This web page was produced as an assignment for an undergraduate course
at Davidson College
MacDNAsis Analysis of Histone H1
by David Lamar
MacDNAsis was used to analyze the
structure of human histone H1. Human cDNA for the histone H1 gene
was used. To see the results of a GenBank search for histone
H1 click here.
Figure 1 is the result of analysis
of the human histone H1 cDNA to determine the largest open reading frame
(ORF) for the gene. The blue-highlighted row is the largest ORF.
To view the cDNA and amino acid sequence for the human
histone H1 gene click here.
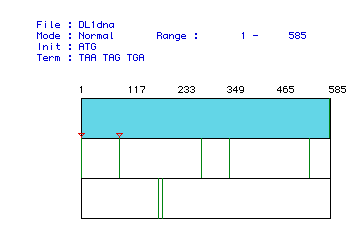
Figure 1: Representation of the Largest Open Reading Frame for Human
Histone H1:
The red triangles indicate start codons and the green lines represent stop
codons. The
blue row indicates the open reading frame from histone H1. The gene
appears to ext-
end from base pair 1 to base pair 585.
The ORF determined from Figure
1 was translated to predict the molecular weight of Histone H1. The
protein was predicted to consist of 195 amino acids and to weigh 20,861.81
daltons (Data not shown).
The predicted amino acid sequence
for histone H1 was used to create a hydropathy plot, an antigenicity plot,
and to predict the protein's secondary structure. Figure 2 is a hydropathy
plot. It can be used to predict whether or not protein is an integral
membrane protein. According to Kyte and Doolittle, a reading above
2.00 on the plot suggests that that domain of the protein is a transmembrane
domain.
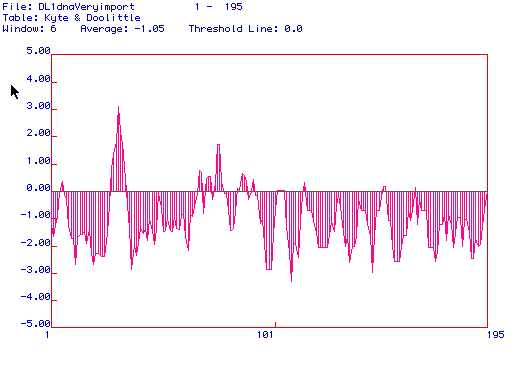
Figure 2: Kyte and Doolittle Hydropathy Plot for Histone
H1: It appears that histone H1 could be
loosely associated to the membrane due to the peak near amino acid 30.
But because this is the
only peak that reaches the threshold and histone H1 is known to be bound
to DNA, it is unlikely
to be an integral membrane protein.
Figure 3 is a Hopp and Woods antigenicity
plot. This plot is used to determine what domains of the protein
would be good candidates against which one could make an effective monoclonal
antibody. The highest peaks indicate the most hydrophilic domains
which would not be likely to be transmembrane domains or tucked in the
interior of the protein. One would assume that domains such as these
would accessible to an antibody in vivo.
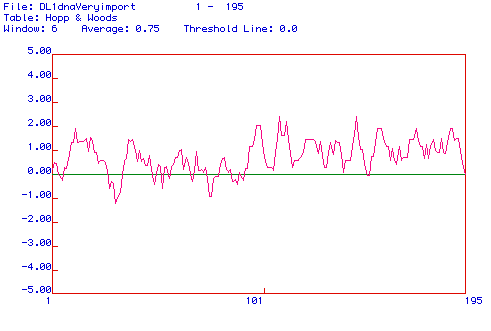
Figure 3: Hopp and Woods Antigenicity Plot for Histone H1:
This plot suggests many
possibilities for an antibody binding domain. The peak near amino
acid 110 and amino
acid 150 are potentially successful domains.
Figure 4 is a Chou, Fasman, and
Rose analysis to predict the secondary structure of histone H1.

Figure 4: Chou, Fasman, and Rose Prediction of Histone H1 Secondary
Structure.
Figure 5 are the results from a
comparison of the amino acid sequences of histone H1 from five different
species: the human (Homo
sapiens), the sea urchin (Echinolampas
crassa), the house mouse (Mus
musculus), the fruit fly (Drosophila
melanogaster), the mussel (Mytilus
edulis). Click on the genus-species name for each animal to view
their histone H1 amino acid sequence.

Figure 5: Comparison of Histone H1 Amino Acid Sequences for Five Different
Species:
The common names of the animals used for the comparison are given to the
right. The
black-outlined, yellow-lettered single letter amino acid symbols indicate
an amino acid
whose place in the sequence is identical with at least one other species.
Figure 6 is a phylogeny of the
five species (Homo
sapiens, Echinolampas
crassa, Mus
musculus, Drosophila
melanogaster, Mytilus
edulis) used in the comparison based on amino acid sequence similarity
for histone H1. Click on the genus-species names above to view the
histone H1 amino acid sequence for each species.
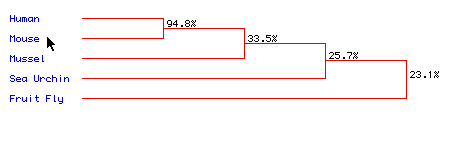
Figure 6: Phylogenetic Tree of Histone H1 similarity Among Five Species:
As
one would expect, human and mouse histone H1 are very similar. All
animals
seems to have high similarity. This could mean that the shape of
histone H1 can
not very greatly and still function correctly.
Back to Top of Page
Molecular Biology Home
Page
Back
to My Home Page

© Copyright 2000 Department of Biology,
Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to:
dalamar@davidson.edu





