> Accessed 2006 Oct 5.
UCSC Genome Browser v143. 2006. S. Cerevisiae chr10:234,239-236,273. <http://genome.ucsc.edu> Accessed 2006 Oct 4.
UCSC Genome Browser v143. 2006. S. cerevisiae chr10:223,069-228,117 <http://genome.ucsc.edu/cgi-bin/hgTracks?hgsid=78733252&hgt.in2=+3x+&position=chr10%3A223069-228117> Accessed 2006 Oct 5.
Wikipedia. 2006. Glutathione. <http://en.wikipedia.org/wiki/Glutathione> Accessed 2006 Oct 5.